Abstract
MicroRNAs (miRNAs) are a class of short (∼22-nt) noncoding RNA molecules that downregulate expression of their mRNA targets. Since their discovery as regulators of developmental timing in Caenorhabditis elegans, hundreds of miRNAs have been identified in both animals and plants1. Here, we report a technique for visualizing detailed miRNA expression patterns in mouse embryos. We elucidate the tissue-specific expression of several miRNAs during embryogenesis, including two encoded by genes embedded in homeobox (Hox) clusters, miR-10a and miR-196a. These two miRNAs are expressed in patterns that are markedly reminiscent of those of Hox genes. Furthermore, miR-196a negatively regulates Hoxb8, indicating that its restricted expression pattern probably reflects a role in the patterning function of the Hox complex.
Similar content being viewed by others
Main
So far, miRNA expression studies in mammals have relied on northern blots, tissue-specific RNA cloning and microarrays2,3,4,5. Though powerful, these techniques lack the resolution to show precise spatiotemporal expression patterns. Standard methods of in situ hybridization are problematic owing to the small size of miRNAs, and although a miRNA in situ hybridization method was recently reported for Arabidopsis thaliana6 and maize7, it has proven difficult to adapt it to vertebrate embryos. Promoter fusion constructs have been useful for visualizing miRNA expression patterns in C. elegans8, but this requires previous knowledge of promoter elements.
Using an alternative approach previously reported for Drosophila melanogaster9, we generated a series of reporter transgenes ('sensors') designed to detect the presence of specific miRNAs in an embryo. Each sensor contains a constitutively expressed reporter (lacZ) bearing sequences complementary to a given miRNA in the 3′ untranslated region (UTR). In cells lacking the miRNA that the sensor is designed to detect, the transgene RNA is stable and allows reporter expression (Fig. 1a). In contrast, in cells that express the miRNA, its perfect complementarity to sequences in the 3′ UTR of the sensor targets the sensor mRNA to the RNA interference (RNAi) pathway, resulting in an absence of β-galactosidase (β-gal) activity10 (Fig. 1b). Thus, in transgenic embryos carrying a sensor, tissues stained for β-gal are white where the target miRNA is present and blue everywhere else. As a control, we used a sensor lacking known miRNA-complementary sites and found that it drove robust β-gal expression throughout the embryo at embryonic day (E) 9.5 and later (Fig. 1c).
(a) A lacZ reporter containing miRNA-complementary sequences in the 3′ UTR constitutively expresses β-gal in cells lacking the complementary miRNA. (b) In cells expressing the miRNA (red), the lacZ message is targeted for degradation by the RNAi pathway. (c) A control sensor lacking any known miRNA complementary sequences drives ubiquitous lacZ expression at E10.5.
We first focused on two vertebrate homologs of the C. elegans let-7 miRNA. This miRNA regulates several genes in the heterochronic pathway, and its sequence and developmentally regulated expression are well conserved across animal phyla11,12. The mouse genome encodes 11 let-7 family members (including miR-let-7a–miR-let-7f, also known as Mirnlet7a–Mirnlet7f in the mouse)1,3. Northern-blot analysis showed that at least four precursor bands, as well as a high level of mature miR-let-7 RNA, were expressed in limbs of developing mice (Fig. 2a). We generated sensors to determine when and where miR-let-7c and miR-let-7e were expressed in the limb and whether different miR-let-7 isoforms were differentially regulated. In embryos carrying either the miR-let-7c or miR-let-7e sensor, β-gal expression was downregulated in limb buds (Fig. 2b,c). In both cases, β-gal expression was more strongly repressed in the distal than the proximal limb, suggesting that miR-let-7c and miR-let-7e had higher expression distally, perhaps relating to the maintenance of an undifferentiated cell population in this region.
