Abstract
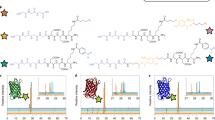
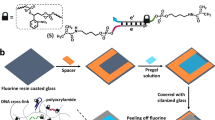
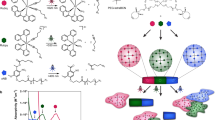
Although biochemically patterned hydrogels are capable of recapitulating many critical aspects of the heterogeneous cellular niche, exercising spatial and temporal control of the presentation and removal of biomolecular signalling cues in such systems has proved difficult. Here, we demonstrate a synthetic strategy that exploits two bioorthogonal photochemistries to achieve reversible immobilization of bioactive full-length proteins with good spatial and temporal control within synthetic, cell-laden biomimetic scaffolds. A photodeprotection–oxime-ligation sequence permits user-defined quantities of proteins to be anchored within distinct subvolumes of a three-dimensional matrix, and an ortho-nitrobenzyl ester photoscission reaction facilitates subsequent protein removal. By using this approach to pattern the presentation of the extracellular matrix protein vitronectin, we accomplished reversible differentiation of human mesenchymal stem cells to osteoblasts in a spatially defined manner. Our protein-patterning approach should provide further avenues to probe and direct changes in cell physiology in response to dynamic biochemical signalling.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
DeForest, C. A. & Anseth, K. S. Advances in bioactive hydrogels to probe and direct cell fate. Annu. Rev. Chem. Biomol. 3, 421–444 (2012).
Langer, R. & Tirrell, D. A. Designing materials for biology and medicine. Nature 428, 487–492 (2004).
Lutolf, M. P., Gilbert, P. M. & Blau, H. M. Designing materials to direct stem-cell fate. Nature 462, 433–441 (2009).
Engler, A. J., Sen, S., Sweeney, H. L. & Discher, D. E. Matrix elasticity directs stem cell lineage specification. Cell 126, 677–689 (2006).
Huebsch, N. et al. Harnessing traction-mediated manipulation of the cell/matrix interface to control stem-cell fate. Nature Mater. 9, 518–526 (2010).
Yang, C., Tibbitt, M. W., Basta, L. & Anseth, K. S. Mechanical memory and dosing influence stem cell fate. Nature Mater. 13, 645–652 (2014).
Chen, C. S., Mrksich, M., Huang, S., Whitesides, G. M. & Ingber, D. E. Geometric control of cell life and death. Science 276, 1425–1428 (1997).
Petersen, O. W., Ronnovjessen, L., Howlett, A. R. & Bissell, M. J. Interaction with basement-membrane serves to rapidly distinguish growth and differentiation pattern of normal and malignant human breast epithelial cells. Proc. Natl Acad. Sci. USA 89, 9064–9068 (1992).
Burdick, J. A. & Murphy, W. L. Moving from static to dynamic complexity in hydrogel design. Nature Commun. 3, 1269 (2012).
Lutolf, M. P. & Hubbell, J. A. Synthetic biomaterials as instructive extracellular microenvironments for morphogenesis in tissue engineering. Nature Biotechnol. 23, 47–55 (2005).
Rompolas, P., Mesa, K. R. & Greco, V. Spatial organization within a niche as a determinant of stem-cell fate. Nature 502, 513–518 (2013).
Katz, J. S. & Burdick, J. A. Light-responsive biomaterials: Development and applications. Macromol. Biosci. 10, 339–348 (2010).
DeLong, S. A., Moon, J. J. & West, J. L. Covalently immobilized gradients of bFGF on hydrogel scaffolds for directed cell migration. Biomaterials 26, 3227–3234 (2005).
Hahn, M. S., Miller, J. S. & West, J. L. Three-dimensional biochemical and biomechanical patterning of hydrogels for guiding cell behavior. Adv. Mater. 18, 2679–2684 (2006).
DeForest, C. A., Polizzotti, B. D. & Anseth, K. S. Sequential click reactions for synthesizing and patterning three-dimensional cell microenvironments. Nature Mater. 8, 659–664 (2009).
DeForest, C. A., Sims, E. A. & Anseth, K. S. Peptide-functionalized click hydrogels with independently tunable mechanics and chemical functionality for 3D cell culture. Chem. Mater. 22, 4783–4790 (2010).
DeForest, C. A. & Anseth, K. S. Cytocompatible click-based hydrogels with dynamically tunable properties through orthogonal photocoupling and photodegradation reactions. Nature Chem. 3, 925–931 (2011).
Luo, Y. & Shoichet, M. S. A photolabile hydrogel for guided three-dimensional cell growth and migration. Nature Mater. 3, 249–253 (2004).
