Abstract
We examine the allometric (comparative scaling) relationships between rates of neurodegeneration resulting from equivalent mutations in a diverse group of genes from five mammalian species with different maximum lifespan potentials. In both retina and brain, rates of neurodegeneration vary by as much as two orders of magnitude and are strongly correlated with maximum lifespan potential and rates of formation of mitochondrial reactive oxygen and nitrogen species (RONS). Cell death in these disorders is directly or indirectly regulated by the intrinsic mitochondrial cell death pathway. Mitochondria are the main source of RONS production and integrate cellular stress signals to coordinate the intrinsic pathway. We propose that these two functions are intimately related and that steady-state RONS-mediated signaling or damage to the mitochondrial stress-integration machinery is the principal factor setting the probability of cell death in response to a diverse range of cellular stressors. This provides a new and unifying framework for investigating neurodegenerative disorders.
Similar content being viewed by others
Main
Cell death in inherited neurodegenerative disorders most commonly occurs by apoptosis1,2,3,4. The evidence is often indirect and inferred on the basis of characteristic morphological changes, such as chromatin compaction and cell shrinkage, or evidence of caspase activation, but in many cases it is more reliably based on multiple lines of evidence1,2,3,4. Caspase-independent cell death also occurs, although the precise mechanisms are less well understood and evidence that this occurs in inherited neurodegenerations is sparse. In these disorders, cell death occurs with a probability that is constant through part or all of the adult lifespan, consistent with a steady-state (one-hit or exponential kinetics) rather than a cumulative damage model5,6. The retina is a particularly well understood model of neurodegeneration. Mutations in more than 100 genes are known to cause neurodegeneration of retinal photoreceptors (RetNet; see URL below); these affect virtually all areas of metabolism and all parts of the cell7,8,9.
Two main pathways lead to the activation of cysteine proteases or caspases that execute the apoptotic death program10,11,12. The extrinsic pathway involves cell-surface signaling complexes (e.g., Fas or FasL), which trigger the cell death cascade by activating caspase 8 and downstream caspases. The intrinsic, or mitochondrial, pathway controls activation of caspase 9, through the adaptor molecule Apaf-1, by regulating release of cytochrome c from mitochondria. Proapoptotic and antiapoptotic members of the Bcl-2 family and various stress and survival signals regulate the release of cytochrome c. Proapoptotic signals can also release proteins such as Smac and Omi, which antagonize IAP (inhibitor of apoptosis) proteins to activate cell death caspases, from the mitochondrial intermembrane space.
Evidence from animal models of human photoreceptor degeneration suggests that the mitochondrial, rather than the extrinsic, cell death pathway has a predominant role3,13,14. The intrinsic pathway also seems to predominate in the brain1,2, but extrinsic and caspase-independent pathways, such as autophagy, may also have a role15. These pathways are often regulated and amplified by mitochondria10,15, emphasizing the crucial role of these organelles in integrating cellular stress signals11,12.
Mitochondria are also the main source of RONS and are implicated in cell and organismal aging, neurodegeneration and apoptosis16,17,18. This raises the question as to whether differences in rates of degeneration between species could be related to differences in mitochondrial RONS production, which is thought to be the primary influence on maximum lifespan potential (LSP)16,17,18. We therefore examined the rates of neurodegeneration caused by functionally equivalent mutations in orthologous genes across mammalian species with different LSPs, here used as a surrogate for constitutive RONS formation.
Measuring rates of degeneration
We first estimated rates of human retinal degenerations using cross-sectional data obtained from a single clinical research laboratory. We then compared rates of photoreceptor degeneration caused by functionally equivalent mutations in five different genes with highly conserved functions in mammalian retina7,8,9 across five species with different LSPs. The mutations we examined cause retinal degeneration either as a primary disorder of rod photoreceptors (RHO, rhodopsin) or as a disorder of both rod and cone photoreceptors (RPGR, retinitis pigmentosa GTPase regulator; RDS (also called PRPH2), retinal degeneration slow or peripherin 2; TULP1, tubby-like protein 1; ATXN7 (also called SCA7), ataxin 7 or spinocerebellar ataxia 7). Most cause rates of degeneration that are consistent with a one-hit exponential decay model of cell death5,6,9.
