Abstract
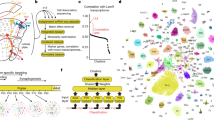
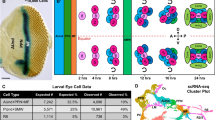
We developed a multicolor neuron labeling technique in Drosophila melanogaster that combines the power to specifically target different neural populations with the label diversity provided by stochastic color choice. This adaptation of vertebrate Brainbow uses recombination to select one of three epitope-tagged proteins detectable by immunofluorescence. Two copies of this construct yield six bright, separable colors. We used Drosophila Brainbow to study the innervation patterns of multiple antennal lobe projection neuron lineages in the same preparation and to observe the relative trajectories of individual aminergic neurons. Nerve bundles, and even individual neurites hundreds of micrometers long, can be followed with definitive color labeling. We traced motor neurons in the subesophageal ganglion and correlated them to neuromuscular junctions to identify their specific proboscis muscle targets. The ability to independently visualize multiple lineage or neuron projections in the same preparation greatly advances the goal of mapping how neurons connect into circuits.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
Change history
16 February 2011
In the version of this article initially published online, accession codes were not included. The error has been corrected for the print, PDF and HTML versions of this article.
30 August 2012
The schematic of the dBrainbow vector construct presented in the original Figure 1 of the paper and its description had incorrect epitope tags. This error has not been corrected in the HTML or PDF versions of the article, but a correct version of the figure as well as full details of the error can be found in the corrigendum.
03 August 2015
In the version of this article initially published, the sequence reported for the dBrainbow construct was incorrect. The blue fluorescent protein was reported as EBFP2; it is mTFP1. The mKO2 protein is tagged with a V5 epitope and not a Myc epitope, and the EGFP protein is tagged with HSV, not with V5, as detailed in a previous correction to this paper. The red channel reflects endogenous mKO2 fluorescence, not anti-Myc staining. The mKO2 protein is produced by a codon-optimized sequence. These errors have been corrected in the HTML and PDF versions of the article, and the correct sequence is provided.
References
Lichtman, J.W., Livet, J. & Sanes, J.R. A technicolour approach to the connectome. Nat. Rev. Neurosci. 9, 417–422 (2008).
Simpson, J.H. Mapping and manipulating neural circuits in the fly brain. Adv. Genet. 65, 79–143 (2009).
Bellen, H.J., Tong, C. & Tsuda, H. 100 years of Drosophila research and its impact on vertebrate neuroscience: a history lesson for the future. Nat. Rev. Neurosci. 11, 514–522 (2010).
Lai, S.L. & Lee, T. Genetic mosaic with dual binary transcriptional systems in Drosophila. Nat. Neurosci. 9, 703–709 (2006).
Yu, H.H., Chen, C.H., Shi, L., Huang, Y. & Lee, T. Twin-spot MARCM to reveal the developmental origin and identity of neurons. Nat. Neurosci. 12, 947–953 (2009).
Livet, J. et al. Transgenic strategies for combinatorial expression of fluorescent proteins in the nervous system. Nature 450, 56–62 (2007).
Sakaue-Sawano, A. et al. Visualizing spatiotemporal dynamics of multicellular cell-cycle progression. Cell 132, 487–498 (2008).
Ai, H.-W., Henderson, J.N., Remington, S.J. & Campbell, R.E. Directed evolution of a monomeric, bright and photostable version of Clavularia cyan fluorescent protein: structural characterization and applications in fluorescence imaging. Biochem. J. 400, 531–540 (2006).
Busch, S., Selcho, M., Ito, K. & Tanimoto, H. A map of octopaminergic neurons in the Drosophila brain. J. Comp. Neurol. 513, 643–667 (2009).
Masse, N.Y., Turner, G.C. & Jefferis, G.S. Olfactory information processing in Drosophila. Curr. Biol. 19, R700–R713 (2009).
Jefferis, G.S., Marin, E.C., Stocker, R.F. & Luo, L. Target neuron prespecification in the olfactory map of Drosophila. Nature 414, 204–208 (2001).
Marin, E.C., Jefferis, G.S., Komiyama, T., Zhu, H. & Luo, L. Representation of the glomerular olfactory map in the Drosophila brain. Cell 109, 243–255 (2002).
