Abstract
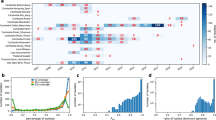
Genetic variation allows the malaria parasite Plasmodium falciparum to overcome chemotherapeutic agents, vaccines and vector control strategies and remain a leading cause of global morbidity and mortality1. Here we describe an initial survey of genetic variation across the P. falciparum genome. We performed extensive sequencing of 16 geographically diverse parasites and identified 46,937 SNPs, demonstrating rich diversity among P. falciparum parasites (π = 1.16 × 10−3) and strong correlation with gene function. We identified multiple regions with signatures of selective sweeps in drug-resistant parasites, including a previously unidentified 160-kb region with extremely low polymorphism in pyrimethamine-resistant parasites. We further characterized 54 worldwide isolates by genotyping SNPs across 20 genomic regions. These data begin to define population structure among African, Asian and American groups and illustrate the degree of linkage disequilibrium, which extends over relatively short distances in African parasites but over longer distances in Asian parasites. We provide an initial map of genetic diversity in P. falciparum and demonstrate its potential utility in identifying genes subject to recent natural selection and in understanding the population genetics of this parasite.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
World Health Organization. WHO expert committee on malaria. World Health Organ. Tech. Rep. Ser. 892, 1–74 (2000).
Gardner, M.J. et al. Genome sequence of the human malaria parasite Plasmodium falciparum. Nature 419, 498–511 (2002).
Nair, S. et al. A selective sweep driven by pyrimethamine treatment in southeast asian malaria parasites. Mol. Biol. Evol. 20, 1526–1536 (2003).
Wootton, J.C. et al. Genetic diversity and chloroquine selective sweeps in Plasmodium falciparum. Nature 418, 320–323 (2002).
Polley, S.D., Chokejindachai, W. & Conway, D.J. Allele frequency-based analyses robustly map sequence sites under balancing selection in a malaria vaccine candidate antigen. Genetics 165, 555–561 (2003).
Baum, J., Thomas, A.W. & Conway, D.J. Evidence for diversifying selection on erythrocyte-binding antigens of Plasmodium falciparum and P. vivax. Genetics 163, 1327–1336 (2003).
Ferreira, M.U., Ribeiro, W.L., Tonon, A.P., Kawamoto, F. & Rich, S.M. Sequence diversity and evolution of the malaria vaccine candidate merozoite surface protein-1 (MSP-1) of Plasmodium falciparum. Gene 304, 65–75 (2003).
Mu, J. et al. Recombination hotspots and population structure in Plasmodium falciparum. PLoS Biol. 3, e335 (2005).
Rich, S.M., Licht, M.C., Hudson, R.R. & Ayala, F.J. Malaria's Eve: evidence of a recent population bottleneck throughout the world populations of Plasmodium falciparum. Proc. Natl. Acad. Sci. USA 95, 4425–4430 (1998).
Hughes, A.L. & Verra, F. Extensive polymorphism and ancient origin of Plasmodium falciparum. Trends Parasitol. 18, 348–351 (2002).
Volkman, S.K. et al. Recent origin of Plasmodium falciparum from a single progenitor. Science 293, 482–484 (2001).
Conway, D.J. et al. Origin of Plasmodium falciparum malaria is traced by mitochondrial DNA. Mol. Biochem. Parasitol. 111, 163–171 (2000).
Joy, D.A. et al. Early origin and recent expansion of Plasmodium falciparum. Science 300, 318–321 (2003).
Delemarre, B.J. & van der Kaay, H.J. Tropical malaria contracted the natural way in the Netherlands. Ned. Tijdschr. Geneeskd. 123, 1981–1982 (1979).
Robson, K.J., Hall, J.R., Davies, L.C., Crisanti, A., Hill, A.V. & Wellems, T.E. Polymorphism of the TRAP gene of Plasmodium falciparum. Proc. Biol. Sci. 242, 205–216 (1990).
Kaneko, O., Soubes, S.C. & Miller, L.H. Plasmodium falciparum: invasion of Aotus monkey red blood cells and adaptation to Aotus monkeys. Exp. Parasitol. 93, 116–119 (1999).
Lewontin, R.C. On measures of gametic disequilibrium. Genetics 120, 849–852 (1988).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat. Genet. 25, 25–29 (2000).
Volkman, S.K. et al. Excess polymorphisms in genes for membrane proteins in Plasmodium falciparum. Science 298, 216–218 (2002).
Mu, J. et al. Genome-wide variation and identification of vaccine targets in the Plasmodium falciparum genome. Nat. Genet. advance online publication 10 December 2006 (doi:10.1038/ng1924).
Sabeti, P.C. et al. Positive natural selection in the human lineage. Science 312, 1614–1620 (2006).
Roper, C. et al. Antifolate antimalarial resistance in southeast Africa: a population-based analysis. Lancet 361, 1174–1181 (2003).
Roper, C. et al. Intercontinental spread of pyrimethamine-resistant malaria. Science 305, 1124 (2004).
Reich, D.E. et al. Linkage disequilibrium in the human genome. Nature 411, 199–204 (2001).
Su, X. et al. A genetic map and recombination parameters of the human malaria parasite Plasmodium falciparum. Science 286, 1351–1353 (1999).
International HapMap Consortium. A haplotype map of the human genome. Nature 437, 1299–1320 (2005).
Waterston, R.H. et al. Initial sequencing and comparative analysis of the mouse genome. Nature 420, 520–562 (2002).