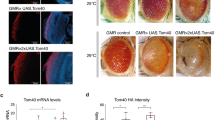
We estimated rates of photoreceptor degeneration by averaging the dark-adapted thresholds sampled across the retina in individuals classified according to the type of mutation they carry (Fig. 1a). We categorized disease severity as mild, moderate or severe to permit comparison with cell-loss data from other species (Supplementary Methods online). One of the most rapidly progressive human disorders, resulting in severe disease by age 27.8 ± 6.3 y (mean ± s.d.; n = 19 from 12 unrelated families), was caused by mutations in RPGR. RHO mutations have been divided into two classes on the basis of psychophysical testing19,20. Class A mutations cause severely abnormal rod function early in life, whereas Class B mutations are compatible with functional rods either regionally (Class B1) or uniformly (Class B2) across the retina. Class A mutations resulted in severe disease by age 31.4 ± 5.4 y (n = 20 from 11 families), whereas class B1 mutations resulted in moderate disease by age 43.2 ± 9.4 y and in severe disease by age 47.4 ± 8.3 y (n = 33 from 12 families). RDS mutations caused a spectrum of phenotypes, including moderate disease by age 58.8 ± 8.4 y (n = 34 from 15 families). We obtained similar data for TULP1 and ATXN7 mutations, although the sample sizes were smaller.
(a) Estimating rates of human retinal degeneration. Mean dark-adapted sensitivity of the retina as a function of age (left panels) in individuals with hereditary retinal degenerations due to mutations in RPGR, RHO and RDS. Visual field maps of dark-adapted sensitivity on a grayscale (right panels) are shown for representative individuals (symbols with crosshairs) in each group. The mean sensitivity abnormalities derived from individual maps are divided into three categories: mild, moderate and severe. Age (mean ± s.d.) is shown for categories with more than five individuals (colored symbols with error bars). Range of normal (N) sensitivity is shown. (b) Relationship between age to reach a criterion level of neurodegeneration and maximum LSP in five species49,50: mouse (3.5 y), rat (4.7 y), dog (20 y), pig (27 y) and human (122 y). Retinal degeneration is shown in the main panel and brain degeneration in the inset. Human retina results are from a. Faster rates of neurodegeneration are represented by fewer years to reach the criterion level of degeneration on the ordinate. Linear regressions are shown. Smaller symbols and dashed lines for TULP1 (n = 4) and ATXN7 (n = 2) indicate the small numbers of individuals in these groups.
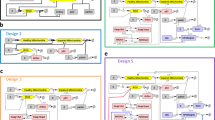
We examined the relationship between severity of degeneration (estimated as time to reach mild, moderate or severe degeneration) and LSP using the data above from humans and published data from animal models (mouse, rat, dog, pig) with mutations of the same class, based on functional criteria, in the orthologous gene (Fig. 1b and Supplementary Table 1 online). We observed a markedly similar relationship across genes and across species. For example, equivalent mutations in RPGR and its orthologs cause severe retinal degeneration in mice at age 0.275 y, dogs at age 2 y and humans at age 28 y. Rates of degeneration within gene or mutational classes were consistent with an allometric equation21, TD = a × LSPn, where TD is the time (in y) to reach a given level of degeneration, a is a gene- or mutation class–specific constant (increasing with disease severity) and n is the allometric exponent. The mean value of n across all five genes was 1.22 ± 0.11 (range 1.00–1.45; Table 1), implying that TD increases in almost direct proportion to LSP. Hence, the rate of degeneration is inversely related to LSP in all these genes, which have disparate functions8,9 (Fig. 1b).