(a) Northern-blot analysis of total RNA from forelimb (Fl) and hindlimb (Hl) of an E11.5 mouse embryo probed for miR-let-7a. At least four miR-let-7 precursor (pre-miR-let-7) bands are expressed, and processed miR-let-7 is strongly expressed in both limbs. (b) Whole-mount forelimb from an E11.0 embryo bearing a miR-let-7c sensor. β-gal expression is downregulated in the anterior, distal region of the limb, but not in the posterior region, suggestive of an anterior concentration of miR-let-7c. miR-let-7c is absent from the apical ectodermal ridge, as shown by strong β-gal expression (arrow). (c) Transverse section through the forelimb of an embryo bearing a miR-let-7e sensor. β-gal activity is slightly downregulated throughout the distal portion of the forelimb mesenchyme and is absent (suggestive of high miR-let-7e expression) in the ectoderm (arrow). (d) miR-let-7c and miR-let-7e differ by two nucleotides. The different expression patterns shown by miR-let-7c and miR-let-7e sensors suggest that their regulation by the corresponding miRNAs is sequence-specific.
Notably, in other respects, the limb expression patterns of these two miR-let-7 miRNAs, which differ in sequence by only two nucleotides (Fig. 2d)3, were very different. miR-let-7c was expressed highly in the anterior limb bud but comparatively weakly in the posterior limb and in the chondrogenic core (Fig. 2a and data not shown). miR-let-7c was absent from the apical ectodermal ridge, as evidenced by high β-gal activity in this tissue (Fig. 2a). In contrast, miR-let-7e was expressed weakly in the limb mesenchyme and robustly throughout the limb ectoderm (Fig. 2b).
Data from mouse-organ total RNA libraries identified another miRNA, miR-1, that is highly and specifically expressed in the adult heart3. Consistent with this finding, and in contrast to controls, embryos transiently transgenic with respect to an miR-1-responsive sensor showed a marked reduction in β-gal expression specifically in the heart (Figs. 1c and 3a,b). Notably, β-gal activity was lowest (suggestive of highest miR-1 expression) in the chamber myocardium, particularly the ventricles, and was somewhat higher in the atrioventricular canal, which gives rise to the valves, and in the future ventricular septum and outflow tract (Fig. 3a,c,d). Northern-blot analysis confirmed that miR-1 was present specifically in RNA extracted from heart in both mouse and chick embryos (Fig. 3e).
(a) lacZ expression from a sensor containing miR-1 complementary sequences is specifically downregulated in the heart of an E10.5 embryo. Downregulation (corresponding to expression of miR-1) seems limited to the chambers and not the outflow tract (arrow). (b) Sagittal section through the heart of an E10.5 embryo bearing a control sensor. lacZ is highly expressed in both the atrium (upper chamber) and ventricle (lower chamber). (c) Sagittal section through the heart of an E10.5 embryo bearing an miR-1 sensor. β-gal activity is downregulated in both chambers, suggestive of high miR-1 expression. (d) Frontal section through the heart of an E10.5 embryo bearing an miR-1 sensor, showing a four-chambered view. β-gal expression is downregulated in the chambers, particularly the ventricles (lower chambers), and is less downregulated (suggestive of lower levels of miR-1 expression) in the atrioventricular canal (black arrow) and in cells that give rise to the ventricular septum (white arrow). (e) A northern blot of forelimb (Fl), heart (Ht) and whole-embryo (E) RNA from Hamburger-Hamilton stage-25 chick embryos (Ch)27 and E10.5 mouse embryos (M) probed with miR-1 indicates that miR-1 is specifically expressed in the heart in embryos from both species.
Finally, we assayed the expression of two miRNAs embedded in the Hox clusters, miR-10a and miR-196a13,14,15 (Fig. 4). In the mouse, miR-10a and miR-10b are embedded in equivalent locations in the Hoxb and Hoxd clusters, respectively, and this chromosomal location is conserved in invertebrates13. We constructed a sensor for miR-10a, which lies upstream of Hoxb4, and found that their expression patterns were markedly similar. The miR-10a sensor directed high levels of β-gal activity in the head and anterior trunk but was downregulated in the posterior trunk, with an anterior expression limit similar to that of Hoxb4 (Fig. 5a,b)16. miR-10a was most highly expressed (as indicated by absence of β-gal activity) in a restricted region of the posterior trunk surrounding the hindlimb buds. Notably, Hoxb4 RNA is downregulated by a post-transcriptional mechanism in the same specific region (Fig. 5a,b)17, although obvious miR-10a binding sites were not detected in the Hoxb4 transcript. Nevertheless, the similarities between the expression patterns of miR-10a and Hoxb4 suggest that miR-10a may have a role in establishing or maintaining rostrocaudal cell fates in conjunction with the Hox-Hom complex.
