Abstract
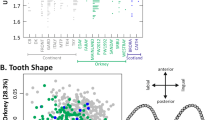
The genome of the laboratory mouse is thought to be a mosaic of regions with distinct subspecific origins. We have developed a high-resolution map of the origin of the laboratory mouse by generating 25,400 phylogenetic trees at 100-kb intervals spanning the genome. On average, 92% of the genome is of Mus musculus domesticus origin, and the distribution of diversity is markedly nonrandom among the chromosomes. There are large regions of extremely low diversity, which represent blind spots for studies of natural variation and complex traits, and hot spots of diversity. In contrast with the mosaic model, we found that most of the genome has intermediate levels of variation of intrasubspecific origin. Finally, mouse strains derived from the wild that are supposed to represent different mouse subspecies show substantial intersubspecific introgression, which has strong implications for evolutionary studies that assume these are pure representatives of a given subspecies.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Paigen, K. One hundred years of mouse genetics: an intellectual history. I. The classical period (1902–1980). Genetics 163, 1–7 (2003).
Paigen, K. One hundred years of mouse genetics: an intellectual history. II. The molecular revolution (1981–2002). Genetics 163, 1227–1235 (2003).
Beck, J.A. et al. Genealogies of mouse inbred strains. Nat. Genet. 24, 23–25 (2000).
Ferris, S.D., Sage, R.D. & Wilson, A.C. Evidence from mtDNA sequences that common laboratory strains of inbred mice are descended from a single female. Nature 295, 163–165 (1982).
Bishop, C.E., Boursot, P., Baron, B., Bonhomme, F. & Hatat, D. Most classical Mus musculus domesticus laboratory mouse strains carry a Mus musculus musculus Y chromosome. Nature 315, 70–72 (1985).
Nagamine, C.M. et al. The musculus-type Y chromosome of the laboratory mouse is of Asian origin. Mamm. Genome 3, 84–91 (1992).
Bonhomme, F., Guenet, J.-L., Dod, B., Moriwaki, K. & Bulfield, G. The polyphyletic of the laboratory inbred mice and their rate of evolution. J. Linn. Soc. 30, 51–58 (1987).
Wade, C.M. et al. The mosaic structure of variation in the laboratory mouse genome. Nature 420, 574–578 (2002).
Wiltshire, T. et al. Genome-wide single-nucleotide polymorphism analysis defines haplotype patterns in mouse. Proc. Natl. Acad. Sci. USA 100, 3380–3385 (2003).
Pletcher, M.T. et al. Use of a dense single nucleotide polymorphism map for in silico mapping in the mouse. PLoS Biol. [online] 2, e393 (2004) (doi:10.1371/journal.pbio.0020393).
Frazer, K.A. et al. Segmental phylogenetic relationships of inbred mouse strains revealed by fine-scale analysis of sequence variation across 4.6 Mb of mouse genome. Genome Res. 14, 1493–1500 (2004).
Petkov, P.M. et al. Evidence of a large-scale functional organization of mammalian chromosomes. PLoS Genet. [online] 1, e33 (2005) (doi:10.1371/journal.pgen.0010033).
Yalcin, B. et al. Unexpected complexity in the haplotypes of commonly used inbred strains of laboratory mice. Proc. Natl. Acad. Sci. USA 101, 9734–9739 (2004).
Harr, B. Genomic islands of differentiation between house mouse subspecies. Genome Res. 16, 730–737 (2006).
Harr, B. Regions of high differentiation–worth a check. Genome Res. 16, 1193–1194 (2006).
Boursot, P. & Belkhir, K. Mouse SNPs for evolutionary biology: beware of ascertainment biases. Genome Res. 16, 1191–1192 (2006).
Zhang, J. et al. A high-resolution multistrain haplotype analysis of laboratory mouse genome reveals three distinctive genetic variation patterns. Genome Res. 15, 241–249 (2005).
Ideraabdullah, F.Y. et al. Genetic and haplotype diversity among wild derived mouse inbred strains. Genome Res. 14, 1880–1887 (2004).
Boissinot, S. & Boursot, P. Discordant phylogeographic patterns between the Y chromosome and mitochondrial DNA in the house mouse: selection on the Y chromosome? Genetics 146, 1019–1034 (1997).
Bonhomme, F. et al. Species-wide distribution of highly polymorphic minisatellite markers suggests past and present genetic exchanges among House Mouse subspecies. Genome Biol. [online] 8, R80 (2007) (doi:10.1186/gb-2007-8-5-r80).
Frazer, K.A. et al. A sequence-based variation map of 8.27 million SNPs in inbred mouse strains. Nature, advance online publication 29 July 2007 (doi:10.1038/nature06067).
Genetic Variants and Strains of the Laboratory Mouse (Lyon, M.F., Rastan, S. & Brown, S.D.M., eds.) 3rd edn. (Oxford University Press, Oxford 1996).
Yonekawa, H. et al. Hybrid origin of Japanese mice “Mus musculus molossinus”: evidence from restriction analysis of mitochondrial DNA. Mol. Biol. Evol. 5, 63–78 (1988).
Sakai, T. et al. Origins of mouse inbred strains deduced from whole-genome scanning by polymorphic microsatellite loci. Mamm. Genome 16, 11–19 (2005).