Lindblad-Toh, K. et al. Genome sequence, comparative analysis and haplotype structure of the domestic dog. Nature 438, 803–819 (2005).
Sabeti, P.C. et al. Detecting recent positive selection in the human genome from haplotype structure. Nature 419, 832–837 (2002).
Jeffares, D.C. et al. Genome variation and evolution of the malaria parasite Plasmodium falciparum. Nat. Genet. advance online publication 10 December 2006 (doi:10.1038/ng1931).
Acknowledgements
We thank all members of the sample collection team in Senegal (A. Ahouidi, L. Ndiaye, O. Ly, Y. Diedhiou, T. Sene, A. Mbaye and D. Diop). We thank T. Taylor, K. Seydel, J. Montgomery, E. Dembo, M. Molyneux and S. Rogerson for help collecting the samples from Malawi. We also thank all the members of the Broad Sequencing Platform and M. Koehrsen, D. Richter and O. Shamovsky for sequence analysis help. Our thanks to M. Goyette and T. Rachupka for help with Sequenom genotyping. Thanks to N. Mahesh and G. Ramirez for help with sample preparation and parasite cultures and to J. Mu and X. Su for typing samples. We thank MR4 for providing us with malaria parasites contributed by the following depositors: W.E. Collins (MRA-347); D.E. Kyle (MRA-154, MRA-157, MRA-159, MRA-176, MRA-207, MRA-284, MRA-285); L.H. Miller and D. Baruch (MRA-331, MRA-326, MRA-328); W. Trager (MRA-731, MRA-732, MRA-733); D. Walliker (MRA-151, MRA-153, MRA-200); T.E. Wellems (MRA-155, MRA-156, MRA-309, MRA-464); and Y. Wu (MRA-201). Special thanks to J. Barnwell for providing P. reichenowi DNA. Our thanks to PlasmoDB (http://www.plasmodb.org) for access to the 3D7 genome sequence. The authors are supported by the US National Institutes of Health, SPARC funding of The Broad Institute of MIT and Harvard, the Burroughs-Wellcome Fund, The Bill and Melinda Gates Foundation, the NIAID Microbial Sequencing Center, the Ellison Medical Foundation, Fogarty International and the Exxon Mobil Foundation. P.C.S. is funded by the Damon Runyon Cancer Fellowship and the L'Oreal for Women in Science Award.
Author information
Authors and Affiliations
Contributions
S.K.V. designed experiments, prepared samples, carried out Sequenom genotyping and PCR resequencing, analyzed genotyping data and wrote the paper. P.C.S. designed experiments; carried out Sequenom genotyping and PCR resequencing; analyzed data for linkage disequilibrium, allele frequency and long-range haplotypes and wrote the paper. D.D.C. analyzed sequence data, prepared sequence data for subsequent analysis and defined polymorphisms. D.E.N. analyzed data for allele frequency and nucleotide diversity by GO category. S.F.S. analyzed data for nucleotide diversity and selective sweeps. D.A.M., J.P.D., O.S., D.N., O.N., S.M. and M.T.D. helped with sample collection. J.P.D. and M.T.D. helped with culture adaptation. D.A.M. and A.L. helped with parasite cultures. A.D. helped with GO function analysis. N.S.-T. and J.Z. prepared libraries for sequencing. S.W. and R.O. helped with PCR resequencing. L.Z. helped with Sequenom genotyping. E.M., S.G. and D.B.J. created the genome assemblies for HB3 and Dd2. R.C.W. coordinated project flow. B.W.B. supervised sequencing. D.L.H. consulted on population genetic analysis. J.E.G. supervised and advised on sequence analysis. E.S.L. designed experiments, consulted on project outcomes and wrote the paper. D.F.W. designed experiments, coordinated all efforts, supervised project at all levels, consulted on project outcomes and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Sample correspondence for SNPs. (PDF 408 kb)
Supplementary Fig. 2
Separation of continental populations using genotyping SNPs. (PDF 396 kb)
Supplementary Fig. 3
Allele frequency in spectra. (PDF 207 kb)
Supplementary Table 1
Parasites used in the study. (PDF 427 kb)
Supplementary Table 2
Twenty chromosomal regions selected for LD study. (PDF 264 kb)
Supplementary Table 3
Nucleotide diversity across the genome. (PDF 258 kb)
Supplementary Table 4
Comparison of genotyping from whole genome–amplified samples and the original unamplified sample. (PDF 243 kb)
Rights and permissions
About this article
Cite this article
Volkman, S., Sabeti, P., DeCaprio, D. et al. A genome-wide map of diversity in Plasmodium falciparum. Nat Genet 39, 113–119 (2007). https://doi.org/10.1038/ng1930
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng1930
This article is cited by
-
Multiplexed ddPCR-amplicon sequencing reveals isolated Plasmodium falciparum populations amenable to local elimination in Zanzibar, Tanzania
Nature Communications (2023)
-
Genomic analysis reveals independent evolution of Plasmodium falciparum populations in Ethiopia
Malaria Journal (2021)
-
Advances and opportunities in malaria population genomics
Nature Reviews Genetics (2021)
-
Genetic diversity of Plasmodium falciparum in Grande Comore Island
Malaria Journal (2020)
-
Plasticity and genetic variation in traits underpinning asexual replication of the rodent malaria parasite, Plasmodium chabaudi
Malaria Journal (2019)