Abstract
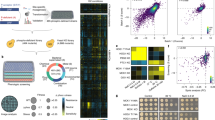
The functional role of protein phosphorylation is impacted by its fractional stoichiometry. Thus, a comprehensive strategy to study phosphorylation dynamics should include an assessment of site stoichiometry. Here we report an integrated method that relies on phosphatase treatment and stable-isotope labeling to determine absolute stoichiometries of protein phosphorylation on a large scale. This approach requires the measurement of only a single ratio relating phosphatase-treated and mock-treated samples. Using this strategy we determined stoichiometries for 5,033 phosphorylation sites in triplicate analyses from Saccharomyces cerevisiae growing through mid-log phase. We validated stoichiometries at ten sites that represented the full range of values obtained using synthetic phosphopeptides and found excellent agreement. Using bioinformatics, we characterized the biological properties associated with phosphorylation sites with vastly differing absolute stoichiometries.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Grimsrud, P.A., Swaney, D.L., Wenger, C.D., Beauchene, N.A. & Coon, J.J. Phosphoproteomics for the masses. ACS Chem. Biol. 5, 105–119 (2010).
Macek, B., Mann, M. & Olsen, J.V. Global and site-specific quantitative phosphoproteomics: principles and applications. Annu. Rev. Pharmacol. Toxicol. 49, 199–221 (2009).
Thingholm, T.E., Jensen, O.N. & Larsen, M.R. Analytical strategies for phosphoproteomics. Proteomics 9, 1451–1468 (2009).
Boersema, P.J., Mohammed, S. & Heck, A.J.R. Phosphopeptide fragmentation and analysis by mass spectrometry. J. Mass Spectrom. 44, 861–878 (2009).
Witze, E.S., Old, W.M., Resing, K.A. & Ahn, N.G. Mapping protein post-translational modifications with mass spectrometry. Nat. Methods 4, 798–806 (2007).
Choudhary, C. et al. Mislocalized activation of oncogenic RTKs switches downstream signaling outcomes. Mol. Cell 36, 326–339 (2009).
Holt, L.J. et al. Global analysis of Cdk1 substrate phosphorylation sites provides insights into evolution. Science 325, 1682–1686 (2009).
Huttlin, E.L. et al. A tissue-specific atlas of mouse protein phosphorylation and expression. Cell 143, 1174–1189 (2010).
Olsen, J.V. et al. Quantitative phosphoproteomics reveals widespread full phosphorylation site occupancy during mitosis. Sci. Signal. 3, ra3 (2010).
Wu, R. et al. Correct interpretation of comprehensive phosphorylation dynamics requires normalization by protein expression changes. Mol. Cell. Proteomics published online, doi:10.1074/mcp.M111.009654 (7 May 2011).
Cooper, J.A. & Hunter, T. Identification and characterization of cellular targets for tyrosine protein kinases. J. Biol. Chem. 258, 1108–1115 (1983).
Stukenberg, P.T. et al. Systematic identification of mitotic phosphoproteins. Curr. Biol. 7, 338–348 (1997).
Gerber, S.A., Rush, J., Stemman, O., Kirschner, M.W. & Gygi, S.P. Absolute quantification of proteins and phosphoproteins from cell lysates by tandem MS. Proc. Natl. Acad. Sci. USA 100, 6940–6945 (2003).
Steen, H., Jebanathirajah, J.A., Springer, M. & Kirschner, M.W. Stable isotope-free relative and absolute quantitation of protein phosphorylation stoichiometry by MS. Proc. Natl. Acad. Sci. USA 102, 3948–3953 (2005).
Carr, S.A., Huddleston, M.J. & Annan, R.S. Selective detection and sequencing of phosphopeptides at the femtomole level by mass spectrometry. Anal. Biochem. 239, 180–192 (1996).
Guo, L. et al. Studies of ligand-induced site-specific phosphorylation of epidermal growth factor receptor. J. Am. Soc. Mass Spectrom. 14, 1022–1031 (2003).
Steen, J.A.J. et al. Different phosphorylation states of the anaphase promoting complex in response to antimitotic drugs: a quantitative proteomic analysis. Proc. Natl. Acad. Sci. USA 105, 6069–6074 (2008).
Jin, L.L. et al. Measurement of protein phosphorylation stoichiometry by selected reaction monitoring mass spectrometry. J. Proteome Res. 9, 2752–2761 (2010).
Stemmann, O., Zou, H., Gerber, S.A., Gygi, S.P. & Kirschner, M.W. Dual inhibition of sister chromatid separation at metaphase. Cell 107, 715–726 (2001).
Atrih, A. et al. Stoichiometric quantification of Akt phosphorylation using LC-MS/MS. J. Proteome Res. 9, 743–751 (2010).
Zhang, X., Jin, Q.K., Carr, S.A. & Annan, R.S. N-terminal peptide labeling strategy for incorporation of isotopic tags: a method for the determination of site-specific absolute phosphorylation stoichiometry. Rapid Commun. Mass Spectrom. 16, 2325–2332 (2002).
Bonenfant, D. et al. Quantitation of changes in protein phosphorylation: a simple method based on stable isotope labeling and mass spectrometry. Proc. Natl. Acad. Sci. USA 100, 880–885 (2003).
Domanski, D., Murphy, L.C. & Borchers, C.H. Assay development for the determination of phosphorylation stoichiometry using multiple reaction monitoring methods with and without phosphatase treatment: application to breast cancer signaling pathways. Anal. Chem. 82, 5610–5620 (2010).
