Abstract
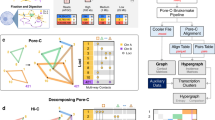
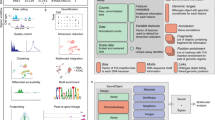
We trained Segway, a dynamic Bayesian network method, simultaneously on chromatin data from multiple experiments, including positions of histone modifications, transcription-factor binding and open chromatin, all derived from a human chronic myeloid leukemia cell line. In an unsupervised fashion, we identified patterns associated with transcription start sites, gene ends, enhancers, transcriptional regulator CTCF-binding regions and repressed regions. Software and genome browser tracks are at http://noble.gs.washington.edu/proj/segway/.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
ENCODE Project Consortium. PLoS Biol. 9, e1001046 (2011).
Day, N., Hemmaplardh, A., Thurman, R.E., Stamatoyannopoulos, J.A. & Noble, W.S. Bioinformatics 23, 1424–1426 (2007).
Erdman, C. & Emerson, J.W. Bioinformatics 24, 2143–2148 (2008).
Jaschek, R. & Tanay, A. in Research in Computational Molecular Biology, Lecture Notes in Computer Science Vol. 5541 (ed. Batzoglou, S.) 170–183 (Springer, Berlin, 2009).
Ernst, J. & Kellis, M. Nat. Biotechnol. 28, 817–825 (2010).
Filion, G.J. et al. Cell 143, 212–224 (2010).
Kharchenko, P.V. et al. Nature 471, 480–485 (2011).
Bilmes, J. & Bartels, C. IEEE Signal Process. Mag. 22, 89–100 (2005).
Reynolds, S.M., Käll, L., Riffle, M.E., Bilmes, J.A. & Noble, W.S. PLOS Comput. Biol. 4, e1000213 (2008).
Wang, Z., Schones, D.E. & Zhao, K. Curr. Opin. Genet. Dev. 19, 127–134 (2009).
Hon, G., Ren, B. & Wang, W. PLOS Comput. Biol. 4, e1000201 (2008).
Raney, B.J. et al. Nucleic Acids Res. 39, D871–D875 (2011).
Hoffman, M.M., Buske, O.J. & Noble, W.S. Bioinformatics 26, 1458–1459 (2010).
Johnson, N.L. Biometrika 36, 149–176 (1949).
Bilmes, J. in UAI '00: Proceedings of the 16th Conference on Uncertainty in Artificial Intelligence (eds. Boutilier, C. & Goldszmidt, M.) 38–45 (Morgan Kaufmann, San Francisco, 2000).
Grundy, W.N., Bailey, T.L., Elkan, C.P. & Baker, M.E. Comput. Appl. Biosci. 13, 397–406 (1997).
Bilmes, J. & Bartels, C. in UAI '03, Proceedings of the 19th Conference in Uncertainty in Artificial Intelligence (eds. Meek, C. & Kjærulff, U.) 47–56 (Morgan Kaufmann Publishers, San Francisco, 2003).
Dempster, A.P., Laird, N.M. & Rubin, D.B. J. Royal Stat. Soc. B 39, 1–22 (1977).
Viterbi, A.J. IEEE Trans. Inf. Theory 13, 260–269 (1967).
Fujita, P.A. et al. Nucleic Acids Res. 39, D876–D882 (2011).
Harrow, J. et al. Genome Biol. 7, S4.1–S4.9 (2006).
Takahashi, H., Kato, S., Murata, M. & Carninci, P. Methods Mol. Biol. 786, 181–200 (2012).
Siepel, A. et al. Genome Res. 15, 1034–1050 (2005).
Buske, O.J., Hoffman, M.M., Ponts, N., Roch, K.G.L. & Noble, W.S. BMC Bioinformatics 12, 415 (2011).
Davis, J. & Goadrich, M. in Proceedings of the 23rd International Conference on Machine Learning 233–240 (ACM, New York, 2006).
Flicek, P. et al. Nucleic Acids Res. 39, D800–D806 (2011).
UniProt Consortium. Nucleic Acids Res. 39, D214–D219 (2011).
Berriz, G.F., Beaver, J.E., Cenik, C., Tasan, M. & Roth, F.P. Bioinformatics 25, 3043–3044 (2009).
Wingender, E. et al. Nucleic Acids Res. 28, 316–319 (2000).
Sandelin, A., Alkema, W., Engstrom, P., Wasserman, W. & Lenhard, B. Nucleic Acids Res. 32, D91–D94 (2004).
Grant, C.E., Bailey, T.L. & Noble, W.S. Bioinformatics 27, 1017–1018 (2011).
Bickel, P.J., Boley, N., Brown, J.B., Huang, H. & Zhang, N.R. Ann. Appl. Stat. 4, 1660–1697 (2010).
Acknowledgements
We thank P.J. Collins for assistance with transient transfection assays, S. Djebali for processing data, C.E. Grant for motif analysis, A. Kundaje for helpful suggestions, and members of the ENCODE Project Consortium, the ENCODE Data Coordination Center and the US National Human Genome Research Institute for providing early public access to the unpublished data used in this work. This work used data produced in the laboratories of B.E. Bernstein (Broad Institute of the Massachusetts Institute of Technology and Harvard University), M.P. Snyder (Stanford University), R.M. Myers (HudsonAlpha Institute for Biotechnology), P.J. Farnham (University of Southern California), V.R. Iyer (University of Texas at Austin), G.E. Crawford (Duke University), J.D. Lieb and T.S. Furey (University of North Carolina at Chapel Hill), J.A. Stamatoyannopoulos (University of Washington), P. Carninci (RIKEN), T.R. Gingeras (Cold Spring Harbor Laboratory), and A. Sidow (Stanford University). This publication was made possible by grants 004695, 004561 and 006259 from National Human Genome Research Institute.
Author information
Authors and Affiliations
Contributions
M.M.H., W.S.N. and J.A.B. conceived of the project; M.M.H., W.S.N. and Z.W. designed computational and biological experiments. M.M.H., J.A.B., O.J.B. and J.W. developed software used in this work; M.M.H., O.J.B. and J.W. conducted computational experiments and analyzed data; and M.M.H., W.S.N., Z.W., J.A.B., O.J.B. and J.W. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–11, Supplementary Tables 1–4, Supplementary Results, Supplementary Discussion (PDF 1780 kb)
Rights and permissions
About this article
Cite this article
Hoffman, M., Buske, O., Wang, J. et al. Unsupervised pattern discovery in human chromatin structure through genomic segmentation. Nat Methods 9, 473–476 (2012). https://doi.org/10.1038/nmeth.1937
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.1937