Abstract
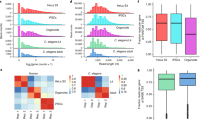
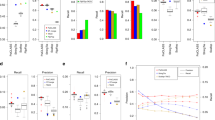
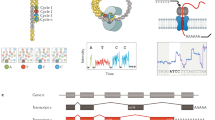
We have mapped and quantified mouse transcriptomes by deeply sequencing them and recording how frequently each gene is represented in the sequence sample (RNA-Seq). This provides a digital measure of the presence and prevalence of transcripts from known and previously unknown genes. We report reference measurements composed of 41–52 million mapped 25-base-pair reads for poly(A)-selected RNA from adult mouse brain, liver and skeletal muscle tissues. We used RNA standards to quantify transcript prevalence and to test the linear range of transcript detection, which spanned five orders of magnitude. Although >90% of uniquely mapped reads fell within known exons, the remaining data suggest new and revised gene models, including changed or additional promoters, exons and 3′ untranscribed regions, as well as new candidate microRNA precursors. RNA splice events, which are not readily measured by standard gene expression microarray or serial analysis of gene expression methods, were detected directly by mapping splice-crossing sequence reads. We observed 1.45 × 105 distinct splices, and alternative splices were prominent, with 3,500 different genes expressing one or more alternate internal splices.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Casneuf, T., Van de Peer, Y. & Huber, W. In situ analysis of cross-hybridisation on microarrays and inference of expression correlation. BMC Bioinformatics 8, 461 (2007).
Eklund, A.C. et al. Replacing cRNA targets with cDNA reduces microarray cross-hybridization. Nat. Biotechnol. 24, 1071–1073 (2006).
Okoniewski, M.J. & Miller, C.J. Hybridization interactions between probesets in short oligo microarrays lead to spurious correlations. BMC Bioinformatics 7, 276 (2006).
Velculescu, V.E., Zhang, L., Vogelstein, B. & Kinzler, K. Serial analysis of gene expression. Science 270, 484–487 (1995).
Harbers, M. & Carninci, P. Tag-based approaches for transcriptome research and genome annotation. Nat. Methods 2, 495–502 (2005).
Brenner, S. et al. Gene expression analysis by massively parallel signature sequencing (MPSS) on microbead arrays. Nat. Biotechnol. 18, 630–634 (2000).
Boguski, M.S. & Toltoshev, C.M. Gene discovery in dbEST. Science 265, 1993–1994 (1994).
Gerhard, D.S. et al. The status, quality, and expansion of the NIH full-length cDNA project: The Mammalian Gene Collection (MGC). Genome Res. 14, 2121–2127 (2004).
Dias Neto, E.D. et al. Shotgun sequencing of the human transcriptome with ORF expressed sequence tags. Proc. Natl. Acad. Sci. USA 97, 3491–3496 (2000).
Bertone, P. et al. Global identification of human transcribed sequences with genome tiling arrays. Science 306, 2242–2246 (2004).
Cheng, J. et al. Transcription maps of 10 human chromosomes at 5-nucleotide resolution. Science 308, 1149–1154 (2005).
Kapranov, P. et al. RNA maps reveal new RNA classes and a possible function for pervasive transcription. Science 316, 1484–1488 (2007).
Royce, T.E. et al. Issues in the analysis of oligonucleotide tiling microarrays for transcript mapping. Trends Genet. 21, 466–475 (2005).
Kapranov, P., Willingham, A.T. & Gingeras, T.R. Genome-wide transcription and the implications for genomic organization. Nat. Rev. Genet. 8, 413–423 (2007).
Nagalakshmi, U. et al. The transcriptional landscape of the yeast genome defined by RNA sequencing. Science published online, doi:10.1126/science.1158441 (1 May 2008).
Wilhelm, B.T. et al. Dynamic repertoire of a eukaryotic transcriptome surveyed at single nucleotide resolution. Nature advance online publication, doi:10.1038/nature07002 (2008).
Lister, R. et al. Highly integrated single-base resolution maps of the epigenome in Arabidopsis. Cell 133, 523–536 (2008).
Altschul, S.F., Gish, W., Miller, W., Myers, E.W. & Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Galau, G.A., Klein, W.H., Britten, R.J. & Davidson, E.H. Significance of rare mRNA sequences in liver. Arch. Biochem. Biophys. 179, 584–599 (1977).
Kapur, K., Xing, Y., Ouyang, Z. & Wong, W.H. Exon arrays provide accurate assessments of gene expression. Genome Biol. 8, R82 (2007).
Lee, C. & Roy, M. Analysis of alternative splicing with microarrays: successes and challenges. Genome Biol. 5, 231 (2004).
Johnson, J.M. et al. Genome-wide survey of human alternative pre-mRNA splicing with exon junction microarrays. Science 302, 2141–2144 (2003).
Martin, J.F. et al. A Mef2 gene that generates a muscle-specific isoform via alternative mRNA splicing. Mol. Cell. Biol. 14, 1647–1656 (1994).
Acknowledgements
This work was supported by The Beckman Foundation, The Simons Foundation and US National Institutes of Health (NIH) grant U54 HG004576 to B.W. and R. Myers. A.M. was supported by an NIH training grant. The authors especially thank D. Trout and B. King for professional data handling and G. Schroth, I. Khrebtukova and S. Luo, of Illumina, for exchanges of preliminary data and protocols under development. M. Liu and J.L. Riechmann, along with others from the laboratories of B. Wold, R. Myers, J. Allman and P. Sternberg, are gratefully acknowledged for many helpful discussions, as are R. Myers and S. Mango for manuscript assistance.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6, Supplementary Tables 1–4 and Supplementary Methods (PDF 500 kb)
Supplementary Dataset 1
Intermediate and final RPKM values for mouse brain. (TXT 1429 kb)
Supplementary Dataset 2
Intermediate and final RPKM values for mouse liver. (TXT 1423 kb)
Supplementary Dataset 3
Intermediate and final RPKM values for mouse muscle. (TXT 1423 kb)
Supplementary Dataset 4
Top 500 genes with strong multiread contributions in mouse liver. (XLS 52 kb)
Rights and permissions
About this article
Cite this article
Mortazavi, A., Williams, B., McCue, K. et al. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat Methods 5, 621–628 (2008). https://doi.org/10.1038/nmeth.1226
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.1226
This article is cited by
-
GTax: improving de novo transcriptome assembly by removing foreign RNA contamination
Genome Biology (2024)
-
Enhanced protein secretion in reduced genome strains of Streptomyces lividans
Microbial Cell Factories (2024)
-
Analysis of differentially expressed genes in torn rotator cuff tendon tissues in diabetic patients through RNA-sequencing
BMC Musculoskeletal Disorders (2024)
-
Full-length transcriptome and RNA-Seq analyses reveal the resistance mechanism of sesame in response to Corynespora cassiicola
BMC Plant Biology (2024)
-
Comparative physiological, metabolomic and transcriptomic analyses reveal the mechanisms of differences in pear fruit quality between distinct training systems
BMC Plant Biology (2024)