Hegeman, A.D., Harms, A.C., Sussman, M.R., Bunner, A.E. & Harper, J.F. An isotope labeling strategy for quantifying the degree of phosphorylation at multiple sites in proteins. J. Am. Soc. Mass Spectrom. 15, 647–653 (2004).
Johnson, H., Eyers, C.E., Eyers, P.A., Beynon, R.J. & Gaskell, S.J. Rigorous determination of the stoichiometry of protein phosphorylation using mass spectrometry. J. Am. Soc. Mass Spectrom. 20, 2211–2220 (2009).
Kanshin, E. et al. The stoichiometry of protein phosphorylation in adipocyte lipid droplets: Analysis by N-terminal isotope tagging and enzymatic dephosphorylation. Proteomics 9, 5067–5077 (2009).
Pflieger, D. et al. Quantitative proteomic analysis of protein complexes. Mol. Cell. Proteomics 7, 326–346 (2008).
Previs, M.J. et al. Quantification of protein phosphorylation by liquid chromatography-mass spectrometry. Anal. Chem. 80, 5864–5872 (2008).
Broberg, A. High-performance liquid chromatography/electrospray ionization ion-trap mass spectrometry for analysis of oligosaccharides derivatized by reductive amination and N,N-dimethylation. Carbohydr. Res. 342, 1462–1469 (2007).
Elias, J.E. & Gygi, S.P. Target-decoy search strategy for increased confidence in large-scale protein identifications by mass spectrometry. Nat. Methods 4, 207–214 (2007).
Albuquerque, C.P. et al. A multidimensional chromatography technology for in-depth phosphoproteome analysis. Mol. Cell. Proteomics 7, 1389–1396 (2008).
Gnad, F. et al. High-accuracy identification and bioinformatic analysis of in vivo protein phosphorylation sites in yeast. Proteomics 9, 4642–4652 (2009).
Li, X. et al. Large-scale phosphorylation analysis of alpha-factor-arrested Saccharomyces cerevisiae. J. Proteome Res. 6, 1190–1197 (2007).
Peng, K., Radivojac, P., Vucetic, S., Dunker, A.K. & Obradovic, Z. Length-dependent prediction of protein intrinsic disorder. BMC Bioinformatics 7, 208 (2006).
Taylor, S.S., Knighton, D.R., Zheng, J.H., Teneyck, L.F. & Sowadski, J.M. Structural framework for the protein-kinase family. Annu. Rev. Cell Biol. 8, 429–462 (1992).
Huang, W., Sherman, B.T. & Lempicki, R.A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Guerra, B. & Issinger, O.G. Protein kinase CK2 in human diseases. Curr. Med. Chem. 15, 1870–1886 (2008).
Haas, W. et al. Optimization and use of peptide mass measurement accuracy in shotgun proteomics. Mol. Cell. Proteomics 5, 1326–1337 (2006).
Eng, J.K., McCormack, A.L. & Yates, J.R. An approach to correlate tandem mass-spectral data of peptides with amino-acid-sequences in a protein database. J. Am. Soc. Mass Spectrom. 5, 976–989 (1994).
Bakalarski, C.E. et al. The impact of peptide abundance and dynamic range on stable-isotope-based quantitative proteomic analyses. J. Proteome Res. 7, 4756–4765 (2008).
Villen, J. & Gygi, S.P. The SCX/IMAC enrichment approach for global phosphorylation analysis by mass spectrometry. Nat. Protoc. 3, 1630–1638 (2008).
Acknowledgements
This work was supported in part by US National Institutes of Health grants (HG3456) to S.P.G. We thank all members of the Gygi lab for help, especially R.A. Everley for his help with instrumentation and L. Ting for critically reading the manuscript.
Author information
Authors and Affiliations
Contributions
S.P.G. and R.W. designed the research. R.W., W.H., N.D., E.L.H., B.Z., M.E.S. and S.P.G. participated in the data generation, analysis and interpretation. R.W. and S.P.G. wrote the manuscript and all authors edited it.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–4, Supplementary Tables 1 and 5 (PDF 381 kb)
Supplementary Table 2
Peptides identified in experiment 1. (XLSX 14310 kb)
Supplementary Table 3
Peptides identified in experiment 2. (XLSX 14521 kb)
Supplementary Table 4
Peptides identified in experiment 3. (XLSX 15000 kb)
Supplementary Table 6
Site stoichiometries obtained in biological triplicate experiments. (XLSX 584 kb)
Supplementary Table 7
Site stoichiometries for events described as Cdk1substrates. (XLSX 21 kb)
Supplementary Table 8
Examples of phosphorylation sites with high stoichiometries. (XLSX 11 kb)
Rights and permissions
About this article
Cite this article
Wu, R., Haas, W., Dephoure, N. et al. A large-scale method to measure absolute protein phosphorylation stoichiometries. Nat Methods 8, 677–683 (2011). https://doi.org/10.1038/nmeth.1636
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.1636
This article is cited by
-
A quantitative and site-specific atlas of the citrullinome reveals widespread existence of citrullination and insights into PADI4 substrates
Nature Structural & Molecular Biology (2024)
-
A streamlined tandem tip-based workflow for sensitive nanoscale phosphoproteomics
Communications Biology (2023)
-
A time-resolved multi-omics atlas of Acanthamoeba castellanii encystment
Nature Communications (2022)
-
Global profiling of distinct cysteine redox forms reveals wide-ranging redox regulation in C. elegans
Nature Communications (2021)
-
Wittig reagents for chemoselective sulfenic acid ligation enables global site stoichiometry analysis and redox-controlled mitochondrial targeting
Nature Chemistry (2021)