Abstract
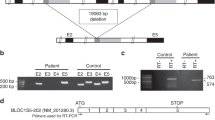
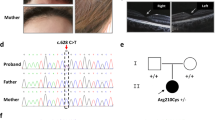
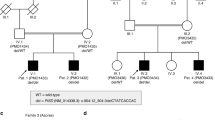
Hermansky-Pudlak syndrome (HPS) is a disorder of organelle biogenesis in which oculocutaneous albinism, bleeding and pulmonary fibrosis result from defects of melanosomes, platelet dense granules and lysosomes1,2,3,4. HPS is common in Puerto Rico5,6, where it is caused by mutations in the genes HPS17 and, less often, HPS3 (ref. 8). In contrast, only half of non–Puerto Rican individuals with HPS have mutations in HPS1 (ref. 9), and very few in HPS3 (ref. 10). In the mouse, more than 15 loci manifest mutant phenotypes similar to human HPS11,12, including pale ear (ep), the mouse homolog of HPS1 (refs 13,14). Mouse ep has a phenotype identical to another mutant, light ear (le)15,16,17,18, which suggests that the human homolog of le is a possible human HPS locus. We have identified and found mutations of the human le homolog, HPS4, in a number of non–Puerto Rican individuals with HPS, establishing HPS4 as an important HPS locus in humans. In addition to their identical phenotypes, le and ep mutant mice have identical abnormalities of melanosomes, and in transfected melanoma cells the HPS4 and HPS1 proteins partially co-localize in vesicles of the cell body. In addition, the HPS1 protein is absent in tissues of le mutant mice. These results suggest that the HPS4 and HPS1 proteins may function in the same pathway of organelle biogenesis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Hermansky, F. & Pudlak, P. Albinism associated with hemorrhagic diathesis and unusual pigmented reticular cells in the bone marrow. Blood 14, 162–169 (1959).
Spritz, R.A. Multi-organellar disorders of pigmentation: tied up in traffic. Clin. Genet. 55, 309–317 (1999).
Spritz, R.A. Multi-organellar disorders of pigmentation: intracellular traffic jams in mammals, flies, and yeast. Trends Genet. 15, 337–340 (1999).
Spritz, R.A. Hermansky-Pudlak syndrome and pale ear: melanosome-making for the millennium. Pig. Cell. Res. 13, 15–20 (2000).
Witkop, C.J., Almadovar, C., Pineiro, B. & Babcock, M.N. Hermansky-Pudlak syndrome (HPS). An epidemiologic study. Ophth. Paediatr. Genet. 11, 245–250 (1990).
Witkop, C.J. et al. Albinism and Hermansky-Pudlak syndrome in Puerto Rico. Bol. Assoc. Med. P. R. 82, 333–339 (1990).
Oh, J. et al. Positional cloning of a gene for Hermansky-Pudlak syndrome, a disorder of cytoplasmic organelles. Nature Genet. 14, 300–306 (1996).
Anikster, Y. et al. Mutation of a new gene causes a unique form of Hermansky-Pudlak syndrome in a genetic isolate of central Puerto Rico. Nature Genet. 28, 376–380 (2001).
Oh, J. et al. Mutation analysis of patients with Hermansky-Pudlak syndrome defines a frameshift hotspot in the HPS gene as well as apparent locus heterogeneity. Am. J. Hum. Genet. 62, 593–598 (1998).
Suzuki, T. et al. The gene mutated in cocoa mice, carrying a defect of organelle biogenesis, is a homologue of the human Hermansky-Pudlak Syndrome-3 gene. Genomics 78, 1–8 (2001).
Bennett, D. Genetics, development, and malignancy of melanocytes. Int. Rev. Cytol. 146, 191–260 (1993).
Swank, R.T., Novak, E.K., McGarry, M.P., Rusiniak, M.E. & Feng, L. Mouse models of Hermansky-Pudlak syndrome: a review. Pig. Cell. Res. 11, 60–80 (1998).
Feng, G.H., Bailin, T., Oh, J. & Spritz, R.A. Mouse pale ear (ep) is homologous to human Hermansky-Pudlak syndrome and contains a rare “AT-AC” intron. Hum. Mol. Genet. 6, 793–797 (1997).
Gardner, J.M. et al. The mouse pale ear (ep) mutation is the homologue of human Hermansky Pudlak syndrome (HPS). Proc. Natl Acad. Sci. USA 94, 9238–9243 (1997).
Lane, P.W. Linkage groups 3 and XVII in the mouse and the position of the light-ear locus. J. Hered. 58, 21–24 (1967).
