Abstract
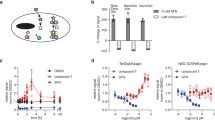
The ability to measure multiple cellular signaling events is essential to better understand the underlying complex biological processes that occur in living cells. Microarray-based technologies are now commonly used to study changes in transcription. This information, however, is not sufficient to understand the regulatory mechanisms that lead to gene expression changes. Here we present an approach to monitor signaling events upstream of gene expression. We coupled different reporter gene assays to unique expressed oligonucleotide tags (EXTs) that serve as identifiers and quantitative reporters. Multiple EXT reporters can be isolated as a pool and analyzed by hybridization to microarrays. To test the feasibility of our approach, we integrated complementation assays based on a protease from tobacco etch virus (TEV protease) and transcription factor activity profiling. Thereby, we simultaneously monitored Neuregulin-dependent mouse ErbB receptor tyrosine kinase dimerization, effector recruitment and downstream signaling.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Papin, J.A., Hunter, T., Palsson, B.O. & Subramaniam, S. Reconstruction of cellular signalling networks and analysis of their properties. Nat. Rev. Mol. Cell Biol. 6, 99–111 (2005).
Citri, A. & Yarden, Y. EGF-ERBB signalling: towards the systems level. Nat. Rev. Mol. Cell Biol. 7, 505–516 (2006).
Mei, L. & Xiong, W.C. Neuregulin 1 in neural development, synaptic plasticity and schizophrenia. Nat. Rev. Neurosci. 9, 437–452 (2008).
Jones, R.B., Gordus, A., Krall, J.A. & MacBeath, G. A quantitative protein interaction network for the ErbB receptors using protein microarrays. Nature 439, 168–174 (2006).
Schulze, W.X., Deng, L. & Mann, M. Phosphotyrosine interactome of the ErbB-receptor kinase family. Mol Syst Biol 1, 0008 (2005).
Krutzik, P.O. & Nolan, G.P. Fluorescent cell barcoding in flow cytometry allows high-throughput drug screening and signaling profiling. Nat. Methods 3, 361–368 (2006).
Glory, E. & Murphy, R.F. Automated subcellular location determination and high-throughput microscopy. Dev. Cell 12, 7–16 (2007).
MacDonald, M.L. et al. Identifying off-target effects and hidden phenotypes of drugs in human cells. Nat. Chem. Biol. 2, 329–337 (2006).
Wu, R.Z., Bailey, S.N. & Sabatini, D.M. Cell-biological applications of transfected-cell microarrays. Trends Cell Biol. 12, 485–488 (2002).
Wehr, M.C. et al. Monitoring regulated protein-protein interactions using split TEV. Nat. Methods 3, 985–993 (2006).
Wehr, M.C., Reinecke, L., Botvinnik, A. & Rossner, M.J. Analysis of transient phosphorylation-dependent protein-protein interactions in living mammalian cells using split-TEV. BMC Biotechnol. 8, 55 (2008).
Bulyk, M.L. DNA microarray technologies for measuring protein-DNA interactions. Curr. Opin. Biotechnol. 17, 422–430 (2006).
Wang, Z., Gerstein, M. & Snyder, M. RNA-seq: a revolutionary tool for transcriptomics. Nat. Rev. Genet. 10, 57–63 (2009).
Romanov, S. et al. Homogeneous reporter system enables quantitative functional assessment of multiple transcription factors. Nat. Methods 5, 253–260 (2008).
Brenner, S. et al. In vitro cloning of complex mixtures of DNA on microbeads: physical separation of differentially expressed cDNAs. Proc. Natl. Acad. Sci. USA 97, 1665–1670 (2000).
Vaskovsky, A., Lupowitz, Z., Erlich, S. & Pinkas-Kramarski, R. ErbB-4 activation promotes neurite outgrowth in PC12 cells. J. Neurochem. 74, 979–987 (2000).
Di Fiore, P.P. et al. erbB-2 is a potent oncogene when overexpressed in NIH/3T3 cells. Science 237, 178–182 (1987).
Xu, Q., Schlabach, M.R., Hannon, G.J. & Elledge, S.J. Design of 240,000 orthogonal 25mer DNA barcode probes. Proc. Natl. Acad. Sci. USA 106, 2289–2294 (2009).
