Abstract
Little is known of the molecular mechanisms whereby spermatogonia, mitotic germ cells of the testis, self-renew and differentiate into sperm1,2. Here we show that Zfp145, encoding the transcriptional repressor Plzf, has a crucial role in spermatogenesis. Zfp145 expression was restricted to gonocytes and undifferentiated spermatogonia and was absent in tubules of W/Wv mutants that lack these cells. Mice lacking Zfp145 underwent a progressive loss of spermatogonia with age, associated with increases in apoptosis and subsequent loss of tubule structure but without overt differentiation defects or loss of the supporting Sertoli cells. Spermatogonial transplantation experiments revealed a depletion of spermatogonial stem cells in the adult. Microarray analysis of isolated spermatogonia from Zfp145-null mice before testis degeneration showed alterations in the expression profile of genes associated with spermatogenesis. These results identify Plzf as a spermatogonia-specific transcription factor in the testis that is required to regulate self-renewal and maintenance of the stem cell pool.
Similar content being viewed by others
Main
Zfp145 (also known as Plzf, promyelocytic leukemia zinc-finger) was identified by virtue of its involvement in chromosomal translocations with the gene Rara (retinoic acid receptor-α), which is associated with acute promyelocytic leukemia3. These translocations generate PLZF-RARA and RARA-PLZF fusion proteins, which can interfere with PLZF function3. PLZF exerts local and long-range chromatin remodeling activity through its ability to homodimerize and recruit DNA histone deacetylases and nuclear corepressors4. Plzf is dynamically expressed during embryogenesis5 and has a crucial role in patterning the developing limb and axial skeletal structures6. In addition, Plzf exerts growth-suppressive activities accompanied by accumulation of cells in the G0/G1 compartment of the cell cycle6,7.
Spermatogenesis is a highly organized cyclic process with distinct phases: mitosis, meiosis and spermiogenesis. The mitotic germ cells in the adult testis are the spermatogonia, which originate from primordial germ cells (PGCs) during embryogenesis1,2. In the mouse, the PGCs proliferate and expand in number between embryonic day (E) 8.5 and E13.5. PGCs then enter a cell cycle arrest and become gonocytes in the postnatal testis. Gonocytes resume proliferation around postnatal day (P) 3, giving rise to spermatogonia. Although spermatogenesis represents a classical stem cell model, its complexity and the lack of in vitro culture systems make it difficult to study the molecular aspects underlying spermatogonial stem cell behavior.
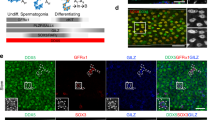
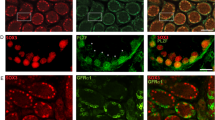
Plzf was expressed in the male gonad during mouse embryogenesis and postnatal life (Fig. 1a). Immunohistochemistry analysis detected Plzf in the testis at E17.5. In the postnatal testis, protein expression peaked at 1 week of age and was localized to the gonocytes and spermatogonia. At this stage of development, when expression of Plzf is highest, gonocytes exit their cell cycle arrest and enter a wave of proliferation. Expression of Plzf decreased through time and was confined to selected spermatogonia (Fig. 1a). Plzf expression was restricted to spermatogonia, as shown by the absence of Plzf staining in W/Wv mutants that lack these cells except in rare tubules8 and by in situ hybridization (Fig. 1a,b). To understand the type of spermatogonia that express Plzf, we immunostained serial sections of testis for Plzf, cyclin D1 (a marker for mitotically active spermatogonia) and cyclin D2, which is expressed as undifferentiated spermatogonia differentiate into A1-type cells9 (Fig. 1c). Expression of Plzf was restricted to spermatogonia that are negative for both cyclin D1 and cyclin D2 and must therefore be largely quiescent and nondifferentiating cells, characteristic of progenitor-type stem cells.