Wong, A.M., Wang, J.W. & Axel, R. Spatial representation of the glomerular map in the Drosophila protocerebrum. Cell 109, 229–241 (2002).
Lai, S.L., Awasaki, T., Ito, K. & Lee, T. Clonal analysis of Drosophila antennal lobe neurons: diverse neuronal architectures in the lateral neuroblast lineage. Development 135, 2883–2893 (2008).
Siegal, M.L. & Hartl, D.L. Transgene Coplacement and high efficiency site-specific recombination with the Cre/loxP system in Drosophila. Genetics 144, 715–726 (1996).
Heidmann, D. & Lehner, C.F. Reduction of Cre recombinase toxicity in proliferating Drosophila cells by estrogen-dependent activity regulation. Dev. Genes Evol. 211, 458–465 (2001).
Jefferis, G.S. et al. Comprehensive maps of Drosophila higher olfactory centers: spatially segregated fruit and pheromone representation. Cell 128, 1187–1203 (2007).
Ito, K. & Awasaki, T. Clonal unit architecture of the adult fly brain. in Brain Development in Drosophila melanogaster Vol. 628. (ed., G.M. Technau) 137–159 (Landes Bioscience and Springer Science and Business Media, 2008).
Lee, T., Lee, A. & Luo, L. Development of the Drosophila mushroom bodies: sequential generation of three distinct types of neurons from a neuroblast. Development 126, 4065–4076 (1999).
Siegal, M.L. & Hartl, D.L. Application of Cre/loxP in Drosophila. Site-specific recombination and transgene coplacement. Methods Mol. Biol. 136, 487–495 (2000).
Yagi, R., Mayer, F. & Basler, K. Refined LexA transactivators and their use in combination with the Drosophila Gal4 system. Proc. Natl. Acad. Sci. USA 107, 16166–16171 (2010).
de Velasco, B. et al. Specification and development of the pars intercerebralis and pars lateralis, neuroendocrine command centers in the Drosophila brain. Dev. Biol. 302, 309–323 (2007).
Potter, C.J. & Luo, L. Octopamine fuels fighting flies. Nat. Neurosci. 11, 989–990 (2008).
Busch, S. & Tanimoto, H. Cellular configuration of single octopamine neurons in Drosophila. J. Comp. Neurol. 518, 2355–2364 (2010).
Manoli, D.S. et al. Male-specific fruitless specifies the neural substrates of Drosophila courtship behaviour. Nature 436, 395–400 (2005).
Demir, E. & Dickson, B.J. Fruitless splicing specifies male courtship behavior in Drosophila. Cell 121, 785–794 (2005).
Kimura, K., Hachiya, T., Koganezawa, M., Tazawa, T. & Yamamoto, D. Fruitless and doublesex coordinate to generate male-specific neurons that can initiate courtship. Neuron 59, 759–769 (2008).
Cachero, S., Ostrovsky, A.D., Yu, J.Y., Dickson, B.J. & Jefferis, G.S. Sexual dimorphism in the fly brain. Curr. Biol. 20, 1589–1601 (2010).
Yu, J.Y., Kanai, M.I., Demir, E., Jefferis, G.S. & Dickson, B.J. Cellular organization of the neural circuit that drives Drosophila courtship behavior. Curr. Biol. 20, 1602–1614 (2010).
Isono, K. & Morita, H. Molecular and cellular designs of insect taste receptor system. Front. Cell Neurosci. 4, 20 (2010).
Vosshall, L.B. & Stocker, R.F. Molecular architecture of smell and taste in Drosophila. Annu. Rev. Neurosci. 30, 505–533 (2007).
Rajashekhar, K.P. & Singh, R.N. Neuroarchitecture of the tritocerebrum of Drosophila melanogaster. J. Comp. Neurol. 349, 633–645 (1994).
Tissot, M., Gendre, N. & Stocker, R.F. Drosophila P[Gal4] lines reveal that motor neurons involved in feeding persist through metamorphosis. J. Neurobiol. 37, 237–250 (1998).
Gordon, M.D. & Scott, K. Motor Control in a Drosophila Taste Circuit. Neuron 61, 373–384 (2009).