We made similar observations for inherited neurodegenerations of the brain. Many different neurodegeneration models have been constructed, only some of which are directly comparable with the human disorders (Supplementary Table 1 online). Again, the selected transgenic mice faithfully modeled the human disorders, implying that their functions are well conserved. For example, transgenic mice expressing the A53T amino acid substitution in α-synuclein (Snca) developed signs of disease within 2 y, about 20 times faster than humans with familial Parkinson disease carrying the same SNCA mutation. We therefore examined cross-species data for neurodegenerations resulting from mutations in the genes HD (Huntington disease), SOD1 (familial amyotrophic lateral sclerosis), PRNP (Gerstmann-Straussler-Scheinker syndrome), SNCA (familial Parkinson disease) and APP (familial Alzheimer disease; Fig. 1b and Table 1). The mean value of the allometric exponent, n, in the equation given above across these five genes was 0.92 ± 0.21 (range 0.50–1.34), which is not significantly different from the value across genes involved in retinal degeneration and is again consistent with a direct (inverse) relationship between LSP and rate of degeneration. The mean allometric exponent for all ten genes was 1.1 ± 0.2 (range 0.7–1.5, n = 12 comparisons; Table 1).
The models included 14 transgenic mice, 3 naturally occurring mutations and 1 knockout mouse. To avoid bias resulting from transgene overexpression, we used models in which the expression levels of the transgene were comparable with those of the native allele. Of the 18 models, 5 express the transgene product at slightly less than the native level, 10 express it at the native level, and 3 express it at two to three times the native level (Supplementary Table 1 online). Therefore, transgene overexpression is unlikely to be a significant confounding factor.
To validate the use of LSP as a surrogate for mitochondrial RONS production16,17,18, we plotted published rates of RONS formation in 11 species and 4 postmitotic tissues (kidney, heart, brain and liver)22,23,24,25,26 to show that a strong (r2 = 0.71 ± 0.15) inverse relationship exists with LSP, with a mean allometric exponent of −0.84 ± 0.34 (Fig. 2).
Rates of RONS formation are shown for kidney, heart, brain and liver in mouse (LSP = 3.5 y), hamster (4 y), rat (4.5 y), guinea pig (7.5 y), pigeon (8 y),white-footed mouse (Peromyscus leucopus; 8 y), rabbit (18 y), parakeet (21 y), canary (24 y), pig (27 y) and cow (30 y). Data points were redrawn from data in refs. 22–26. Linear regressions are shown. Superoxide formation data in the liver are given at 1% of their true values.
Interpreting allometric differences
Our analysis suggests that the mechanisms concerned with allometric differences in rates of degeneration are both independent of the precise mode or kinetics of cell death and general to the nervous system. The neurodegenerative disease models that we examined vary in terms of the kinetics of neuronal loss. Some clearly differ from the simple exponential model seen in most retinal degenerations. For example, the rate of substantia nigra cell loss in Parkinson disease is exponential beginning early in life but shows a ten-fold increased exponential rate beginning ∼5 y before the onset of symptoms27. The precise cell death pathway may also vary between disorders. For example, although the animal model of juvenile Huntington disease included in this analysis showed apoptotic neurodegeneration, models containing large numbers of glutamine repeats show evidence of nonapoptotic cell death28. Autophagy is a caspase-independent mode of cell death that seems to be involved in some neurodegenerations15, but both caspases and autophagy can be simultaneously activated in dying cells, and so different cell types and contexts may trigger the use of caspases, the ubiquitin-proteasome pathway, autophagy or perhaps other mechanisms to destroy the cell4,15. Most existing evidence suggests that cell death occurs by a predominantly apoptotic mechanism that is directly or indirectly regulated by the mitochondrial pathway1,2,3,4,8,9,13,14. Our data support this notion and suggest that, although other proteolytic mechanisms may operate to varying degrees in the final execution phase4,15, mitochondrial regulation lies at the core of the cell death pathway in neurodegeneration.