(a) Immuno-histochemistry with an antibody to Plzf on sections from the embryonic genital ridge at E13.5 and E17.5 and on the developing postnatal testis (at 1, 2 and 3 weeks). Expression of Plzf was first detected in the germ cells at E17.5 during development and peaked at around 1 week postnatal. Plzf expression was localized to the spermatogonia and Plzf-positive cells became progressively sparser in the seminiferous tubules with age. Insets show higher magnifications of the basal spermatogonia layer. Tubules from the W/Wv mouse that did not contain spermatogonia lacked expression of Plzf. (b) Bright-field image of in situ hybridization of testis of juvenile (3-week-old) mice with sense (S) and antisense (As) probes to Zfp145. Arrowheads in the antisense panel show localization of signal to selected spermatogonia; the sense control probe produced no specific signal. (c) Serial sections (6 μm) of testis from juvenile (3-week-old) Zfp145+/+ mice immunostained for cyclin D1 and Plzf and for cyclin D2 and Plzf (paired sections). Scale bars, 50 μm. Plzf expression is restricted to spermatogonia negative for both cyclin D1 and cyclin D2 (mitotically inactive and nondifferentiating spermatogonia).
The testes of Zfp145−/− mice were much smaller than those of Zfp145+/+ and Zfp145+/− mice (Fig. 2a). Histological analysis of testis sections from Zfp145−/− mice showed degenerated tubules with absence of the gametes at the basal membrane (Fig. 2b). This progressive lack of spermatogonia in the tubules impaired the spermatogenic process and consequently decreased production of mature spermatozoa (Fig. 2c).
(a) The reproductive tracts from 3-month-old Zfp145+/+, Zfp145+/− and Zfp145−/− male littermates. The testes from Zfp145−/− mice are markedly smaller than those from Zfp145+/+ or Zfp145+/− controls (P < 0.02); attached tissues show no significant differences in size. Graph shows testis mass from sets of Zfp145+/+ and Zfp145−/− adult mice. (b) Sections of testis from adult Zfp145+/+ and Zfp145−/− littermates stained with hematoxylin and eosin to show morphology of the seminiferous tubules. A substantial proportion of tubules in the testis of Zfp145−/− mice had degenerated and no longer contained spermatogonia or differentiating spermatocytes (asterisk). Occasionally, degenerated tubules contained elongated spermatids (arrow). Higher magnifications of the basal spermatogonia layer of the tubules are shown as insets. Graph shows the numbers of degenerating tubules found in Zfp145−/− adult mice (P < 0.05), defined as tubules that lacked spermatogonia. (c) Tubule degeneration in Zfp145−/− mice was associated with reduced production of mature sperm. Hematoxylin and eosin–stained sections through the epididymis of Zfp145+/+ and Zfp145−/− adult mice show reduced levels of sperm, which correlated with decreased sperm counts (right panels), both in terms of total sperm number (P < 0.02) and the numbers of motile, actively progressive sperm (P < 0.02). (d) Immunohistochemistry with antibodies to PCNA (upper panels) and p27 (lower panels) on sections of testis from adult Zfp145+/+ and Zfp145−/− mice. Black arrowheads indicate the supporting Sertoli cells identified either through their distinctive morphology or p27 immunoreactivity. White arrowheads indicate the spermatogonia, which are PCNA-positive. Degenerating tubules in testis of Zfp145−/− mice lack PCNA-positive spermatogonia but retain the Sertoli cells. Scale bars, 50 μm.
Zfp145−/− mice do not have fewer gonocytes, suggesting that there is no defect in the production of PGCs during embryogenesis. To determine whether the gonocytes in testis of Zfp145−/− mice could initiate and complete spermatogenesis, we examined postnatal testis of Zfp145+/+, Zfp145+/− and Zfp145−/− mice. Gonocytes resumed proliferation to become spermatogonia and migrated from the center of the testis cords to the basement membrane of the testis tubules during the first postnatal days in mice of all genotypes (Supplementary Fig. 1 online). Testis of adult Zfp145−/− mice contained tubules with multilayered seminiferous epithelia, including some with mature spermatids lining the lumen (Fig. 2b), making it possible to identify the diverse cell stages (Supplementary Fig. 1 online). This indicates that Zfp145−/− germ cells could initiate and complete the entire spermatogenic process to produce apparently normal sperm. But the epididymal spermatozoa number and sperm counts of Zfp145−/− mice were much lower than those of littermate controls (Fig. 2c). We wanted to determine whether the low numbers of mature sperm produced by Zfp145−/− mice were still functional, but the numbers of viable, actively motile sperm from sets of Zfp145−/− mice were too low for us to carry out a standard in vitro fertilization procedure successfully (data not shown and Fig. 2c).