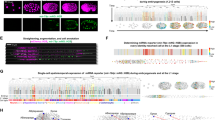
(a) lacZ expression from a sensor containing miR-10a complementary sequences is downregulated in the trunk of an E10.0 embryo. The anterior limit of miR-10a expression is similar to that of the flanking gene, Hoxb4, at the border of mesodermal expression (black arrow). In the posterior part of the embryo, miR-10a expression is complementary to that of Hoxb4. It is most highly expressed in a region posterior to the limb buds but anterior to the tail (white arrow), where Hoxb4 is downregulated by a post-transcriptional mechanism. (b) An in situ hybridization for Hoxb4 shows the previously reported anterior expression limit (top arrow) of Hoxb4 RNA. The transcript is not detected in the posterior trunk (bottom arrow) and is expressed in the tail. (c) A northern blot of Hamburger-Hamilton stage-25 chick (Ch) and E10.5 mouse (M) embryos, bisected at the anterior limit of miR-10a suggested by the sensor. The blot was probed for miR-10a, confirming that in both species expression is high in the posterior (P) and very low in the anterior (A) of the embryo (whole embryo; E). (d) Like miR-10a, miR-196a is expressed in a Hox-like pattern. An miR-196a sensor is downregulated in the posterior trunk and tail, with an anterior expression limit similar to the posterior limit of expression for Hoxb8 RNA. (e) In situ hybridization for Hoxb8, showing previously reported downregulation in the posterior trunk and tail.
We confirmed the posteriorly restricted pattern of expression of miR-10a by northern-blot analysis; this expression pattern was also conserved in chick embryos (Fig. 5c). Because mouse miR-10a and miR-10b differ by a single nucleotide, they are expected to cross-hybridize with the northern-blot probe; thus, both paralogs probably are restricted to the posterior of the embryo. In at least one case (miR-let-7c and miR-let-7e), we found that sensor transgenes could discriminate a difference of two nucleotides, but we do not know whether this is true for a single-nucleotide difference. Therefore, the expression pattern of the miR-10a sensor may be a composite of the expression patterns of miR-10a and miR-10b.
The similarity between the expression patterns of miR-10a and Hoxb4 suggests that transcription of miR-10a may be regulated by cis elements that also regulate Hox genes. Whereas miR-10a and miR-10b lie upstream of HOXB4 and HOXD4 in humans, miR-10b is located in the last intron of Hoxd4 in the mouse, further supporting this hypothesis13.
We also constructed a sensor transgene for the Hox complex–embedded miRNA miR-196a. This sensor was expected to uncover a composite pattern of expression of miR-196a1 and miR-196a2, which have identical mature sequences and are located upstream of Hoxb9 and Hoxc9, respectively14,15. In embryos transgenic with respect to this sensor, β-gal activity was high in the head and anterior trunk and was downregulated in the posterior trunk (Fig. 5d). This pattern is reminiscent of that of Hox gene expression, suggesting that miR-196a may be influenced by regulatory controls imposed on the Hox clusters. But Hox9-group genes, located immediately upstream of miR-196, all have more anterior expression limits than miR-196 (ref. 18), suggesting that miR-196 family members are not regulated simply by control elements shared with the nearest Hox gene.
We did note that the posterior limit of Hoxb8 expression was similar to the anterior limit of miR-196a expression identified by the sensor (Fig. 5e)19. This is notable because we identified, using a new miRNA target prediction program to be described elsewhere, a site in the Hoxb8 3′ UTR containing 21 nucleotides of complementarity (including one G–U base pair) to miR-196a (Fig. 6a). This site is highly conserved across vertebrate species (Fig. 6b).
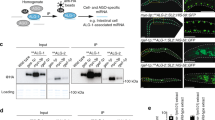
(a) Complementarity between miR-196a and the Hoxb8 3′ UTR was predicted by a new miRNA target prediction program, FASTH (J.S., M.T.M., P.A.S. and M.Z., unpublished data). (b) The miR-196a putative binding site is conserved in Hoxb8 orthologs from multiple species. Asterisks indicate conserved nucleotides. (c) miR-196a downregulates a GFP reporter bearing the miR-196 complementary site from the Hoxb8 3′ UTR in cell culture. (d) Hoxb8 RNA is a target of RISC-mediated RNA cleavage in vivo. RNA was extracted from E11.5 mouse embryos and subjected to a modified form of 5′ RACE. Most (11 of 16) sequenced Hoxb8 clones were cleaved at the expected site. The remaining clones may represent artifacts owing to 5′ exonuclease activity or may be indicative of alternative processing of miR-196a isoforms. Our data does not distinguish between these possibilities.