The strong correlations between LSP, mitochondrial RONS formation and rate of neurodegeneration across distinct genes and species could result from indirect or chance factors, but these do not stand up well to scrutiny. First, species differences in the genes or their pathways might alone account for the differing rates of neurodegeneration. But the genes that we analyzed all act within functional pathways that are highly conserved across mammalian species1,2,8,9, and so this factor cannot account for the consistency of the relationship across genes and species or for the large allometric differences in rates of degeneration (spanning two orders of magnitude; Fig. 1b). Second, because physiological time runs faster in smaller than in larger mammals, rates of degeneration should perhaps behave accordingly21, thus accounting for the observed correlations. This would not be the case, however, if disease pathogenesis is independent of the physiological mechanisms regulating metabolic rates and longevity. The principal factors driving species differences in basal metabolic rate (BMR) are still not known29,30,31. BMR reflects a series of linked metabolic processes, as shown by the similarity in allometric scaling exponents in the equation relating BMR per cell (Bc) to body mass (M), such that Bc α M−1/4 for many such processes21,29. A proposed 'pacemaker' for such interlinked processes is the degree of polyunsaturation of membrane phospholipids, which regulates the activities that contribute the most to cellular energy demand, such as maintenance of sodium and mitochondrial proton gradients (which together account for ∼50% of BMR)29,30,31,32. The degree of polyunsaturation of most membrane lipids declines with LSP, but this is not the case in the brain or retina, which are unusual in their high requirement for polyunsaturated membrane lipids29,30. Another possible source of allometric differences in BMR is basal proton leak through the mitochondrial inner membrane, which is responsible for ∼20% of BMR and correlates well with LSP in mammals (but not birds)31,33. Whatever the mechanisms driving species differences in BMR, they probably do not have a large influence on neurodegeneration, as BMR is poorly correlated with lifespan16,17,18.
The reasons for species differences in LSP are only partially understood, but evolutionary studies have clarified their origins18,34. They seem to be the evolutionary consequence of different habitats and lifestyles16,17,18,34. High extrinsic mortality rates, as in mice, lead to weak selection for genetic modifications that prolong reproductive and lifespans34. In species with low extrinsic mortality rates, a key target for evolutionary modification of LSP seems to be mitochondrial RONS formation (rather than BMR or body size), which is strongly correlated with LSP16,17,18. In contrast, whereas antioxidant defense mechanisms are prominent in aerobic tissues such as retina and brain, cross-species comparisons show that they are either uncorrelated or negatively correlated with LSP18. Transgenic overexpression of antioxidant enzymes, such as superoxide dismutase, can have large effects on lifespan35,36,37, but naturally occurring antioxidant defenses do not readily account for differences in LSP18.
Environmental and genetic influences that extend lifespan, such as caloric restriction and >40 mutations associated with longevity in nematodes, are each associated with increased resistance to oxidative stress35. Both caloric restriction and many of these mutants use a common pathway involving SIRT protein deacetylases and the forkhead transcription factor family (FOXO or Daf-16), which can upregulate mitochondrial Sod2 and other antioxidant defenses and reduce apoptotic cell death36,37. Antioxidant defenses are therefore important influences on both cell death and longevity, but evolution seems to have favored limiting RONS production rather than increasing their destruction18, perhaps because RONS also have cellular signaling roles.
This suggests a third possible explanation for the correlation between neurodegeneration rate and LSP: that oxidative stress is a key pathogenetic factor common to each of the neurodegenerations. This explanation does not fit the data well, as most of the genes known to cause inherited neurodegeneration in the retina do not directly influence RONS formation, implying that they have an indirect effect, at most, on oxidative stress. In the brain, the situation is more complex as protein aggregation and oxidative stress have been invoked to varying degrees in all of the neurodegenerations, but there is no consensus that oxidative stress is other than one of several pathogenetic influences1,2,38. There are certainly few common pathogenetic factors among the >100 genes involved in retinal degeneration in mammals7,8,9, and similar diversity is emerging in other forms of neurodegeneration1,2. The genes involved affect most aspects of metabolism and most subcellular compartments. In the retina, exposure of photoreceptors to real or potential oxidative stress, arising from phototoxicity, high aerobic metabolism and membranes rich in polyunsaturates, is shared by all mammalian retinas and thus cannot explain the species differences. The situation in brain is similar. This vulnerability may, however, place these tissues closer than less aerobic tissues to the apoptotic threshold. The only common factor among the diversity of mutations causing neurodegeneration (which affect signal transduction, maintenance of intracellular structures, protein turnover, intracellular transport, transcriptional regulation or RNA splicing or cause protein aggregation and the unfolded protein response) is that all act as cellular stressors1,2,8,9.