Lane, P.W. & Green, E.L. Pale ear and light ear in the house mouse: mimic mutations in linkage groups XII and XVII. J. Hered. 58, 17–20 (1967).
Meisler, M., Levy, J., Sansone, F. & Gordon, M. Morphologic and biochemical abnormalities of kidney lysosomes in mice with an inherited albinism. Am. J. Pathol. 101, 581–594 (1980).
Novak, E.K., Hui, S.-W. & Swank, R.T. Platelet storage pool deficiency in mouse pigment mutations associated with seven distinct genetic loci. Blood 63, 536–544 (1984).
Meisler, M.H., Wanner, L. & Strahler, J. Pigmentation and lysosomal phenotypes in mice doubly homozygous for both light-ear and pale-ear mutant alleles. J. Hered. 75, 103–106 (1984).
Chapman, V.M., Noell, W.K. & Adler, D. Alb-1. Mouse News Lett. 53, 61 (1975).
Buckler, A.J. et al. Exon amplification: a strategy to isolate mammalian genes based on RNA splicing. Proc. Natl Acad. Sci. USA 88, 4005–4009 (1991).
Altschul, S.F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402 (1997).
Dunham, I. et al. The DNA sequence of human chromosome 22. Nature 402, 489–495 (1999).
Lee, S.-T., Park, S.-K., Lee, K.-H., Holmes, S.A. & Spritz, R.A. A non-radioactive method for simultaneous detection of single-strand conformation polymorphisms (SSCPs) and heteroduplexes. Molecules Cells 5, 668–672 (1995).
Spritz, R.A. Chediak-Higashi syndrome. in Primary Immunodeficiency Diseases (eds Ochs, H.D., Smith, C.I.E. & Puck, J.M.) 389–396 (Oxford University Press, New York, 1999).
Feng, L. et al. The β3A subunit gene (Ap3b1) of the AP-3 adaptor complex is altered in the mouse hypopigmentation mutant pearl, a model for Hermansky-Pudlak syndrome and night blindness. Hum. Mol. Genet. 8, 323–330 (1999).
Dell'Angelica, E.C., Shotelersuk, V., Aguilar, R.C., Gahl, W.A. & Bonifacino, J.S. Altered trafficking of lysosomal proteins in Hermansky-Pudlak syndrome due to mutations in the b3A subunit of the AP-3 adaptor. Mol. Cell 3, 1–20 (1999).
Oh, J., Liu, Z.-X., Feng, G.H., Raposo, G. & Spritz, R.A. The Hermansky-Pudlak syndrome (HPS) protein is part of a high molecular weight complex involved in biogenesis of early melanosomes. Hum. Mol. Genet. 9, 375–385 (2000).
Cuomo, M. et al. Production and characterization of the murine monoclonal antibody 2G10 to a human T4-tyrosinase epitope. J. Invest. Dermatol. 96, 446–451 (1991).
Robinson, M.S. & Bonifacino, J.S. Adaptor-related proteins. Curr. Opin. Cell Biol. 13, 444–453 (2001).
Acknowledgements
This work was supported by grants from the National Insitutes of Health (to R.A.S, R.T.S. and the Roswell Park Cancer Institute) and the Wellcome Trust (to E.V.S.). We thank A. Wilson, Y. Jiang, L. Zhen, D. Tabaczynski, D. Poslinski and M.K. Ellsworth for technical assistance.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Suzuki, T., Li, W., Zhang, Q. et al. Hermansky-Pudlak syndrome is caused by mutations in HPS4, the human homolog of the mouse light-ear gene. Nat Genet 30, 321–324 (2002). https://doi.org/10.1038/ng835
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng835
This article is cited by
-
Novel HPS6 mutations identified by whole-exome sequencing in two Japanese sisters with suspected ocular albinism
Journal of Human Genetics (2016)
-
Association of the Hermansky–Pudlak syndrome type 4 (HPS4) gene variants with cognitive function in patients with schizophrenia and healthy subjects
BMC Psychiatry (2013)
-
The construction of transgenic and gene knockout/knockin mouse models of human disease
Transgenic Research (2012)
-
Homozygosity Mapping and Whole-Exome Sequencing to Detect SLC45A2 and G6PC3 Mutations in a Single Patient with Oculocutaneous Albinism and Neutropenia
Journal of Investigative Dermatology (2011)
-
Clinical, Molecular, and Cellular Features of Non-Puerto Rican Hermansky–Pudlak Syndrome Patients of Hispanic Descent
Journal of Investigative Dermatology (2011)