To determine whether this miR-196 complementary site could function as a target for miR-196 regulation, we cloned it downstream of a green fluorescent protein (GFP) reporter and assayed GFP expression in cultured cells in the presence or absence of miR-196a. Reporter expression was six to ten times lower in the presence of miR-196a, indicating that miR-196a can interact with the Hoxb8 3′ UTR (Fig. 6c).
In all previously reported cases in animals, miRNAs bind their targets with incomplete complementarity, resulting in translational repression; however, it has been shown that the degree of complementarity between the miRNA and its target determines whether the target RNA will be translationally repressed or degraded through the RNAi pathway10,20. The perfect complementarity between miR-196a and the Hoxb8 3′ UTR suggests that miR-196a could mediate degradation rather than translational repression of Hoxb8 mRNA. A recent study using a modified form of 5′ RACE showed this to be the case15. We also tested this possibility using the 5′ RACE assay, which is designed to detect RNA-induced silencing complex (RISC)-mediated RNA cleavage products21,22. We detected Hoxb8 RNA cleavage products in total RNA extracted from E11.5 mouse embryos and found that the transcript was cleaved at the expected site in the region of miR-196a complementarity (Fig. 6d). Our results confirm the previous results15 and indicate that miR-196 directs Hoxb8 cleavage through a small interfering RNA mechanism. The previous study, moreover, used reporter assays to show that mRNA cleavage is the primary mechanism by which miR-196 regulates Hoxb8 gene activity15.
Our results show that a miRNA-responsive transgene can be used to glean detailed information about the expression of various miRNAs during mouse embryonic development. Although our approach relied on transient transgenics, in which each embryo represented an independent insertion of the transgene, our results were reproducible. For example, four of four embryos transgenic with respect to the miR-1 sensor, three of three embryos transgenic with respect to the miR-let-7c sensor and three of four embryos transgenic with respect to the miR-10a sensor showed identical β-gal-negative domains in their hearts, limbs and axes, respectively. The expression patterns of miR-let-7c and miR-let-7e were identical in all four limbs of the embryos analyzed. In addition to being reproducible, within the parameters of our study, this approach was specific. Only two nucleotides separate the miR-let-7c and miR-let-7e sequences, yet their corresponding sensors showed different patterns of β-gal activity. We have not, however, tested whether single-base differences can be differentially detected by our reporter system, or whether all two-base differences can be distinguished by sensors. Nonetheless, the reproducible and distinct patterns uncovered by the miR-let-7c and miR-let-7e sensors showed that miRNAs have a high degree of specificity in vivo and suggested that closely related miRNA family members may have distinct targets.
We found that miR-10a is expressed in a pattern similar to that of the neighboring gene Hoxb4, possibly indicative of similar regulatory mechanisms or functions. We observed a Hox-like expression pattern for miR-196a, with anterior limits that differed from those of its nearest neighbors, possibly suggestive of a different regulatory mechanism. Nevertheless, the expression pattern of miR-196a is the inverse of that of Hoxb8 RNA, and Hoxb8 RNA contains a miR-196a complementary site. In agreement with recent findings15, we found that miR-196a bound to the Hoxb8 3′ UTR in tissue culture and that RISC-mediated Hoxb8 RNA cleavage products could be detected in vivo. Taken together, these data indicate that miR-196a represses Hoxb8 expression in the posterior trunk and tail and predict that the restriction of miR-196a to this posterior region is important for proper embryonic patterning.
Understanding the spatial and temporal expression patterns of vertebrate miRNAs is an important first step towards understanding their functions. Expression data will be particularly crucial for designing misexpression studies in redundant miRNA families, where knockouts might be uninformative. The method we describe is rapid when done in transient transgenics, and it should allow efficient screening of hundreds of miRNAs for those involved in a particular process or organ system of interest.
Methods
Sensor transgene construction.
