Abstract
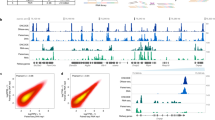
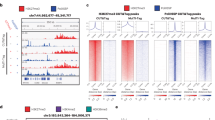
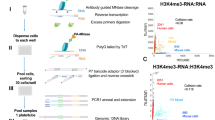
Current methods for whole-genome mapping of protein-DNA interactions, performed by coupling chromatin immunoprecipitation with high-throughput sequencing (ChIP-seq), require large amounts of starting materials, which precludes their application to rare cell types. Here we combine a high-sensitivity ChIP assay with a new library preparation procedure to map histone modifications in as few as 10,000 cells. We used the technique to characterize mouse hematopoietic progenitors and thereby gain insight into their developmental program.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
References
Park, P.J. Nat. Rev. Genet. 10, 669–680 (2009).
Acevedo, L.G. et al. Biotechniques 43, 791–797 (2007).
O'Neill, L.P., VerMilyea, M.D. & Turner, B.M. Nat. Genet. 38, 835–841 (2006).
Attema, J.L. et al. Proc. Natl. Acad. Sci. USA 104, 12371–12376 (2007).
Dahl, J.A. & Collas, P. Nucleic Acids Res. 36, e15 (2008).
Goren, A. et al. Nat. Methods 7, 47–49 (2010).
Lieb, J.D., Liu, X., Botstein, D. & Brown, P.O. Nat. Genet. 28, 327–334 (2001).
Mikkelsen, T.S. et al. Nature 448, 553–560 (2007).
Li, B., Carey, M. & Workman, J.L. Cell 128, 707–719 (2007).
Tothova, Z. et al. Cell 128, 325–339 (2007).
Azuara, V. et al. Nat. Cell Biol. 8, 532–538 (2006).
Bernstein, B.E. et al. Cell 125, 315–326 (2006).
Cui, K. et al. Cell Stem Cell 4, 80–93 (2009).
Weishaupt, H., Sigvardsson, M. & Attema, J.L. Blood 115, 247–256 (2010).
Wei, G. et al. Immunity 30, 155–167 (2009).
Chambers, S.M. et al. Cell Stem Cell 1, 578–591 (2007).
Bottardi, S., Ghiam, A.F., Bergeron, F. & Milot, E. Cell Cycle 6, 1035–1039 (2007).
Uchida, N., Aguila, H.L., Fleming, W.H., Jerabek, L. & Weissman, I.L. Blood 83, 3758–3779 (1994).
Morrison, S.J. & Weissman, I.L. Immunity 1, 661–673 (1994).
Varnum-Finney, B., Dallas, M.H., Kato, K. & Bernstein, I.D. Blood 111, 2615–2620 (2008).
Acknowledgements
We thank D. Flowers and I. Bernstein for expertise, reagents and assistance in isolation of hematopoietic cells, N. Shoresh and T. Mikkelsen for computational assistance and B. Knoechel, E. Mendenhall, A. Goren and A. Chi for constructive discussions and critical reading of the manuscript. This research was supported by funds from the Starr Cancer Consortium, a Charles E. Culpeper Scholarship, the US National Human Genome Research Institute, the National Institutes of Health Roadmap for Epigenomics and the National Heart, Lung, and Blood Institute.
Author information
Authors and Affiliations
Contributions
M.A. and B.E.B. designed the method. M.A. performed the experiments. M.A., J.Z. and B.E.B. analyzed the data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
M.A. and B.E.B. have filed a patent application describing the methods presented here (US 12/699,508).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and Supplementary Tables 1–3 (PDF 4521 kb)
Rights and permissions
About this article
Cite this article
Adli, M., Zhu, J. & Bernstein, B. Genome-wide chromatin maps derived from limited numbers of hematopoietic progenitors. Nat Methods 7, 615–618 (2010). https://doi.org/10.1038/nmeth.1478
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.1478
This article is cited by
-
Histone variant H3.3 maintains adult haematopoietic stem cell homeostasis by enforcing chromatin adaptability
Nature Cell Biology (2022)
-
Understanding histone H3 lysine 36 methylation and its deregulation in disease
Cellular and Molecular Life Sciences (2019)
-
Cell Type-Specific Survey of Epigenetic Modifications by Tandem Chromatin Immunoprecipitation Sequencing
Scientific Reports (2018)
-
Purification of nanogram-range immunoprecipitated DNA in ChIP-seq application
BMC Genomics (2017)
-
Live cell imaging of low- and non-repetitive chromosome loci using CRISPR-Cas9
Nature Communications (2017)