Abstract
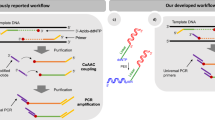
Here we report a highly efficient and simplified strategy to preadenylate bar-coded oligonucleotides designed for microRNA (miRNA) capture and multiplex analysis. Using this approach, we enzymatically preadenylated bar-coded oligonucleotides with high efficiency when compared to the chemical method currently used by miRNA investigators. As a case study, we used these oligonucleotides in an ATP-independent ligation to miRNAs, suggesting the utility of our method in end-capture protocols and high-throughput sequencing applications.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Ambros, V. Nature 431, 350–355 (2004).
Bartel, D.P. Cell 116, 281–297 (2004).
Ambros, V. & Chen, X. Development 134, 1635–1641 (2007).
Lau, N.C. et al. Science 294, 858–862 (2001).
Pfeffer, S., Lagos-Quintana, M. & Tuschl, T. Curr. Protoc. Mol. Biol. 26.4 (2005).
Pak, J. & Fire, A. Science 315, 241–244 (2007).
Wood, Z.A., Sabatini, R.S. & Hajduk, S.L. Mol. Cell 13, 455–456 (2004).
Lehman, I.R. Science 186, 790–797 (1974).
Ohtsuka, E. et al. Nucleic Acids Res. 3, 1613–1623 (1976).
McLaughlin, L.W., Piel, N. & Graeser, E. Biochemistry 24, 267–273 (1985).
Patel, M.P., Baum, D.A. & Silverman, S.K. Bioorg. Chem. 36, 46–56 (2008).
Wang, Y. & Silverman, S.K. RNA 12, 1142–1146 (2006).
Chiuman, W. & Li, Y. Bioorg. Chem. 30, 332–349 (2002).
Silverman, S.K. RNA 10, 731–746 (2004).
Acknowledgements
This work was supported by the Center for Excellence in Genome Sciences grant from the US National Human Genome Research Institute. We thank A.Z. Fire for helpful discussions and assistance in this project. F.V. is supported by a Canadian Institutes of Health Research Fellowship and A.M.S. is supported by the US National Institutes of Health Ruth L. Kirschstein Fellowship.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–2, Supplementary Table 1, Supplementary Methods (PDF 601 kb)
Rights and permissions
About this article
Cite this article
Vigneault, F., Sismour, A. & Church, G. Efficient microRNA capture and bar-coding via enzymatic oligonucleotide adenylation. Nat Methods 5, 777–779 (2008). https://doi.org/10.1038/nmeth.1244
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.1244
This article is cited by
-
Capture and sequencing of NAD-capped RNA sequences with NAD captureSeq
Nature Protocols (2017)
-
Substantial and robust changes in microRNA transcriptome support postnatal development of the hypothalamus in rat
Scientific Reports (2016)
-
Efficient synthesis of stably adenylated DNA and RNA adapters for microRNA capture using T4 RNA ligase 1
Scientific Reports (2015)
-
Abiotic stress-associated microRNAs in plants: discovery, expression analysis, and evolution
Frontiers in Biology (2013)
-
A cost-effective method for Illumina small RNA-Seq library preparation using T4 RNA ligase 1 adenylated adapters
Plant Methods (2012)