We constructed sensors from a plasmid containing the lacZ coding sequence with a 5′ nuclear localization signal followed by an SV40 3′ UTR containing a polyadenylation signal (a gift from S. Dymecki; Harvard Medical School). First, we added a PstI site to the 5′ end of the SV40 3′ UTR by PCR, such that the 3′ UTR was flanked by unique PstI and NotI sites. Next, we amplified a second copy of the 3′ UTR by PCR to add a NotI site upstream of the PstI site and a XhoII site at the 3′ end and cloned it downstream of the first copy of the 3′ UTR. We also cloned an annealed oligonucleotide fragment bearing an FRT sequence followed by PacI and FseI sites into the PstI site of the first 3′ UTR and cloned annealed oligonucleotides bearing only an FRT sequence into the second 3′ UTR (oligonucleotide sequences available on request). Finally, we cloned annealed oligonucleotides bearing two copies of a miRNA-complementary sequence, separated by an AscI site, into the PacI and FseI sites in the first copy of the 3′ UTR (oligonucleotide sequences are available on request; for the sequence of each miRNA see the Sanger Institute's miRNA registry).
Although they are not described in this study, we included the second 3′ UTR and FRT sites to allow the miRNA complementary sequence to be deleted in stably transgenic lines after introduction of Flp recombinase. Although all embryos reported in this work were transiently transgenic, and thus any consistent patterns can be attributed to transgene regulation by the cognate miRNA, in stable lines, Flp-mediated deletion should control for position effects associated with individual insertion sites by rescuing ubiquitous lacZ expression. Alternatively, to avoid position effects and the potential difficulty of Flp-mediated recombination in cases where multiple copies of the transgene insert, the sensor construct could be knocked-in to a ubiquitously expressed locus.
Finally, we subcloned each miRNA sensor into pInsCAG (also a gift from S. Dymecki), which contains the ubiquitous CAG promoter (chicken β-actin promoter, human cytomegalovirus enhancer)23 and four copies of a chicken β-globin insulator sequence24, such that the final constructs are flanked by two insulator sequences (the utility of which was not verified for our assay) on each side. Although our analysis focused on E9.5–E10.5 embryos, the CAG promoter drives expression in virtually all tissues during later embryogenesis and in adults25; the sensor transgene should therefore prove useful for analyzing miRNA expression patterns in later stages as well. Differences in levels of miRNA expression between embryos cannot be inferred from these experiments because each transgene insertion site will drive unique levels of lacZ expression.
lacZ staining.
We collected embryos in phosphate-buffered saline (PBS) and fixed them for 30 min at 4 °C in 2% paraformaldehyde and 0.2% gluteraldehyde in PBS. We washed embryos several times in PBS and then three times for 15–30 min each in lacZ wash buffer (100 mM sodium phosphate at pH 7.4, 0.1% sodium deoxycholate, 2 mM MgCl2 and 0.2% Nonidet P-40). We then stained them overnight at 37 °C in lacZ wash buffer containing 5 mM K3Fe(CN)6, 5 mM K4Fe(CN)6 and 1 mg ml−1 X-gal. We then washed embryos several times in PBS and photographed them. For sectioning, we equilibrated stained embryos in 20% sucrose and then embedded them in 7.5% gelatin and 15% sucrose overnight at 4 °C. We froze blocks by submerging them in 2-methylbutane chilled in a dry ice–ethanol bath and stored them at −80 °C until sectioning. We cut cryosections (20 μm each) and photographed them.
Mice.
Mice were housed and handled in accordance with protocols approved by the Institutional Animal Care and Use Committee of Harvard Medical School.
Northern-blot analysis.
We carried out northern-blot analysis as previously described12 or with minor modifications as follows. We extracted total RNA from fresh, homogenized tissue with Trizol reagent (Invitrogen) following the manufacturer's instructions, denatured it at 65 °C for 15 min, loaded 30 μg per lane onto a precast 15% polyacrylamide Tris-borate-EDTA–urea gel (Novex) and separated it using 0.5 × Tris-borate-EDTA as running buffer. We electroblotted RNA to a Hybond N+ membrane (Amersham) at 200 mA overnight at 4 °C, crosslinked it with ultraviolet radiation using a Stratalinker and then baked at 80 °C for 1 h. Blots were prehybridized for at least 4 h at 50 °C in hybridization buffer (5 × saline sodium citrate, 7% SDS, 20 mM sodium phosphate at pH 7.2, 1 × Denhardt solution and 20 μg ml−1 denatured salmon sperm DNA), before overnight incubation at 50 °C in hybridization buffer containing [32P]-end-labeled probe containing a 4 × concatamer of miRNA complementary sequence. We washed the blots twice for 10 min each in wash buffer (3 × saline sodium citrate, 5% SDS, 25 mM sodium phosphate at pH 7.2 and 10 × Denhard's solution) at 50 °C and exposed them to film.