Mazurkiewicz, P., Tang, C.M., Boone, C. & Holden, D.W. Signature-tagged mutagenesis: barcoding mutants for genome-wide screens. Nat. Rev. Genet. 7, 929–939 (2006).
Kovacs, D.M. & Kaplan, B.B. Discordant estimates of heterologous promoter activity as determined by reporter gene mRNA levels and enzyme activity. Biochem. Biophys. Res. Commun. 189, 912–918 (1992).
Barreau, C., Paillard, L. & Osborne, H.B. AU-rich elements and associated factors: are there unifying principles? Nucleic Acids Res. 33, 7138–7150 (2005).
Deplancke, B., Dupuy, D., Vidal, M. & Walhout, A.J. A gateway-compatible yeast one-hybrid system. Genome Res. 14, 2093–2101 (2004).
Fields, S. & Song, O. A novel genetic system to detect protein-protein interactions. Nature 340, 245–246 (1989).
Kanno, A., Ozawa, T. & Umezawa, Y. Intein-mediated reporter gene assay for detecting protein-protein interactions in living mammalian cells. Anal. Chem. 78, 556–560 (2006).
Stagljar, I., Korostensky, C., Johnsson, N. & te Heesen, S. A genetic system based on split-ubiquitin for the analysis of interactions between membrane proteins in vivo. Proc. Natl. Acad. Sci. USA 95, 5187–5192 (1998).
Eyckerman, S. et al. Design and use of a mammalian protein-protein interaction trap (MAPPIT). Sci. STKE 2002, pl18 (2002).
Barnea, G. et al. The genetic design of signaling cascades to record receptor activation. Proc. Natl. Acad. Sci. USA 105, 64–69 (2008).
Nomenclature committee of the International Union of Biochemistry (NC-IUB). Nomenclature for incompletely specified bases in nucleic acid sequences. Recommendations 1984. J. Biol. Chem. 261, 13–17 (1986).
Matys, V. et al. TRANSFAC and its module TRANSCompel: transcriptional gene regulation in eukaryotes. Nucleic Acids Res. 34, D108–D110 (2006).
Gossen, M. & Bujard, H. Tight control of gene expression in mammalian cells by tetracycline-responsive promoters. Proc. Natl. Acad. Sci. USA 89, 5547–5551 (1992).
Rossner, M.J. et al. Global transcriptome analysis of genetically identified neurons in the adult cortex. J. Neurosci. 26, 9956–9966 (2006).
Acknowledgements
We acknowledge the contributions of F. Benseler and all members of the Sequencing Core Facility (Max Planck Institute of Experimental Medicine, Götttingen) for the EXT synthesis and excellent services, and the expert support by staff of the Microarray Core Lab of the University of Göttingen, namely R. Hitt, G. Salinas-Riester and L. Opitz. We thank R. Reinhardt, S. Klages and S. Scheer for high-throughput sequencing and primary data analysis, K.-A. Nave for support, F. Melchior, E. Wimmer, as well as all lab members for stimulating discussions, and M. Wehr for critically reading the manuscript and providing valuable feedback. This study was supported by grants of the Bundesministerium für Bildung und Forschung (FKZ0315180A) and partially by the European Union (LSHM-CT-2005-018637) to M.J.R.
Author information
Authors and Affiliations
Contributions
A.B. cloned EXT constructs, performed the assays, analyzed the data, contributed conceptually and wrote the manuscript. S.P.W. performed bioinformatic calculations and analyzed the data. T.M.F. cloned expression constructs. M.J.R. conceived the study, analyzed the data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
M.J.R. filed a patent application covering the principles of EXTassays.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–15, Supplementary Tables 1–5, Supplementary Note 1 and Supplementary Discussion (PDF 3237 kb)
Rights and permissions
About this article
Cite this article
Botvinnik, A., Wichert, S., Fischer, T. et al. Integrated analysis of receptor activation and downstream signaling with EXTassays. Nat Methods 7, 74–80 (2010). https://doi.org/10.1038/nmeth.1407
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.1407
This article is cited by
-
Multiplexed profiling of GPCR activities by combining split TEV assays and EXT-based barcoded readouts
Scientific Reports (2018)
-
Pathway sensor-based functional genomics screening identifies modulators of neuronal activity
Scientific Reports (2018)