Abstract
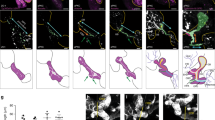
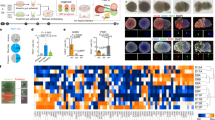
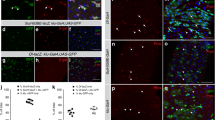
During organogenesis, the foregut endoderm gives rise to the many different cell types that comprise the hepatopancreatic system, including hepatic, pancreatic and gallbladder cells, as well as the epithelial cells of the hepatopancreatic ductal system that connects these organs together and with the intestine. However, the mechanisms responsible for demarcating ducts versus organs are poorly understood. Here, we show that Fgf10 signaling from the adjacent mesenchyme is responsible for refining the boundaries between the hepatopancreatic duct and organs. In zebrafish fgf10 mutants, the hepatopancreatic ductal epithelium is severely dysmorphic, and cells of the hepatopancreatic ductal system and adjacent intestine misdifferentiate toward hepatic and pancreatic fates. Furthermore, Fgf10 also functions to prevent the differentiation of the proximal pancreas and liver into hepatic and pancreatic cells, respectively. These data shed light onto how the multipotent cells of the foregut endoderm, and subsequently those of the hepatopancreatic duct, are directed toward different organ fates.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Sosa-Pineda, B., Wigle, J.T. & Oliver, G. Hepatocyte migration during liver development requires Prox1. Nat. Genet. 25, 254–255 (2000).
Wang, J. et al. Prox1 activity controls pancreas morphogenesis and participates in the production of “secondary transition” pancreatic endocrine cells. Dev. Biol. 286, 182–194 (2005).
Li, J., Ning, G. & Duncan, S.A. Mammalian hepatocyte differentiation requires the transcription factor HNF-4. Genes Dev. 14, 464–474 (2000).
Jacquemin, P. et al. An endothelial–mesenchymal relay pathway regulates early phases of pancreas development. Dev. Biol. 290, 189–199 (2006).
Norton, W.H.J., Ledin, J., Grandel, H. & Neumann, C.J. HSPG synthesis by zebrafish Ext2 and Extl3 is required for Fgf10 signalling during limb development. Development 132, 4963–4973 (2005).
Ahlgren, U., Pfaff, S.L., Jessell, T.M., Edlund, T. & Edlund, H. Independent requirement for ISL1 in formation of pancreatic mesenchyme and islet cells. Nature 385, 257–260 (1997).
Lu, W., Luo, Y., Kan, M. & McKeehan, W.L. Fibroblast Growth Factor-10. A second candidate stromal to epithelial cell andromedin in prostate. J. Biol. Chem. 274, 12827–12834 (1999).
Pulkkinen, M.A., Spencer-Dene, B., Dickson, C. & Otonkoski, T. The IIIb isoform of fibroblast growth factor receptor 2 is required for proper growth and branching of pancreatic ductal epithelium but not for differentiation of exocrine or endocrine cells. Mech. Dev. 120, 167–175 (2003).
Tonou-Fujimori, N. et al. Expression of the FGF receptor 2 gene (fgfr2) during embryogenesis in the zebrafish Danio rerio. Mech. Dev. 119 (Suppl.), S173–S178 (2002).
Crosnier, C. et al. Delta-Notch signalling controls commitment to a secretory fate in the zebrafish intestine. Development 132, 1093–1104 (2005).
Bhushan, A. et al. Fgf10 is essential for maintaining the proliferative capacity of epithelial progenitor cells during early pancreatic organogenesis. Development 128, 5109–5117 (2001).
Wan, H. et al. Analyses of pancreas development by generation of gfp transgenic zebrafish using an exocrine pancreas-specific elastaseA gene promoter. Exp. Cell Res. 312, 1526–1539 (2006).
Ng, A.N.Y. et al. Formation of the digestive system in zebrafish: III. Intestinal morphogenesis. Dev. Biol. 286, 114–135 (2005).
Sussel, L. et al. Mice lacking the homeodomain transcription factor Nkx2.2 have diabetes due to arrested differentiation of pancreatic β cells. Development 125, 2213–2221 (1998).
Naya, F.J. et al. Diabetes, defective pancreatic morphogenesis and abnormal enteroendocrine differentiation in BETA2/NeuroD-deficient mice. Genes Dev. 11, 2323–2334 (1997).