To further define the progressive loss of spermatogonia and the status of the Sertoli cells (a morphologically distinct cell population that support the spermatogonia), we used proliferating cell nuclear antigen (PCNA) and p27 as distinct markers for the proliferative spermatogonia10 and Sertoli cells11, respectively. Many tubules of Zfp145−/− mice lacked PCNA-positive cells but retained p27-positive Sertoli cells along the basement membrane (Fig. 2d).
In addition to some normal tubules and tubules containing only Sertoli cells, we also observed tubules that contained only elongated spermatids in testis of adult Zfp145−/− mice (Fig. 2b). This indicated that spermatogenesis had occurred in these tubules, leading to production of spermatids, but that earlier stages of subsequent rounds of the cycling seminiferous epithelium were lost.
We also noticed that as the tubules of Zfp145−/− mice degenerated, the surrounding testosterone-synthesizing Leydig cell population was able to expand. This correlated with increased levels of serum testosterone in these mice, ruling out an impairment in the pituitary-testis axis (Supplementary Fig. 2 online)12.
Testis degeneration can be induced by an increase in spermatogonial proliferation, which leads to aberrant entry of the cells into meiosis and subsequent activation of meiotic checkpoints13. This can result in an increase in germ cell apoptosis and a reduction in the proliferative population in the seminiferous tubules with time. We therefore assessed the proliferation of spermatogonia in testis of Zfp145−/− mice by immunostaining for cyclin D1 (ref. 9) and phosphorylated histone H3 and by assessing incorporation of bromodeoxyuridine (BrdU) at different postnatal ages. We observed a significant and progressive decrease in the overall number of proliferative spermatogonia in testis of Zfp145−/− mice, which started at 2 weeks of age and was most notable in the adult, when overt tubule degeneration was readily apparent (Fig. 3a,b and Supplementary Fig. 3 online; data on phosphorylated H3 not shown). This is in complete agreement with the progressive reduction in spermatogonia observed in testis of Zfp145−/− mutants. One might expect adult Zfp145−/− mice to have fewer proliferating cells, as many tubules have lost their spermatogonia, but we observed no obvious degeneration of the tubules until 2 weeks of age, although numbers of proliferating spermatogonia began to decrease. This suggests that the proliferative capacity of spermatogonia was already compromised at this early age. At the first postnatal week, however, even though testis of Zfp145−/− and Zfp145+/+ mice had similar numbers of proliferating cells per tubule (Fig. 3a,b), we observed many fewer inactive tubules in Zfp145−/− mice (Fig. 3c). This suggests that inactivation of Zfp145 causes an initial increased burst of proliferation followed by the rapid exhaustion of the proliferative spermatogonial compartment.
(a) Representative immunohistochemistry staining of testis from Zfp145+/+ and Zfp145−/− mice of various postnatal ages for cyclin D1, a marker of mitotic spermatogonia. Sections were counterstained with hematoxylin. Scale bar, 50 μm. (b) Number of cyclin D1–positive cells per seminiferous tubule in testis of Zfp145+/+ and Zfp145−/− mice of various ages. The mean value is shown with standard error. More than 40 tubules were scored for each of the first three time points and genotypes, whereas all the tubules from three independent microscope fields (>20 tubules) were scored for the 3-week and >12-week (adult) time points. *P < 0.05 for comparison of Zfp145+/+ and Zfp145−/− mice; **P < 0.02. (c) Graph showing expanded results from the scoring of testis from 1-week-old mice for cyclin D1–positive cells. The value from each individual seminiferous tubule is shown in ascending order. The Zfp145+/+ and Zfp145−/− testes have similar numbers of high-activity tubules (>10 cyclin D1–positive cells per tubule) and these tubules have a similar range of activities (graph lines show extensive overlap for the high-activity tubules). The low-activity tubules (<10 cyclin D1–positive cells per tubule) of Zfp145−/− mice had significantly more proliferating cells than the low-activity of Zfp145+/+ tubules (P < 0.02). In addition, there are fewer inactive tubules (0 cyclin D1–positive cells per tubule) in testis of Zfp145−/− mice than in testis of Zfp145+/+ mice. Dotted line represents arbitrary divider of high- and low-activity tubules. (d) Representative immunohistochemistry staining of testis of Zfp145+/+ and Zfp145−/− mice at various ages with an antibody specific for cleaved, activated caspase 3 as a marker of apoptotic cells. Sections were counterstained with hematoxylin. Scale bar, 50 μm.