Abe, K. et al. Contribution of Asian mouse subspecies Mus musculus molossinus to genomic constitution of strain C57BL/6J, as defined by BAC-end sequence-SNP analysis. Genome Res. 14, 2439–2447 (2004).
Wade, C.M. & Daly, M.J. Genetic variation in laboratory mice. Nat. Genet. 37, 1175–1180 (2005).
Auffray, J.-C., Vanlerberghe, F. & Britton-Davidian, J. The house mouse progression in Eurasia: a palaeontological and archaeozoological approach. Biol. J. Linn. Soc. 41, 13–25 (1990).
Din, W. et al. Origin and radiation of the house mouse: clues from nuclear genes. J. Evol. Biol. 9, 519–539 (1996).
Prager, E.M., Orrego, C. & Sage, R.D. Genetic variation and phylogeography of central Asian and other house mice, including a major new mitochondrial lineage in Yemen. Genetics 150, 835–861 (1998).
Patil, N. et al. Blocks of limited haplotype diversity revealed by high-resolution scanning of human chromosome 21. Science 294, 1719–1723 (2001).
Keightley, P.D., Lercher, M.J. & Eyre-Walker, A. Evidence for widespread degradation of gene control regions in hominid genomes. PLoS Biol. 3, 282–288 (2005).
Forejt, J. Hybrid sterility in the mouse. Trends Genet. 12, 412–417 (1996).
Thrachtulec, Z. et al. Positional cloning of the hybrid sterility 1 gene: fine genetic mapping and evaluation of two candidate genes. Biol. J. Linn. Soc. 84, 637–641 (2005).
Payseur, B.A. & Hoekstra, H.E. Signatures of reproductive isolation in patterns of single nucleotide diversity across inbred strains of mice. Genetics 171, 1905–1916 (2005).
Oka, A. et al. Disruption of genetic interaction between two autosomal regions and the x chromosome causes reproductive isolation between mouse strains derived from different subspecies. Genetics 175, 185–197 (2007).
Dai, J. et al. The absence of mitochondrial DNA diversity among common laboratory inbred mouse strains. J. Exp. Biol. 208, 4445–4450 (2005).
Tucker, P.K., Lee, B.K., Lundrigan, B.L. & Eicher, E.M. Geographic origin of the Y chromosomes in “old” inbred strains of mice. Mamm. Genome 3, 254–261 (1992).
Ferris, S.D. et al. Flow of mitochondrial DNA across a species boundary. Proc. Natl. Acad. Sci. USA 80, 2290–2294 (1983).
Li, J. et al. Genomic segmental polymorphisms in inbred mouse strains. Nat. Genet. 36, 952–954 (2004).
Snijders, A.M. et al. Mapping segmental and sequence variations among laboratory mice using BAC array CGH. Genome Res. 15, 302–311 (2005).
Graubert, T.A. et al. A high-resolution map of segmental DNA copy number variation in the mouse genome. PLoS Genet. [online] 3, e3 (2007) (doi:10.1371/journal.pgen.0030003).
Valdar, W. et al. Genome-wide genetic association of complex traits in heterogeneous stock mice. Nat. Genet. 38, 879–887 (2006).
Churchill, G.A. et al. The Collaborative Cross, a community resource for the genetic analysis of complex traits. Nat. Genet. 36, 1133–1137 (2004).
Churchill, G.A. Stochastic models for heterogeneous DNA sequences. Bull. Math. Biol. 51, 79–94 (1989).
Acknowledgements
We thank S. Ahmed for technical assistance; K. Paigen, K. Broman and B. Payseur for helpful comments during the preparation of the manuscript; J. Felsenstein for advice and L. Wu for assistance with the phylogenetic tree computations and A. Smith for developing a genome browser format for displaying phylogenetic trees. CIM/Pas was provided by F. Bonhomme (University Mont Pellier II). This work was supported by the US National Institute of General Medical Sciences as part of the Center of Excellence in Systems Biology (1P50 GM076468).
Author information
Authors and Affiliations
Contributions
This study was designed by F.P.-M.V. and G.A.C. The genome-wide analyses were carried out by H.Y. The sequence data used to determine the false-negative and false-positive rates and to confirm the presence and direction of intrasubspecific introgression were generated by T.A.B.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6, Supplementary Note, Supplementary Table 1 (PDF 3338 kb)
Rights and permissions
About this article
Cite this article
Yang, H., Bell, T., Churchill, G. et al. On the subspecific origin of the laboratory mouse. Nat Genet 39, 1100–1107 (2007). https://doi.org/10.1038/ng2087
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng2087
This article is cited by
-
Multi-omics analysis identifies drivers of protein phosphorylation
Genome Biology (2023)
-
Population structure and inbreeding in wild house mice (Mus musculus) at different geographic scales
Heredity (2022)
-
A new mouse SNP genotyping assay for speed congenics: combining flexibility, affordability, and power
BMC Genomics (2021)
-
Identification of novel loci controlling inflammatory bowel disease susceptibility utilizing the genetic diversity of wild-derived mice
Genes & Immunity (2020)
-
Wild-derived mice: from genetic diversity to variation in immune responses
Mammalian Genome (2018)