Mitochondrial RONS influence cell death
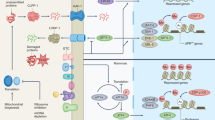
The organelle principally involved in the integration of such stressors, and of proapoptotic and antiapoptotic signals in general, is the mitochondrion, working in close liaison with the endoplasmic reticulum11,12,39. Mitochondria are the barometer of the cell, responding to the state of health or disease in all parts of the cell, from the nucleus to the cytoplasm, integrating the disparate stresses to repress or activate the apoptotic pathway. Mitochondria are therefore implicated in neurodegenerative cell death, suggesting that it is no coincidence that they are also the main source of RONS formation in the cell16,17.
The strong allometric correlations between LSP, mitochondrial RONS production and rates of neurodegeneration are suggestive of a unifying hypothesis in which steady-state RONS production, acting on the mitochondrial stress integration machinery, is not just a minor player but is the principal factor in setting the probability of apoptosis for a given species and cell type in response to a cellular stress (Fig. 3). No other factor easily accounts for the >100-fold difference in neurodegeneration rates across mammalian species.
Species- and cell type–specific differences in constitutive RONS production may act on the apoptotic threshold through two mechanisms: (i) by redox setting of the gain on signaling pathways concerned with cell survival (e.g., neurotrophin signaling); and (ii) by influencing the steady-state equilibrium between oxidative damage and repair to mitochondrial membranes, in which proapoptotic and antiapoptotic protein-lipid interactions regulate the commitment to apoptosis. A diverse range of cellular stressors, such as inherited mutations causing neurodegeneration, are proposed to alter the probability of apoptosis through effects on the intrinsic mitochondrial pathway. The main factor influencing the probability of apoptosis seems to be not the stress itself but rather species-specific influences on the apoptotic threshold. Small mammals with short LSP have a high probability of apoptosis, whereas large mammals with long LSP have a low probability of apoptotic cell death.
Mitochondria are the primary source of constitutive RONS formation, as they consume ∼90% of delivered oxygen and continuously convert 0.1–1% of it to RONS in the inner mitochondrial membrane16,17,18. The main physiological source of RONS such as superoxide has been proposed to be reverse electron flow into complex I, but complex III and other flavoproteins may also be involved40. RONS production is implicated in a wide variety of cell death events16,17,18. We propose that two related aspects of steady-state RONS formation influence the probability of apoptosis in neurodegeneration: (i) RONS effects on signal transduction pathways or (ii) net RONS damage to lipids and macromolecules (including DNA) influencing the mitochondrial stress-integration machinery.
The first possibility, effects of RONS on signal transduction, may act through redox regulation of crucial cysteine thiols, influencing signaling pathways implicated in cell survival, RONS production and apoptosis41,42. Modification of specific transcription factors, protein phosphatases or uncoupling proteins may affect the apoptotic threshold, the point of no return (such as the Bak-Bax gateway12) or other RONS-mediated signaling effects on apoptosis43.
Alternatively, the critical factor may be steady-state RONS damage to key mitochondrial components. This assumes that there is an equilibrium between sublethal damage and repair to macromolecules and lipids influencing mitochondrial membranes, which are close to sites of RONS production and directly interact with both proapoptotic and antiapoptotic proteins44. The sensitivity of the mitochondrial stress-integration machinery to an apoptotic trigger may therefore be regulated by RONS. For example, differences in acyl composition or reduced amounts of mitochondrial cardiolipin may influence cytochrome c release into the intermembrane space45, awaiting a second event, such as Bax- or Bid-induced permeabilization of the outer mitochondrial membrane, to trigger its release into the cytoplasm11,12,44. The degree of membrane lipid polyunsaturation may or may not influence oxygen consumption but is an indicator of susceptibility to RONS damage30, which could have a large effect on Bcl-2 family protein–lipid interactions influencing apoptosis.
The final trigger for the execution pathway may follow a lethal hit5,6, such as RONS damage to a respiratory chain complex. The probability of such a hit will also be influenced by constitutive RONS production and long-term influences on cellular redox tone. Mammals with high constitutive RONS production and short LSP may simply be closer to the apoptotic threshold, with a higher probability of cell death, than those with a long LSP (Fig. 3).