Increased proliferative activity in early postnatal testes is associated with increased cell death and tubule degeneration at later stages2,13. We therefore scored for apoptosis in testis of Zfp145+/+ and Zfp145−/− mice14. At the first and second postnatal weeks, testis of Zfp145−/− mice had more seminiferous tubules that contained apoptotic cells than did testis of Zfp145+/+ controls (Fig. 3d and Table 1; TUNEL data not shown). This accompanied and followed the observed aberrant proliferative activity of spermatogonia from Zfp145−/− mice during the early postnatal stages.
Because the seminiferous tubules of Zfp145−/− mice suffer a progressive degeneration (Fig. 2b,d) and Zfp145 expression is restricted to undifferentiated spermatogonia (Fig. 1c), we hypothesized that Plzf is required to maintain the germline stem cell compartment. To test this, we transplanted cells from testes of Zfp145+/+ and Zfp145−/− mice into testes of recipient mice that had been chemically depleted of germ cells and assayed their ability to repopulate the tubules15,16,17. Only undifferentiated spermatogonial stem cells are able to repopulate tubules in recipient testis. In mice that received transplants from Zfp145−/− donors, we distinguished the possible residual repopulation from the recipient testis by immunostaining for Plzf. We found that transplanted Zfp145−/− cells were substantially less efficient than control cells at repopulation (Fig. 4a,b). In addition, the limited numbers of repopulated tubules from Zfp145−/− cell transplantation contained Plzf-positive spermatogonia (Fig. 4c) and were therefore derived from the low numbers of remaining recipient undifferentiated spermatogonia. Therefore, loss of Zfp145 leads to depletion of the germline stem cells in testis.
(a) Sections of recipient testis after transplantation with spermatogonia from testis of 3–4-month-old Zfp145+/+ and Zfp145−/− mice were stained with hematoxylin and eosin to show repopulated (white asterisk) and remaining depleted (black arrowhead) seminiferous tubules. (b) Graph showing percentages of tubules in recipient testis judged as having restored spermatogenesis. Individual counts from each testis are indicated with triangles; the mean of each data set is plotted with a horizontal bar. At least 200 tubules were counted for each testis. Eight and ten testes transplanted from Zfp145+/+ and Zfp145−/− cells, respectively, were analyzed. The numbers of repopulated tubules from the Zfp145+/+ transplantation was significantly higher than that from the Zfp145−/− (P < 0.05). (c) Immunostaining of repopulated tubules for Plzf. In testis repopulated from Zfp145−/− mutants, the residual spermatogenesis derives entirely from the recipient (Plzf-positive) spermatogonia. Scale bars, 50 μm. Spermatogonia staining positive for Plzf are indicated with black arrowheads.
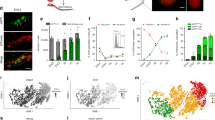
Microarray analysis of spermatogonia isolated by flow cytometry from Zfp145−/− mice before testis degeneration showed alterations in its transcriptomic profile (Fig. 5a and Supplementary Tables 1–3 online). Expression of genes directly regulated by Plzf in other model systems (e.g., Ccna2 (ref. 18), Myc19) was not altered, but we found several genes whose expression was perturbed. Among them were genes involved in metabolism (Gpd1, Cyp11a1, Hsd17b1, Ash2l), RNA binding (Rbm5, Rbm9, Paip1, Pabpc1), cell cycle control (Ccnd2, Ches1), cytoskeletal and cell-junction components (Knsl7, Col4a3bp, Cldn11) and transcription factors (Dmrt2; Fig. 5b). We verified a selection of gene changes by semiquantitative RT-PCR on whole-testis RNA from Zfp145+/+ and Zfp145−/− mice and confirmed that expression of Myc and Ccna2 was not altered (Fig. 5c). Expression of Ccna2 was confirmed to be unaltered by quantitative RT-PCR (data not shown). Several genes aberrantly expressed in the testes of Zfp145−/− mice may have roles in spermatogenesis9,20,21. For instance, RNA-binding proteins are implicated in the regulation of mRNA splicing and timing of translation during spermatogenesis and are affected in some cases of azoospermia in humans. Ches1 has a role in regulating the G2/M checkpoint, important in controlling mitosis and meiosis22, and another set of genes (e.g., Ccnd2 (ref. 9), Morc23 and Tsx24) are implicated in spermatogonia differentiation. Deregulated expression of these genes probably disrupts the tight balance between spermatogonial self-renewal and differentiation. Looking for Plzf binding sites in the promoter regions of these genes, we found only two putative Plzf responsive elements in Tsx, which is downregulated in testis of Zfp145−/− mice. Tsx and the other genes are therefore probably indirect Plzf targets. Alternatively, Tsx is in the restricted array of target genes directly transactivated by Plzf25.