Wylie, R. G. et al. Spatially controlled simultaneous patterning of multiple growth factors in three-dimensional hydrogels. Nature Mater. 10, 799–806 (2011).
Adzima, B. J. et al. Spatial and temporal control of the alkyne–azide cycloaddition by photoinitiated Cu(II) reduction. Nature Chem. 3, 258–261 (2011).
Mosiewicz, K. A. et al. In situ cell manipulation through enzymatic hydrogel photopatterning. Nature Mater. 12, 1072–1078 (2013).
Kloxin, A. M., Kasko, A. M., Salinas, C. N. & Anseth, K. S. Photodegradable hydrogels for dynamic tuning of physical and chemical properties. Science 324, 59–63 (2009).
Wirkner, M. et al. Triggered cell release from materials using bioadhesive photocleavable linkers. Adv. Mater. 23, 3907–3910 (2011).
Griffin, D. R. et al. Synthesis of photodegradable macromers for conjugation and release of bioactive molecules. Biomacromolecules 14, 1199–1207 (2013).
Kloxin, A. M., Tibbitt, M. W. & Anseth, K. S. Synthesis of photodegradable hydrogels as dynamically tunable cell culture platforms. Nature Protoc. 5, 1867–1887 (2010).
Fairbanks, B. D., Singh, S. P., Bowman, C. N. & Anseth, K. S. Photodegradable, photoadaptable hydrogels via radical-mediated disulfide fragmentation reaction. Macromolecules 44, 2444–2450 (2011).
Griffin, D. R. & Kasko, A. M. Photodegradable macromers and hydrogels for live cell encapsulation and release. J. Am. Chem. Soc. 134, 13103–13107 (2012).
Azagarsamy, M. A., Alge, D. L., Radhakrishnan, S. J., Tibbitt, M. W. & Anseth, K. S. Photocontrolled nanoparticles for on-demand release of proteins. Biomacromolecules 13, 2219–2224 (2012).
He, M., Li, J., Tan, S., Wang, R. & Zhang, Y. Photodegradable supramolecular hydrogels with fluorescence turn-on reporter for photomodulation of cellular microenvironments. J. Am. Chem. Soc. 135, 18718–18721 (2013).
DeForest, C. A. & Anseth, K. S. Photoreversible patterning of biomolecules within click-based hydrogels. Angew. Chem. Int. Ed. 51, 1816–1819 (2011).
Lutolf, M. P. Materials science: Cell environments programmed with light. Nature 482, 477–478 (2012).
Alge, D. L. & Anseth, K. S. Bioactive hydrogels: Lighting the way. Nature Mater. 12, 950–952 (2013).
Agard, N. J., Prescher, J. A. & Bertozzi, C. R. A strain-promoted [3 + 2] azide-alkyne cycloaddition for covalent modification of biomolecules in living systems. J. Am. Chem. Soc. 126, 15046–15047 (2004).
Lutolf, M. P. et al. Synthetic matrix metalloproteinase-sensitive hydrogels for the conduction of tissue regeneration: Engineering cell-invasion characteristics. Proc. Natl Acad. Sci. USA 100, 5413–5418 (2003).
Patterson, J. & Hubbell, J. A. Enhanced proteolytic degradation of molecularly engineered PEG hydrogels in response to MMP-1 and MMP-2. Biomaterials 31, 7836–7845 (2010).
Codelli, J. A., Baskin, J. M., Agard, N. J. & Berozzi, C. R. Second-generation difluorinated cyclooctynes for copper-free click chemistry. J. Am. Chem. Soc. 130, 11486–11493 (2008).
Sims, E. A., DeForest, C. A. & Anseth, K. S. A mild, large-scale synthesis of 1,3-cyclooctanedione: Expanding access to difluorinated cyclooctyne for copper-free click chemistry. Tetrahedron Lett. 52, 1871–1873 (2011).
Ning, X. H., Guo, J., Wolfert, M. A. & Boons, G. J. Visualizing metabolically labeled glycoconjugates of living cells by copper-free and fast huisgen cycloadditions. Angew. Chem. Int. Ed. 47, 2253–2255 (2008).
Debets, M. F. et al. Aza-dibenzocyclooctynes for fast and efficient enzyme PEGylation via copper-free (3 + 2) cycloaddition. Chem. Commun. 46, 97–99 (2010).
Tibbitt, M. W., Kloxin, A. M., Sawicki, L. A. & Anseth, K. S. Mechanical properties and degradation of chain and step-polymerized photodegradable hydrogels. Macromolecules 46, 2785–2792 (2013).
Dendane, N. et al. Efficient surface patterning of oligonucleotides inside a glass capillary through oxime bond formation. Bioconjug. Chem. 18, 671–676 (2007).