Targeting antioxidants to mitochondria
There are few situations in which neurodegeneration rates can be altered as substantially as the allometric differences shown here. Therefore, elucidation of the mechanisms concerned may provide new therapeutic approaches and a new framework for investigating neurodegeneration. What are the implications for modifying rates of cell death? First, antioxidant therapy in neurodegenerations may fail because it is inappropriately targeted. Targeting of powerful antioxidant molecules to the mitochondrial matrix or membranes may herald a new era in antioxidant therapy46,47. Rather than counteracting disease-induced oxidative stress, the principal value of such therapy may be to reduce constitutive RONS influences on the apoptotic gateway. Second, identification of redox signaling pathways and key lipid or protein RONS targets affecting the apoptotic gateway could provide new therapeutic targets. Third, metabolically active postmitotic cells in other tissues, such as heart, pancreas and kidney, in which cell death contributes to disease may be targets for similar therapeutic strategies, as they are likely to be subject to similar oxidative influences. Last, longevity is strongly influenced by the gradual attrition of postmitotic cells in vital organs and by the diseases that afflict them36,48. At advanced ages, the probability of cell death is no longer constant but increases with time, as does the risk of cancer in dividing cells. This is no longer a steady-state situation and implies the failure of oxidative, immune or other defenses. Regardless, a reduction in constitutive RONS formation is likely to reduce the probability of cell death, irrespective of age and cellular stress. Efficient targeting of antioxidant molecules to mitochondrial membranes in the most susceptible cells may both slow rates of degeneration and improve the quality of life in old age.
URL. RetNet, the Retinal Information Network, is available at http://www.sph.uth.tmc.edu/Retnet/.
Note: Supplementary information is available on the Nature Genetics website.
References
Mattson, M.P. Apoptosis in neurodegenerative disorders. Nat. Rev. Mol. Cell Biol. 1, 120–129 (2000).
Nijhawan, D., Honarpour, N. & Wang, X. Apoptosis in neural development and disease. Annu. Rev. Neurosci. 23, 73–87 (2000).
Reme, C.E., Grimm, C., Hafezi, F., Marti, A. & Wenzel, A. Apoptotic cell death in retinal degenerations. Prog. Retin. Eye Res. 17, 443–464 (1998).
Abraham, M.C. & Shaham, S. Death without caspases, caspases without death. Trends Cell Biol. 14, 184–193 (2004).
Clarke, G. et al. A one-hit model of cell death in inherited neuronal degenerations. Nature 406, 195–199 (2000).
Clarke, G., Lumsden, C.J. & McInnes, R.R. Inherited neurodegenerative diseases: the one-hit model of neurodegeneration. Hum. Mol. Genet. 10, 2269–2275 (2001).
Daiger, S.P. Identifying retinal disease genes: how far have we come, how far do we have to go? Novartis Found. Symp. 255, 17–27 (2004).
Rattner, A., Sun, H. & Nathans, J. Molecular genetics of human retinal disease. Annu. Rev. Genet. 33, 89–131 (1999).
Pacione, L.R., Szego, M.J., Ikeda, S., Nishina, P.M. & McInnes, R.R. Progress toward understanding the genetic and biochemical mechanisms of inherited photoreceptor degenerations. Annu. Rev. Neurosci. 26, 657–700 (2003).
Wolf, B.B. & Green, D.R. Apoptosis: letting slip the dogs of war. Curr. Biol. 12, R177–R179 (2002).
Newmeyer, D.D. & Ferguson-Miller, S. Mitochondria: releasing power for life and unleashing the machineries of death. Cell 112, 481–490 (2003).
Danial, N.N. & Korsmeyer, S.J. Cell death: critical control points. Cell 116, 205–219 (2004).
LaVail, M.M. et al. Protection of mouse photoreceptors by survival factors in retinal degenerations. Invest. Ophthalmol. Vis. Sci. 39, 592–602 (1998).
Hafezi, F., Grimm, C., Simmen, B.C., Wenzel, A. & Reme, C.E. Molecular ophthalmology: an update on animal models for retinal degenerations and dystrophies. Br. J. Ophthalmol. 84, 922–927 (2000).
Yu, L., Lenardo, M.J. & Baehrecke, E.H. Autophagy and caspases: a new cell death program. Cell Cycle 3, 1124–1126 (2004).