(a) Spermatogonia were sorted by flow cytometry from dispersed testes of 1-week-old Zfp145+/+ and Zfp145−/− mice by virtue of low side scatter (SCC) and high α6-integrin (CD49f; red box). Sorted spermatogonia were pooled from several mice and RNA was extracted for microarray analysis of gene expression. (b) Results were visualized by GeneSpring v6.0 and a selection of genes showing significantly altered expression between Zfp145+/+ and Zfp145−/− spermatogonia are shown. (c) Expression of selected genes identified in the microarray comparison (Ccnd2, Morc, Tsx) and previously suggested targets of Plzf transcriptional repression (Myc, Ccna2) were analyzed by semiquantitative RT-PCR in RNA extracted from the testes of three Zfp145+/+ and three Zfp145−/− 1-week-old mice. (d) Role of Plzf in spermatogenesis. Scheme representing the time line of testis development and production of mature spermatozoa. Red bar in lower panel shows expression levels of Zfp145 over this developmental process. Vertical orange bar represents the stage of spermatogonia collected for the microarray and the point at which the Zfp145−/− testis shows alterations in proliferation and apoptosis before degeneration of the seminiferous tubules.
This study shows that Plzf has an essential role in spermatogonia maintenance (Fig. 5d): its inactivation results in age-dependent germ cell loss leading to testicular degeneration due to a progressive reduction of self-renewal capability of the spermatogonial stem cell compartment. Our findings are in agreement with the observations that a Zfp145-inactivating mutation is found in the luxoid mutant, a mouse strain that has impaired spermatogenesis and progressive loss of spermatogonia26.
Based on our data, it is conceivable that inactivation of Zfp145 leads to unrestricted exit from quiescence of spermatogonia, ensuing inappropriate activation of meiotic checkpoints and increased apoptosis. As these defects closely mimic some characteristics of human male infertility, Zfp145−/− mutants may be a new animal model for such disorders27. Furthermore, in view of the specific biochemical role of Plzf, these data implicate chromatin remodeling and transcriptional regulation in the control of stem cell maintenance in the germ cell compartment. The ability of Plzf to regulate molecular pathways underlying stem cell maintenance may be perturbed in acute promyelocytic leukemia, thus lending leukemic hemopoietic progenitors a proliferative advantage.
Methods
Mice and sperm analysis.
We inactivated Zfp145 in mice as previously described6. All mice were housed at the Memorial Sloan Kettering Cancer Center animal facility in compliance with National Institutes of Health guidelines on the use of laboratory and experimental animals. We carried out sperm counts by microscopic analysis of sperm extracted from the epididymis and vas deferens of sets of Zfp145+/+ and Zfp145−/− adult mice (6 months old). Sperm counts and in vitro fertilization procedures were done at the Memorial Sloan Kettering Transgenic Facility.
Histology and immunohistochemistry.
We fixed testes in Bouin's or formalin fixative solutions and embedded them in paraffin. We cut 8-μm sections, removed the wax from them and stained them with periodic acid Schiff–hematoxylin (Polyscientific) or with hematoxylin and eosin. For immunohistochemistry, we fixed these sections in ice-cold acetone and quenched endogenous peroxidase activity by incubating them with 0.1% hydrogen peroxide. We subjected the sections to antigen retrieval by microwaving in Antigen Unmasking Solution (Vector Laboratories) or Citra Plus (Biogenex) and incubated them overnight at 4 °C with primary antibody. We used antibodies to Plzf (provided by P.G. Pelicci; Department of Experimental Oncology, European Institute of Oncology and FIRC Institute of Molecular Oncology, Milan, Italy), PCNA (clone PC10), p27 (rabbit polyclonal), cyclin D1 (polyclonal), cyclin D2 (polyclonal; Santa Cruz Biotechnology), p27 (monoclonal; BD Transduction Laboratories), activated caspase-3 and phosphorylated H3 (Cell Signaling Technology). We carried out staining with mouse monoclonal antibody using a MOM kit (Vector Laboratories). We washed sections three times in phosphate-buffered saline and then incubated them with the appropriate biotinylated secondary antibody (DAKO and Vector Laboratories) followed by StreptABComplex/HRP (DAKO and Vector Laboratories). We then developed sections using diaminobenzidine and counterstained them with hematoxylin.
