Abstract
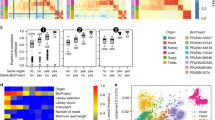
A major challenge in comparative genomics is to understand how phenotypic differences between species are encoded in their genomes. Phenotypic divergence may result from differential transcription of orthologous genes, yet less is known about the involvement of differential translation regulation in species phenotypic divergence. In order to assess translation effects on divergence, we analyzed ∼2,800 orthologous genes in nine yeast genomes. For each gene in each species, we predicted translation efficiency, using a measure of the adaptation of its codons to the organism's tRNA pool. Mining this data set, we found hundreds of genes and gene modules with correlated patterns of translational efficiency across the species. One signal encompassed entire modules that are either needed for oxidative respiration or fermentation and are efficiently translated in aerobic or anaerobic species, respectively. In addition, the efficiency of translation of the mRNA splicing machinery strongly correlates with the number of introns in the various genomes. Altogether, we found extensive selection on synonymous codon usage that modulates translation according to gene function and organism phenotype. We conclude that, like factors such as transcription regulation, translation efficiency affects and is affected by the process of species divergence.
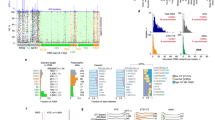
NOTE: In the original version of this paper, subpanels 2c and d were switched, resulting in incorrect information in the legend. Figure 2c shows genes of the tricarboxylic acid (TCA) cycle, while Figure 2d shows glycolysis genes. The error has been corrected in the HTML and PDF versions of the article.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
28 March 2007
NOTE: In the original version of this paper, subpanels 2c and d were switched, resulting in incorrect information in the legend. Figure 2c shows genes of the tricarboxylic acid (TCA) cycle, while Figure 2d shows glycolysis genes. The error has been corrected in the HTML and PDF versions of the article.
References
Wolfe, K.H. Comparative genomics and genome evolution in yeasts. Philos. Trans. R. Soc. Lond. B Biol. Sci. 361, 403–412 (2006).
Gompel, N., Prud'homme, B., Wittkopp, P.J., Kassner, V.A. & Carroll, S.B. Chance caught on the wing: cis–regulatory evolution and the origin of pigment patterns in Drosophila. Nature 433, 481–487 (2005).
Ihmels, J. et al. Rewiring of the yeast transcriptional network through the evolution of motif usage. Science 309, 938–940 (2005).
Powers, D.A. & Schulte, P.M. Evolutionary adaptations of gene structure and expression in natural populations in relation to a changing environment: a multidisciplinary approach to address the million-year saga of a small fish. J. Exp. Zool. 282, 71–94 (1998).
dos Reis, M., Savva, R. & Wernisch, L. Solving the riddle of codon usage preferences: a test for translational selection. Nucleic Acids Res. 32, 5036–5044 (2004).
Sharp, P.M. & Li, W.H. The codon adaptation index–a measure of directional synonymous codon usage bias, and its potential applications. Nucleic Acids Res. 15, 1281–1295 (1987).
Percudani, R., Pavesi, A. & Ottonello, S. Transfer RNA gene redundancy and translational selection in Saccharomyces cerevisiae. J. Mol. Biol. 268, 322–330 (1997).
Lowe, T.M. & Eddy, S.R. tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res. 25, 955–964 (1997).
Tanay, A., Regev, A. & Shamir, R. Conservation and evolvability in regulatory networks: the evolution of ribosomal regulation in yeast. Proc. Natl. Acad. Sci. USA 102, 7203–7208 (2005).
Ghaemmaghami, S. et al. Global analysis of protein expression in yeast. Nature 425, 737–741 (2003).
Fraser, H.B., Hirsh, A.E., Wall, D.P. & Eisen, M.B. Coevolution of gene expression among interacting proteins. Proc. Natl. Acad. Sci. USA 101, 9033–9038 (2004).
Lithwick, G. & Margalit, H. Relative predicted protein levels of functionally associated proteins are conserved across organisms. Nucleic Acids Res. 33, 1051–1057 (2005).
Rice, J. Mathematical Statistics and Data Analysis (Wadsworth Publishing Company, Belmont, Calfiornia, 1995).
Harris, M.A. et al. The Gene Ontology (GO) database and informatics resource. Nucleic Acids Res. 32, D258–D261 (2004).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate – a practical and powerful approach to multiple testing. J. R. S. Soc. Ser. B. Methodol. 57, 289–300 (1995).
Nantel, A. The long hard road to a completed Candida albicans genome. Fungal Genet. Biol. 43, 311–315 (2006).