Halliwell, B. & Gutteridge, J.M.C. Free Radicals in Biology and Medicine 3rd edn. (Oxford University Press, Oxford, 1999).
Beckman, K.B. & Ames, B.N. The free radical theory of aging matures. Physiol. Rev. 78, 547–581 (1998).
Perez-Campo, R., Lopez-Torres, M., Cadenas, S., Rojas, C. & Barja, G. The rate of free radical production as a determinant of the rate of aging: evidence from the comparative approach. J. Comp. Neurol. B 168, 149–158 (1998).
Gal, A., Apfelstedt-Sylla, E., Janecke, A.R. & Zrenner, E. Rhodopsin mutations in inherited retinal dystrophies and dysfunctions. Prog. Retinal Eye Res. 16, 51–79 (1997).
Cideciyan, A.V. et al. Disease sequence from mutant rhodopsin allele to rod and cone photoreceptor degeneration in man. Proc. Natl. Acad. Sci. USA 95, 7103–7108 (1998).
Schmidt-Nielsen, K. Scaling: Why Is Animal Size So Important? (Cambridge University Press, Cambridge, 1984).
Sohal, R.S., Sevensson, I., Sohal, B.H. & Brunk, U.T. Superoxide anion radical production in different animal species. Mech. Ageing Dev. 49, 129–135 (1989).
Sohal, R.S., Svensson, I. & Brunk, U.T. Hydrogen peroxide production by liver mitochondria in different species. Mech. Ageing Dev. 53, 209–215 (1990).
Ku, H.-H., Brunk, U.T. & Sohal, R.S. Relationship between mitochondrial superoxide and hydrogen peroxide production and longevity of mammalian species. Free Radic. Biol. Med. 15, 621–627 (1993).
Barja, G. Mitochondrial free radical production and aging in mammals and birds. Ann. NY Acad. Sci. 854, 224–238 (1998).
Ku, H.-H. & Sohal, R.S. Comparison of mitochondrial pro-oxidant generation and antioxidant defences between rat and pigeon: possible basis of variation in longevity and metabolic potential. Mech. Aging Dev. 72, 67–76 (1993).
Dunnett, S.B. & Bjorklund, A. Prospects for new restorative and neuroprotective treatments in Parkinson's disease. Nature 399 (Suppl), A32–A39 (1999).
Turmaine, M., Raza, A., Mahal, A., Mangiarini, L., Bates, G.P. & Davies, S.W. Nonapoptotic neurodegeneration in a transgenic mouse model of Huntington's disease. Proc. Natl. Acad. Sci. USA 97, 8093–8097 (2000).
Hulbert, A.J. & Else, P.L. Mechanisms underlying the cost of living in animals. Annu. Rev. Physiol. 62, 207–235 (2000).
Pamplona, R., Barja, G. & Portero-Otin, M. Membrane fatty acid unsaturation, protection against oxidative stress, and maximum life span. Ann. NY Acad. Sci. 959, 476–490 (2002).
Hulbert, A.J. & Else, P.L. Membranes as possible pacemakers of metabolism. J. Theor. Biol. 199, 257–274 (1999).
Turner, N., Else, P.L. & Hulbert, A.J. Docosahexaenoic acid (DHA) content of membranes determines molecular acivity of the sodium pump: implications for disease states and metabolism. Naturwissenschaften 90, 521–523 (2003).
Porter, R.K., Hulbert, A.J. & Brand, M.D. Allometry of mitochondrial proton leak: influence of membrane surface area and fatty acid composition. Am. J. Physiol. 271, R1550–R1560 (1996).
Partridge, L. Evolutionary biology and age-related mortality. in Between Zeus and the Salmon (eds. Wachter, K.W. & Finch, C.E.) 78–95 (National Academy Press, Washington, DC, 1997).
Sampayo, J.N., Gill, M.S. & Lithgow, G.J. Oxidative stress and aging—the use of superoxide dismutase/catalase mimetics to extend lifespan. Biochem. Soc. Trans. 31, 1305–1307 (2003).
Antebi, A. Tipping the balance toward longevity. Dev. Cell. 6, 315–316 (2004).