In situ hybridization.
We carried out in situ hybridization as previously described26.
Cell culture assays.
We generated the GFP-Hoxb8 vector by cloning annealed oligonucleotides bearing the miR-196 complementary sequence into the NT-GFP vector (Stratagene; oligonucleotide sequences available on request). miR-196a duplex RNA was manufactured by Dharmacon. We transfected HeLa cells with reporter and control NT-GFP vectors alone or cotransfected them with miR-196a duplex RNA using lipofectamine 2000 (Invitrogen) in accordance with the manufacturer's instructions. We measured GFP fluorescence using a Becton Dickson Facscaliber flow cytometer 36–48 h after transfection.
Cleavage site mapping.
We isolated total RNA from E11.5 embryos with Trizol reagent (Invitrogen) following the manufacturer's instructions. We carried out 5′ RACE with the FirstChoice RLM-RACE kit (Ambion), as described by the manufacturer but omitting both phosphatase steps. After ligation of the adaptor (as described by the manufacturer), we carried out two rounds of PCR using Hoxb8- and adapter-specific primers (sequences are available on request). We cloned RACE products into the TOPO TA vector (Invitrogen) following the manufacturer's instructions and sequenced clones. This modified RACE technique makes use of the fact that, unlike other RNA species, RISC-mediated cleavage products contain a 5′ monophosphate that can be ligated directly to RNA adaptors without enzymatic treatment. The site of substrate cleavage can then be mapped after RT-PCR of cleavage products, and in all reported cases, it corresponds to the tenth nucleotide from the 5′ end of the miRNA21,22.
URL.
The Sanger Institute miRNA registry is available at http://www.sanger.ac.uk/Software/Rfam/mirna/index.shtml.
References
Griffiths-Jones, S. The microRNA Registry. Nucleic Acids Res. 32, D109–D111 (2004).
Lee, R.C. & Ambros, V. An extensive class of small RNAs in Caenorhabditis elegans. Science 294, 862–864 (2001).
Lagos-Quintana, M. et al. Identification of tissue-specific microRNAs from mouse. Curr. Biol. 12, 735–739 (2002).
Lagos-Quintana, M., Rauhut, R., Lendeckel, W. & Tuschl, T. Identification of novel genes coding for small expressed RNAs. Science 294, 853–858 (2001).
Krichevsky, A.M., King, K.S., Donahue, C.P., Khrapko, K. & Kosik, K.S. A microRNA array reveals extensive regulation of microRNAs during brain development. RNA 9, 1274–1281 (2003).
Chen, X. A microRNA as a translational repressor of APETALA2 in Arabidopsis flower development. Science 303, 2022–2025 (2004).
Juarez, M.T., Kui, J.S., Thomas, J., Heller, B.A. & Timmermans, M.C. microRNA-mediated repression of rolled leaf1 specifies maize leaf polarity. Nature 428, 84–88 (2004).
Johnson, S.M., Lin, S.Y. & Slack, F.J. The time of appearance of the C. elegans let-7 microRNA is transcriptionally controlled utilizing a temporal regulatory element in its promoter. Dev. Biol. 259, 364–379 (2003).
Brennecke, J., Hipfner, D.R., Stark, A., Russell, R.B. & Cohen, S.M. bantam encodes a developmentally regulated microRNA that controls cell proliferation and regulates the proapoptotic gene hid in Drosophila. Cell 113, 25–36 (2003).
Hutvágner, G. & Zamore, P.D. A microRNA in a multiple-turnover RNAi enzyme complex. Science 297, 2056–2060 (2002).
Reinhart, B.J. et al. The 21-nucleotide let-7 RNA regulates developmental timing in Caenorhabditis elegans. Nature 403, 901–906 (2000).
