Abstract
We developed an automated system, ScanLag, that measures in parallel the delay in growth (lag time) and growth rate of thousands of cells. Using ScanLag, we detected small subpopulations of bacteria with dramatically increased lag time upon starvation. By screening a library of Escherichia coli deletion mutants, we achieved two-dimensional mapping of growth characteristics, which showed that ScanLag enables multidimensional screens for quantitative characterization and identification of rare phenotypic variants.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Collins, S.R., Weissman, J.S. & Krogan, N.J. Nat. Methods 6, 721–723 (2009).
Bochner, B.R. FEMS Microbiol. Rev. 33, 191–205 (2009).
Elfwing, A., LeMarc, Y., Baranyi, J. & Ballagi, A. Appl. Environ. Microbiol. 70, 675–678 (2004).
Balaban, N.Q., Merrin, J., Chait, R., Kowalik, L. & Leibler, S. Science 305, 1622–1625 (2004).
Guillier, L., Pardon, P. & Augustin, J.C. Appl. Environ. Microbiol. 71, 2940–2948 (2005).
Glaser, D.A. & Wattenburg, W.H. Ann. NY Acad. Sci. 139, 243–257 (1966).
Michel, J.B., Yeh, P.J., Chait, R., Moellering, R.C. & Kishony, R. Proc. Natl. Acad. Sci. USA 105, 14918–14923 (2008).
Barkai, N. & Shilo, B.Z. Mol. Cell 28, 755–760 (2007).
Dubnau, D. & Losick, R. Mol. Microbiol. 61, 564–572 (2006).
Bigger, J.W. Lancet 244, 497–500 (1944).
Hershey, A.D. J. Bacteriol. 37, 285–299 (1939).
Pin, C. & Baranyi, J. Appl. Environ. Microbiol. 74, 2534–2536 (2008).
Niven, G.W., Morton, J.S., Fuks, T. & Mackey, B.A. Appl. Environ. Microbiol. 74, 3757–3763 (2008).
Kato, J. & Hashimoto, M. Mol. Syst. Biol. 3, 132 (2007).
Kaprelyants, A.S. et al. Intercellular Signalling and the Multiplication of Prokaryotes: Bacterial Cytokines (Cambridge University Press, 1999).
Lutz, R. & Bujard, H. Nucleic Acids Res. 25, 1203–1210 (1997).
Gefen, O., Gabay, C., Mumcuoglu, M., Engel, G. & Balaban, N.Q. Proc. Natl. Acad. Sci. USA 105, 6145–6149 (2008).
Hurley, M.A. & Roscoe, M.E. J. Appl. Bacteriol. 55, 159–164 (1983).
Cochran, W.G. Biometrics 6, 105–116 (1950).
Quake, S.R. & Scherer, A. Science 290, 1536–1540 (2000).
Acknowledgements
We thank A. Keynan for invaluable discussions on the importance of the lag phase, D. Azulay and A. Bar-Yaacov for helpful theoretical discussions, I. Rosenshine (Hebrew University) for strains, and the National BioResource Project and National Institute of Genetics, Japan for the large and medium deletion mutants library, S. Silbert for initial software development and E. Rotem and S. Pearl for technical support and comments on the manuscript. The work was supported by the Human Frontier Science Program and by the Israel Science Foundation.
Author information
Authors and Affiliations
Contributions
I.L.-R., O.G., I.R. and D.S. performed the research. O.F. and H.S. contributed analytical tools. N.Q.B., I.L.-R., I.R. and O.G. designed the research. N.Q.B. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and Supplementary Data (PDF 383 kb)
Supplementary Video 1
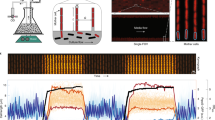
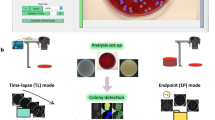
A typical analysis process using ScanLag. Shown are analyses of overnight culture (left) and culture starved for 3 d (right). Colony appearance was automatically detected as soon as colonies reached a threshold size. Each colony was assigned an identifying number and an arbitrary color. The lower panels show the resulting distribution, for each plate, represented as 1-CDF (cumulative distribution function), which represents the fraction of the colonies that have not yet appeared. At the beginning of the movie, no colony has appeared yet, namely that fraction is 1, whereas at the end of the movie it is zero. (MOV 2108 kb)
Supplementary Software 1
ScanningManager software application controlling image acquisition by an array of scanners. (ZIP 534 kb)
Supplementary Software 2
TimeLapse analysis software application (Matlab) for the analysis of images and extraction of colony appearance data. (ZIP 57 kb)
Rights and permissions
About this article
Cite this article
Levin-Reisman, I., Gefen, O., Fridman, O. et al. Automated imaging with ScanLag reveals previously undetectable bacterial growth phenotypes. Nat Methods 7, 737–739 (2010). https://doi.org/10.1038/nmeth.1485
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.1485
This article is cited by
-
Bacterial colony size growth estimation by deep learning
BMC Microbiology (2023)
-
Dynamic proteome trade-offs regulate bacterial cell size and growth in fluctuating nutrient environments
Communications Biology (2023)
-
Pareto optimality between growth-rate and lag-time couples metabolic noise to phenotypic heterogeneity in Escherichia coli
Nature Communications (2021)
-
Observation of universal ageing dynamics in antibiotic persistence
Nature (2021)
-
Efficient microbial colony growth dynamics quantification with ColTapp, an automated image analysis application
Scientific Reports (2020)