Barth, G. & Gaillardin, C. Physiology and genetics of the dimorphic fungus Yarrowia lipolytica. FEMS Microbiol Rev 19, 219–237 (1997).
Braun, B.R. et al. A human-curated annotation of the Candida albicans genome. PLoS Genet 1, 36–57 (2005).
Kellis, M., Birren, B.W. & Lander, E.S. Proof and evolutionary analysis of ancient genome duplication in the yeast Saccharomyces cerevisiae. Nature 428, 617–624 (2004).
Berbee, M. & Taylor, J. Systematics and evolution. in The Mycota, Vol. VIIB (eds. McLaughlin, D., McLaughlin, E. & Lemke, P.) 229–245 (Springer, Berlin, 2001).
Arnaud, M.B. et al. Sequence resources at the Candida Genome Database. Nucleic Acids Res. 35, D452–D456 (2007).
Issel-Tarver, L. et al. Saccharomyces Genome Database. Methods Enzymol 350, 329–346 (2002).
Galagan, J.E. et al. Sequencing of Aspergillus nidulans and comparative analysis with A. fumigatus and A. oryzae. fumigatus and A. oryzae. Nature 438, 1105–1115 (2005).
Pruess, M., Kersey, P. & Apweiler, R. The Integr8 project–a resource for genomic and proteomic data. In Silico Biol. 5, 179–185 (2005).
Kanz, C. et al. The EMBL Nucleotide Sequence Database. Nucleic Acids Res. 33, D29–D33 (2005).
Segal, E. et al. A genomic code for nucleosome positioning. Nature 442, 772–778 (2006).
Wright, F. The 'effective number of codons' used in a gene. Gene 87, 23–29 (1990).
Alexeyenko, A., Tamas, I., Liu, G. & Sonnhammer, E.L. Automatic clustering of orthologs and inparalogs shared by multiple proteomes. Bioinformatics 22, e9–e15 (2006).
Prillinger, H. et al. Phylogeny and systematics of the fungi with special reference to the Ascomycota and Basidiomycota. Chem. Immunol. 81, 207–295 (2002).
Acknowledgements
We thank the Pilpel laboratory for helpful discussions and E. Segal, A. Regev, I. Tirosh, Y. Gilad, N. Barkai, J. Moult and J.L. Sussman for discussions and critical reviews of the manuscript. Y.P. holds the Rothstein Career Development Chair in Genetic Diseases. We thank the Ben May Charitable Trust and EMBRACE, the European Union Network of Excellence in Bioinformatics for grant support.
Author information
Authors and Affiliations
Contributions
Y.P. and O.M. conceived the study, and Y.P. supervised the study. O.M. and Y.P. designed the analyses, O.M. performed the analyses and O.M. and Y.P. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
The relationship between translation efficiency (tAI) and protein levels. (PDF 224 kb)
Supplementary Fig. 2
Physically interacting proteins tend to have similar translation efficiencies across species. (PDF 90 kb)
Supplementary Fig. 3
Clustering of the multispecies translation efficiency profiles. (PDF 407 kb)
Supplementary Fig. 4
Comparison of the effective number of codons (Nc) and the tRNA adaptation index (tAI) for the coding sequences of ten ascomycotic yeast species. (PDF 223 kb)
Supplementary Fig. 5
Translation efficiency profiles of genes related to M-phase of the cell cycle. (PDF 92 kb)
Supplementary Table 1
The tRNA repertoires of the analyzed species. (PDF 76 kb)
Supplementary Table 2
Translation efficiency profiles of orthologous genes across species, their division into 40 clusters, and the functional enrichment within clusters. (PDF 562 kb)
Rights and permissions
About this article
Cite this article
Man, O., Pilpel, Y. Differential translation efficiency of orthologous genes is involved in phenotypic divergence of yeast species. Nat Genet 39, 415–421 (2007). https://doi.org/10.1038/ng1967
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng1967
This article is cited by
-
Multiple intermolecular interactions facilitate rapid evolution of essential genes
Nature Ecology & Evolution (2023)
-
Codon usage bias and environmental adaptation in microbial organisms
Molecular Genetics and Genomics (2021)
-
The evolutionary signal in metagenome phyletic profiles predicts many gene functions
Microbiome (2018)
-
Codon usage and amino acid usage influence genes expression level
Genetica (2018)
-
Complete mitochondrial genome of Gracilaria changii (Rhodophyta: Gracilariaceae)
Journal of Applied Phycology (2017)