Pasquinelli, A.E. et al. Conservation of the sequence and temporal expression of let-7 heterochronic regulatory RNA. Nature 408, 86–89 (2000).
Lagos-Quintana, M., Rauhut, R., Meyer, J., Borkhardt, A. & Tuschl, T. New microRNAs from mouse and human. RNA 9, 175–179 (2003).
Lim, L.P., Glasner, M.E., Yekta, S., Burge, C.B. & Bartel, D.P. Vertebrate microRNA genes. Science 299, 1540 (2003).
Yekta, S., Shih, I.H. & Bartel, D.P. MicroRNA-directed cleavage of HOXB8 mRNA. Science 304, 594–596 (2004).
Whiting, J. et al. Multiple spatially specific enhancers are required to reconstruct the pattern of Hox-2.6 gene expression. Genes Dev. 11, 2048–2059 (1991).
Brend, T., Gilthorpe, J., Summerbell, D. & Rigby, P.W. Multiple levels of transcriptional and post-transcriptional regulation are required to define the domain of Hoxb4 expression. Development 130, 2717–2728 (2003).
Erselius, J.R., Goulding, M.D. & Gruss, P. Structure and expression pattern of the murine Hox-3.2 gene. Development 110, 629–642 (1990).
van den Akker, E. et al. Axial skeletal patterning in mice lacking all paralogous group 8 Hox genes. Development 128, 1911–1921 (2001).
Bartel, D.P. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116, 281–297 (2004).
Llave, C., Xie, Z., Kasschau, K.D. & Carrington, J.C. Cleavage of Scarecrow-like mRNA targets directed by a class of Arabidopsis miRNA. Science 297, 2053–2056 (2002).
Kasschau, K.D. et al. P1/HC–Pro, a viral suppressor of RNA silencing, interferes with Arabidopsis development and miRNA function. Dev. Cell 4, 205–217 (2003).
Niwa, H., Yamamura, K. & Miyazaki, J. Efficient selection for high-expression transfectants with a novel eukaryotic vector. Gene 108, 193–199 (1991).
Chung, J.H., Whiteley, M. & Felsenfeld, G. A 5′ element of the chicken β-globin domain serves as an insulator in human erythroid cells and protects against position effect in Drosophila. Cell 74, 505–514 (1993).
Araki, K., Araki, M., Miyazaki, J. & Vassalli, P. Site-specific recombination of a transgene in fertilized eggs by transient expression of Cre recombinase. Proc. Natl. Acad. Sci. USA 92, 160–164 (1995).
Dietrich, S., Schubert, F.R. & Lumsden, A. Control of dorsoventral pattern in the chick paraxial mesoderm. Development 124, 3895–3908 (1997).
Hamburger, V. & Hamilton, H.L. A series of normal stages in the development of the chick embryo. J. Morphol. 88, 49–92 (1951).
Acknowledgements
We thank S. Dymecki for plasmids; A. Abney and L. Du for pronuclear injections of the transgenes; members of the laboratories of C.J.T., C. Cepko, S. Dymecki and P.A.S. for discussions and advice; and A. Brent for critical reading of the manuscript. This work was supported by a grant from the March of Dimes to C.J.T., and by a US National Institutes of Health MERIT award and grants from the National Cancer Institute to P.A.S. J.H.M. and M.T.M. are supported by postdoctoral fellowships from the National Institutes of Health and the Cancer Research Institute, respectively.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Mansfield, J., Harfe, B., Nissen, R. et al. MicroRNA-responsive 'sensor' transgenes uncover Hox-like and other developmentally regulated patterns of vertebrate microRNA expression. Nat Genet 36, 1079–1083 (2004). https://doi.org/10.1038/ng1421
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng1421
This article is cited by
-
Nr6a1 controls Hox expression dynamics and is a master regulator of vertebrate trunk development
Nature Communications (2022)
-
Molecular features and vulnerabilities of recurrent chordomas
Journal of Experimental & Clinical Cancer Research (2021)
-
The roles of microRNAs in mouse development
Nature Reviews Genetics (2021)
-
MiR-10a and HOXB4 are overexpressed in atypical myeloproliferative neoplasms
BMC Cancer (2018)
-
Transcription and microRNA Profiling of Cultured Human Tympanic Membrane Epidermal Keratinocytes
Journal of the Association for Research in Otolaryngology (2018)