Miller, A. The internal anatomy and histology of the imago of Drosophila melanogaster. in Biology of Drosophila. (ed., Demerec, M.) 420–534 (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York, USA, 1950).
Brizzard, B. Epitope tagging. Biotechniques 44, 693–695 (2008).
Pfeiffer, B.D. et al. Tools for neuroanatomy and neurogenetics in Drosophila. Proc. Natl. Acad. Sci. USA 105, 9715–9720 (2008).
Groth, A.C., Fish, M., Nusse, R. & Calos, M.P. Construction of transgenic Drosophila by using the site-specific integrase from phage phiC31. Genetics 166, 1775–1782 (2004).
Rodin, S. & Georgiev, P. Handling three regulatory elements in one transgene: combined use of cre-lox, FLP-FRT, and I-Scel recombination systems. Biotechniques 39, 871–876 (2005).
Stocker, R.F., Heimbeck, G., Gendre, N. & de Belle, J.S. Neuroblast ablation in Drosophila P[GAL4] lines reveals origins of olfactory interneurons. J. Neurobiol. 32, 443–456 (1997).
Cole, S.H. et al. Two functional but noncomplementing Drosophila tyrosine decarboxylase genes: distinct roles for neural tyramine and octopamine in female fertility. J. Biol. Chem. 280, 14948–14955 (2005).
Connolly, J.B. et al. Associative learning disrupted by impaired Gs signaling in Drosophila mushroom bodies. Science 274, 2104–2107 (1996).
Wu, J.S. & Luo, L. A protocol for dissecting Drosophila melanogaster brains for live imaging or immunostaining. Nat. Protoc. 1, 2110–2115 (2006).
Panchuk-Voloshina, N. et al. Alexa dyes, a series of new fluorescent dyes that yield exceptionally bright, photostable conjugates. J. Histochem. Cytochem. 47, 1179–1188 (1999).
Acknowledgements
We thank A. Arnold, E. Shumsky and A. Soell for help with imaging and spectral separation; B. Pfeiffer, A. Nern and G. Rubin (Janelia Farm Research Campus) for sharing vectors and recombinase lines before publication and for the gift of R12D05-GAL4; V. Hartenstein, B. Gerber, T. Lee, B. Baker and P. Keller for helpful discussions about biological applications of dBrainbow; S. Albin, A. Seeds and E. Hoopfer for scientific discussion of the project; D. Grover for statistical advice; and K. Basler, R. Yagi and C. Lehner (University of Zurich) and M. Siegal (New York University) for additional Cre lines.
Author information
Authors and Affiliations
Contributions
S.H. designed and performed cloning, tested constructs in S2 cells and made the figures. P.C. performed the fly genetics, immunohistochemistry and confocal imaging. C.E.M. generated and analyzed the subesophageal ganglion and proboscis data. D.H. generated the recombinant fly stocks. L.L.L. advised on selection of fluorescent proteins, construct design and the conversion from endogenous fluorescence to antibody. J.H.S. conceived the project, cloned initial test constructs and wrote the paper with help from S.H., P.C., C.E.M. and L.L.L.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–4 and Supplementary Tables 1–4 (PDF 4504 kb)
Rights and permissions
About this article
Cite this article
Hampel, S., Chung, P., McKellar, C. et al. Drosophila Brainbow: a recombinase-based fluorescence labeling technique to subdivide neural expression patterns. Nat Methods 8, 253–259 (2011). https://doi.org/10.1038/nmeth.1566
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.1566
This article is cited by
-
Modelling TDP-43 proteinopathy in Drosophila uncovers shared and neuron-specific targets across ALS and FTD relevant circuits
Acta Neuropathologica Communications (2023)
-
The insect somatostatin pathway gates vitellogenesis progression during reproductive maturation and the post-mating response
Nature Communications (2022)
-
Multicolor strategies for investigating clonal expansion and tissue plasticity
Cellular and Molecular Life Sciences (2022)
-
Deciphering neural heterogeneity through cell lineage tracing
Cellular and Molecular Life Sciences (2021)
-
SPARC enables genetic manipulation of precise proportions of cells
Nature Neuroscience (2020)