Daitoku, H. et al. Silent information regulator 2 potentiates Foxo1-mediated transcription through its deacetylase activity. Proc. Natl. Acad. Sci. USA 101, 10042–10047 (2004).
Andersen, J.K. Oxidative stress in neurodegeneration: cause or consequence? Nat. Med. 10 (Suppl), S18–S25 (2004).
Boehning, D. et al. Cytochrome c binds to inositol (1,4,5) trisphosphate receptors, amplifying calcium-dependent apoptosis. Nat. Cell Biol. 5, 1051–1061 (2003).
St-Pierre, J., Buckingham, J.A., Roebuck, S.J. & Brand, M.D. Topology of superoxide production from different sites in the mitochondrial electron transport chain. J. Biol. Chem. 277, 44784–44790 (2002).
Shiva, S. et al. Redox signalling: from nitric oxide to oxidised lipids. in Free Radicals: Enzymology, Signalling and Disease (eds. Cooper, C.E., Wilson, M.T. & Darley-Usmar, V.M.) 107–120 (Portland, London, 2004).
Kamata, H. & Hirata, H. Redox regulation of cellular signalling. Cell. Signal. 11, 1–14 (1998).
Echtay, K.S. et al. A signalling role for 4-hydroxy-2-nonenal in regulation of mitochondrial uncoupling. EMBO J. 22, 4103–4110 (2003).
Adams, J.M. & Cory, S. The Bcl-2 protein family: arbiters of cell survival. Science 281, 1322–1326 (1998).
Ott, M., Robertson, J.D., Gogvadze, V., Zhivotovsky, B. & Orrenius, S. Cytochrome c release from mitochondria proceeds by a two-step process. Proc. Natl. Acad. Sci. USA 99, 1259–1263 (2002).
Jauslin, M.L., Meier, T., Smith, R.A. & Murphy, M.P. Mitochondria-targeted antioxidants protect Friedreich Ataxia fibroblasts from endogenous oxidative stress more effectively than untargeted antioxidants. FASEB J. 17, 1972–1974 (2003).
Echtay, K.S., Murphy, M.P., Smith, R.A., Talbot, D.A. & Brand, M.D. Superoxide activates mitochondrial uncoupling protein 2 from the matrix side. Studies using targeted antioxidants. J. Biol. Chem. 277, 47129–47135 (2002).
Olshansky, S.J., Carnes, B.A. & Cassel, C. In search of Methuselah: estimating the upper limits to human longevity. Science 250, 634–640 (1990).
Altman, P.L. & Dittmer, D.S. Biology Data Book 2nd edn. (FASEB, Bethesda, Maryland, 1972).
Comfort, A. The Biology of Senescence (Elsevier, New York, 1979).
Acknowledgements
We thank M. Murphy and N. Hastie for critical reading and advice on the manuscript and S. B. Schwartz and E. A. M. Windsor for help with data analyses. We acknowledge the financial support of the British Retinitis Pigmentosa Society, Foundation Fighting Blindness, the US National Institutes of Health (National Eye Institute) and the Macula Vision Research Foundation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Wright, A., Jacobson, S., Cideciyan, A. et al. Lifespan and mitochondrial control of neurodegeneration. Nat Genet 36, 1153–1158 (2004). https://doi.org/10.1038/ng1448
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng1448
This article is cited by
-
Rod function deficit in retained photoreceptors of patients with class B Rhodopsin mutations
Scientific Reports (2020)
-
The effect of 17β-estradiol on the expression of dipeptidyl peptidase III and heme oxygenase 1 in liver of CBA/H mice
Journal of Endocrinological Investigation (2015)
-
Activities of DNA base excision repair enzymes in liver and brain correlate with body mass, but not lifespan
AGE (2012)
-
Cerebral Formation of Free Radicals during Hypoxia Does Not Cause Structural Damage and is Associated with a Reduction in Mitochondrial PO2; Evidence of O2-Sensing in Humans?
Journal of Cerebral Blood Flow & Metabolism (2011)
-
Haploinsufficiency of RanBP2 is neuroprotective against light-elicited and age-dependent degeneration of photoreceptor neurons
Cell Death & Differentiation (2009)