Abstract
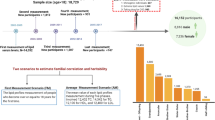
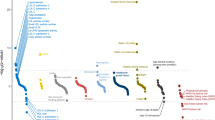
Coronary artery disease and myocardial infarction (MI) are leading causes of death in the western world. Numerous studies have shown that risk factors such as diabetes mellitus, arterial hypertension and hypercholesterolemia contribute to the development of the disease. Although each risk factor by itself is partly under genetic control, a positive family history is an independent predictor, which suggests that there are additional susceptibility genes1. We have scanned the whole genome in 513 families to identify chromosomal regions linked to myocardial infarction and related risk factors that are known to be under genetic control. Here we show, by using variance component analysis and incorporating risk factors, that risk of myocardial infarction maps to a single region on chromosome 14 with a significant lod score of 3.9 (pointwise P=0.00015, genome-wide P<0.05), providing evidence of a principal MI locus. To characterize this locus we analyzed each risk factor by itself. Serum concentrations of lipoprotein (a) show linkage to both the apolipoprotein (a) locus (lod score 26.99) and a new locus on chromosome 1 (lod score 3.8). There is suggestive linkage for diabetes mellitus on chromosome 6 (lod score 2.96), for hypertension on chromosomes 1 and 6, for high-density and low-density lipoprotein cholesterol on chromosomes 1 and 17, and for triglyceride concentrations on chromosome 9. Although some of these risk factors overlap with previously identified loci, none overlaps with the newly identified susceptibility locus for myocardial infarction and coronary artery disease.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Marenberg, M.E., Risch, N., Berkman, L., Floderus, B. & de Faire, U. Genetics susceptibility to death from coronary heart disease in a study of twins. N. Engl. J. Med. 330, 1041–1046 (1994).
Almasy, L. & Blangero, J. Multipoint quantitative trait linkage analysis in general pedigrees. Am. J. Hum. Genet. 62, 1198–1211 (1998).
Blangero, J., Williams, J.T. & Almasy, L. Variance component methods for detecting complex trait loci. Adv. Genet. 42, 151–181 (2001).
Schork, N.J. Extended multipoint identity-by-descent analysis of human quantitative traits: efficiency, power, and modeling considerations. Am. J. Hum. Genet. 53, 1306–1319 (1993).
Thompson, E.A. Sampling and ascertainment in genetic epidemiology: a tutorial review. Technical Report No. 243, Univ. Washington, Dept Statistics, Seattle (1993).
Comuzzie, A. & Williams, J. Correcting for ascertainment bias in the COGA data set. Genet. Epidemiol. 17, S109–S114 (1999).
Executive summary of the third report of the national cholesterol education program (NCEP) expert panel on detection, evaluation, and treatment of high blood cholesterol in adults (Adult Treatment Panel III). J. Am. Med. Assoc. 285, 2486–2497 (2001).
Feingold, E., Brown, P.O. & Siegmund, D. Gaussian models for genetic linkage analysis using complete high-resolution maps of identity by descent. Am. J. Hum. Genet. 53, 234–251 (1993).
Pajukanta, P. et al. Two loci on chromosomes 2 and X for premature coronary heart disease identified in early- and late-settlement populations of Finland. Am. J. Hum. Genet. 67, 1481–1493 (2000).
Dahlen, G.H. et al. The importance of serum lipoprotein (a) as an independent risk factor for premature coronary artery disease in middle-aged black and white women from the United States. J. Intern. Med. 244, 417–424 (1998).
Boerwinkle, E. et al. Apolipoprotein(a) gene accounts for greater than 90% of the variation in plasma lipoprotein(a) concentrations. J. Clin. Invest. 90, 52–60 (1992).
Pajukanta, P. et al. Linkage of familial combined hyperlipidaemia to chromosome 1q21–q23. Nature Genet. 18, 369–373 (1998).
Vionnet, N. et al. Genomewide search for type 2 diabetes-susceptibility genes in French whites: evidence for a novel susceptibility locus for early-onset diabetes on chromosome 3q27-qter and independent replication of a type 2-diabetes locus on chromosome 1q21–q24. Am. J. Hum. Genet. 67, 1470–1480 (2000).