Hart, A., Papadopoulou, S. & Edlund, H. Fgf10 maintains Notch activation, stimulates proliferation, and blocks differentiation of pancreatic epithelial cells. Dev. Dyn. 228, 185–193 (2003).
Norgaard, G.A., Jensen, J.N. & Jensen, J. FGF10 signaling maintains the pancreatic progenitor cell state revealing a novel role of Notch in organ development. Dev. Biol. 264, 323–338 (2003).
Her, G.M., Yeh, Y.H. & Wu, J.L. 435-bp liver regulatory sequence in the liver fatty acid binding protein (L-FABP) gene is sufficient to modulate liver regional expression in transgenic zebrafish. Dev. Dyn. 227, 347–356 (2003).
Kawaguchi, Y. et al. The role of the transcriptional regulator Ptf1a in converting intestinal to pancreatic progenitors. Nat. Genet. 32, 128–134 (2002).
Field, H.A., Ober, E.A., Roeser, T. & Stainier, D.Y.R. Formation of the digestive system in zebrafish. I. Liver morphogenesis. Dev. Biol. 253, 279–290 (2003).
Alexander, J., Stainier, D.Y.R. & Yelon, D. Screening mosaic F1 females for mutations affecting zebrafish heart induction and patterning. Dev. Genet. 22, 288–299 (1998).
Acknowledgements
We thank A. Ayala, S. Waldron and N. Zvenigorodsky for help with the fish; C. Wright, M. Hebrok, M. German, S. Curado, E. Ober, T. Sakaguchi, M. Bagnat and A. Schlegel for discussions and/or critical reading of the manuscript; C. Wright and J. Lewis for antibodies and B. Appel (Vanderbilt University) for the Tg(nkx2.2a:mEGFP) line. P.D.S.D. was supported by the Juvenile Diabetes Research Foundation (32002643) and the Larry L. Hillblom Foundation (2005 1G). C.A.M. was supported by a US National Science Foundation predoctoral fellowship. This work was supported in part by grants from the US National Institutes of Health (National Institute of Diabetes and Digestive and Kidney Diseases), the Juvenile Diabetes Research Foundation and the Packard Foundation (D.Y.R.S.). Requests for materials should be addressed to D.Y.R.S. (didier_stainier@biochem.ucsf.edu).
Author information
Authors and Affiliations
Contributions
P.D.S.D. made the original observation, generated hypotheses, designed experiments, analyzed fgf10 mutants and made the figures; C.A.M. analyzed fgf10 and neuroD expression, performed phenotype assessment and drew the illustration; W.N. and C.J.M. provided fgf10 mutants prior to publication; C.C. provided 2F11 antibodies prior to publication; X.P. and Z.G. provided Tg(lfabp:dsRed) zebrafish prior to publication; D.Y.R.S. oversaw the studies and P.D.S.D., C.A.M. and D.Y.R.S. wrote the manuscript with feedback from the other authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Differential expression of Prox1 and Hfn4α between the hepatopancreatic ducts and organs. (PDF 214 kb)
Supplementary Fig. 2
fgfr2 expression in the endoderm. (PDF 700 kb)
Supplementary Fig. 3
fgf10 mutants lacking the extrapancreatic duct and extrahepatic duct. (PDF 767 kb)
Supplementary Fig. 4
Pdx1 expression in the extrapancreatic duct is not maintained in fgf10 mutants. (PDF 684 kb)
Rights and permissions
About this article
Cite this article
Dong, P., Munson, C., Norton, W. et al. Fgf10 regulates hepatopancreatic ductal system patterning and differentiation. Nat Genet 39, 397–402 (2007). https://doi.org/10.1038/ng1961
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng1961
This article is cited by
-
All routes lead to Rome: multifaceted origin of hepatocytes during liver regeneration
Cell Regeneration (2021)
-
Transgenic fluorescent zebrafish lines that have revolutionized biomedical research
Laboratory Animal Research (2021)
-
AEO-7 surfactant is “super toxic” and induces severe cardiac, liver and locomotion damage in zebrafish embryos
Environmental Sciences Europe (2020)
-
In vivo generation and regeneration of β cells in zebrafish
Cell Regeneration (2020)
-
A morphogenetic EphB/EphrinB code controls hepatopancreatic duct formation
Nature Communications (2019)