Park, S. & Yousaf, M. N. An interfacial oxime reaction to immobilize ligands and cells in patterns and gradients to photoactive surfaces. Langmuir 24, 6201–6207 (2008).
Woll, D. et al. Intramolecular sensitization of photocleavage of the photolabile 2-(2-nitrophenyl)propoxycarbonyl (NPPOC) protecting group: Photoproducts and photokinetics of the release of nucleosides. Chem. Eur. J. 14, 6490–6497 (2008).
Odian, G. G. Principles of Polymerization 4th edn (Wiley-Interscience, 2004).
Jevševar, S., Kunstelj, M. & Porekar, V. G. PEGylation of therapeutic proteins. Biotechnol. J. 5, 113–128 (2010).
Dirksen, A. & Dawson, P. E. Rapid oxime and hydrazone ligations with aromatic aldehydes for biomolecular labeling. Bioconjug. Chem. 19, 2543–2548 (2008).
Blanden, A. R., Mukherjee, K., Dilek, O., Loew, M. & Bane, S. L. 4-aminophenylalanine as a biocompatible nucleophilic catalyst for hydrazone ligations at low temperature and neutral pH. Bioconjug. Chem. 22, 1954–1961 (2011).
Zustiak, S. P. & Leach, J. B. Characterization of protein release from hydrolytically degradable poly(ethylene glycol) hydrogels. Biotechnol. Bioeng. 108, 197–206 (2011).
Phillies, G. D. Diffusion of bovine serum albumin in a neutral polymer solution. Biopolymers 24, 379–386 (1985).
Kopan, R. & Ilagan, M. X. G. The canonical Notch signaling pathway: Unfolding the activation mechanism. Cell 137, 216–233 (2009).
Beckstead, B. L., Santosa, D. M. & Giachelli, C. M. Mimicking cell–cell interactions at the biomaterial-cell interface for control of stem cell differentiation. J. Biomed. Mater. Res. A 79A, 94–103 (2006).
Varnum-Finney, B. et al. Immobilization of Notch ligand, Delta-1, is required for induction of Notch signaling. J. Cell Sci. 113, 4313–4318 (2000).
Khetan, S. et al. Degradation-mediated cellular traction directs stem cell fate in covalently crosslinked three-dimensional hydrogels. Nature Mater. 12, 458–465 (2013).
Hayman, E. G., Pierschbacher, M. D., Ohgren, Y. & Ruoslahti, E. Serum spreading factor (vitronectin) is present at the cell surface and in tissues. Proc. Natl Acad. Sci. USA 80, 4003–4007 (1983).
Lin, C. C., Metters, A. T. & Anseth, K. S. Functional PEG-peptide hydrogels to modulate local inflammation induced by the pro-inflammatory cytokine TNFα. Biomaterials 30, 4907–4914 (2009).
Acknowledgements
The authors thank P. Rapp for discussions on FRAP and FEM analysis, as well as for his constructive comments on the manuscript; M. Shahgholi and N. Torian for assistance with HRMS; J. Heath, J. Pfeilsticker and R. Henning for advice on peptide work and for use of their peptide synthesizer and HPLC; D. Koos and the Caltech Biological Imaging Center for use of confocal microscopes; and K. Beres and C. Murry for assistance with all Notch-related studies. This work was supported by the National Science Foundation Grant NSF-DMR 1206121 and a University of Washington Faculty Startup Grant (C.A.D.).
Author information
Authors and Affiliations
Contributions
C.A.D. and D.A.T. designed the experiments. C.A.D. conducted the experiments. C.A.D. and D.A.T. interpreted the data and composed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Information (PDF 9484 kb)
Supplementary Information
Supplementary Movie 1 (WMV 7072 kb)
Supplementary Information
Supplementary Movie 2 (WMV 6900 kb)
Supplementary Information
Supplementary Movie 3 (WMV 4594 kb)
Supplementary Information
Supplementary Movie 4 (WMV 8097 kb)
Rights and permissions
About this article
Cite this article
DeForest, C., Tirrell, D. A photoreversible protein-patterning approach for guiding stem cell fate in three-dimensional gels. Nature Mater 14, 523–531 (2015). https://doi.org/10.1038/nmat4219
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmat4219
This article is cited by
-
Middle-out methods for spatiotemporal tissue engineering of organoids
Nature Reviews Bioengineering (2023)
-
Cell–extracellular matrix mechanotransduction in 3D
Nature Reviews Molecular Cell Biology (2023)
-
Tricolor visible wavelength-selective photodegradable hydrogel biomaterials
Nature Communications (2023)
-
Next-generation cancer organoids
Nature Materials (2022)
-
Sensitizer-enhanced two-photon patterning of biomolecules in photoinstructive hydrogels
Communications Materials (2022)