Ghosh, S. et al. The Finland-United States investigation of non-insulin-dependent diabetes mellitus genetics (FUSION) study. I. An autosomal genome scan for genes that predispose to type 2 diabetes. Am. J. Hum. Genet. 67, 1174–1185 (2000).
Mansfield, T.A. et al. Multilocus linkage of familial hyperkaliaemia and hypertension, pseudohypoaldosteronism type II, to chromosomes 1q31–42 and 17p11–q21. Nature Genet. 16, 202–205 (1997).
Stoll, M. et al. New target regions for human hypertension via comparative genomics. Genome Res. 10, 473–482 (2000).
Horikawa, Y. et al. Genetic variation in the gene encoding calpain-10 is associated with type 2 diabetes mellitus. Nature Genet. 26, 163–175 (2000).
Hugot, J.-P. et al. Association of NOD2 leucine-rich repeat variants with susceptibility to Crohn's disease. Nature 411, 599–603 (2001).
Ogura, Y. et al. A frameshift mutation in NOD2 associated with susceptibility to Crohn's disease. Nature 411, 603–606 (2001).
Lobel, H., et al. Coronary heart disease case fatality in four countries. A community study. The actual myocardial infarction register teams of Auckland. Circulation 88, 2524–2531 (1993).
Weber, J.L. & Broman, K.W. Genotyping for human whole genome scans: past, present and future. Adv. Genet. 42, 77–96 (2001).
Lee, Y.A., et al. A major susceptibility locus for atopic dermatitis maps to chromosome 3q21. Nature Genet. 26, 470–473 (2000).
Allison, D.B. et al. Testing the robustness of the likelihood-ratio test in a variance- component quantitative-trait loci-mapping procedure. Am. J. Hum. Genet. 65, 531–544 (1999).
Goring, H.H. & Ott, J. Relationship estimation in affected sib pair analysis of late-onset diseases. Eur. J. Hum. Genet. 5, 69–77 (1997).
Amos, C.I. Robust variance-components approach for assessing genetic linkage in pedigrees. Am. J. Hum. Genet. 54, 535–543 (1994).
Fulker, D., Cherny, S., Sham, P. & Hewitt, J. Combined linkage and association sib-pair analysis for quantitative traits. Am. J. Hum. Genet. 64, 259–267 (1999).
Sobel, E. & Lange, K. Descent graphs in pedigree analysis: applications to haplotyping, location scores, and marker-sharing statistics. Am. J. Hum. Genet. 58, 1323–1337 (1996).
Acknowledgements
We thank G. Klein for helping to establish the MI family database; members of these families for cooperation and consent; M. Pöll and S. Kürzinger for technical assistance; U. Hubauer for managing the phenotype evaluation; H. Xia for database assistance; J. Williams and L. Almasy for statistical advice; and J. Weber and the NHLBI Mammalian Genotyping Service for valuable contributions. U.B. and H.J.J. are supported in part by a grant from the National Heart, Lung, and Blood Institute. A.C. and J.B. are supported in part by a grant from the National Institutes of Health. We acknowledge the support from the Wilhelm-Vaillant-Stiftung, the Ernst-und-Berta-Grimmke-Stiftung, the Deutsche Herzstiftung and the Deutsche Forschungsgemeinschaft (to U.B., S.H., C.H. and H.S.).
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Broeckel, U., Hengstenberg, C., Mayer, B. et al. A comprehensive linkage analysis for myocardial infarction and its related risk factors. Nat Genet 30, 210–214 (2002). https://doi.org/10.1038/ng827
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng827
This article is cited by
-
Should We Use Genetic Scores in the Determination of Treatment Strategies to Control Dyslipidemias?
Current Cardiology Reports (2020)
-
First-degree relatives with similar phenotypic characterisation of acute myocardial infarction: a case report and review of the literature
BMC Cardiovascular Disorders (2019)
-
Genome-Wide Linkage Analysis of Large Multiple Multigenerational Families Identifies Novel Genetic Loci for Coronary Artery Disease
Scientific Reports (2017)
-
Implications of ACE (I/D) Gene Variants to the Genetic Susceptibility of Coronary Artery Disease in Asian Indians
Indian Journal of Clinical Biochemistry (2017)
-
Quantitative assessment of the influence of PSMA6 variant (rs1048990) on coronary artery disease risk
Molecular Biology Reports (2013)