For BrdU staining, we injected mice intraperitoneally with 100 mg of BrdU (Sigma) per kg body weight. After 2 h, we dissected, fixed and processed testes in paraffin. We detected BrdU incorporation with monoclonal antibody to BrdU (Roche), incubated samples with biotinylated secondary antibody and streptavidin-HRP and then detected with diaminobenzidine. We carried out staining for activated caspase-3, phosphorylated H3 and BrdU using the automated staining processor Discovery from Ventana Medical Systems.
In situ mRNA hybridization.
We incubated paraffin-embedded testis sections (8 μm) with sense and antisense riboprobes to Zfp145 radioactively labeled with 33P-UTP and hydrolyzed them in alkali as described28. We carried out autoradiography for 2 weeks and counterstained the sections with hematoxylin and eosin.
Testis transplantation.
The spermatogonial transplantation technique allows the study of germline stem cells of the testis15,16. We obtained donor testis cell populations from 3–4-month-old Zfp145+/+ and Zfp145−/− mice and transplanted them into testis of busulphan-depleted recipient nude mice17 (busulphan-treated testes are almost completely devoid of endogenous germ cells). Three months after the transplantation procedure, we collected testes and processed them for hematoxylin and eosin staining and immunohistochemistry. The numbers of repopulated tubules give an indication of the numbers of germline stem cells in the donor cell population, as only the nondifferentiated spermatogonia are able to persist and repopulate the recipient seminiferous tubules. To quantify the percentage of tubules repopulated by the donor, we analyzed sections at three distinct points throughout each testis.
Testosterone assay.
We extracted blood from male adult Zfp145+/+ and Zfp145−/− mice (<6 months old) during the evening when mice are most sexually active. We than assayed the isolated serum for testosterone levels by a specific Enzyme Immunoassay kit (Diagnostic Systems Laboratories).
Cell isolation and RNA preparation.
We obtained single-cell suspensions from mice of the different genotypes at the first postnatal week and prepared them by enzymatic digestion. We pooled dissociated testis cells from more than six mice of each genotype and stained them with primary antibody to α6-integrin (GoH3; BD PharMingen) conjugated to R-phycoerythrin as previously described29. We sorted cells using a FACSVAntage flow cytometer (BD Biosciences). We isolated total RNA with a TRIZOL kit (Life Technology) from cells sorted for high levels of α6-integrin and low levels of side scatter (spermatogonia) and precleaned them with an RNeasy kit (Quiagen) before microarray analysis.
DNA microarray analysis.
We carried out total RNA quality control before labeling. We analyzed 25–50 ng of total RNA on a RNA 6000 Nano Assay (Agilent) using a Bioanlyzer 2100. We then labeled 2 μg of good-quality total RNA according to protocols recommended by manufacturers. After reverse-transcription with an oligo-dT-T7 (Genset), we generated double-stranded cDNA with the Supercript double-stranded cDNA synthesis custom kit (Invitrogen Life Technologies). In an in vitro transcription step (with T7 RNA polymerase (MessageAmp aRNA kit from Ambion), we linearly amplified the cDNA and labeled it with biotinylated nucleotides (Enzo Diagnostics). We then hybridized 10 μg of labeled and fragmented cRNA onto MGU74v2 A/B/C GeneChips (Affymetrix) and processed them as previously described30. We quantified the image using MAS 5.1 (MicroArray Suite, Affymetrix) with the default parameters for the statistical algorithm and all probe set scaling with a target intensity of 500. We then filtered the data so that the absolute value of the relative change was ≥1.5. We also removed genes that were scored as increasing and scored as absent in the numerator (i.e., the experiment in the ratio whose value is the numerator) and genes that were scored as decreasing and scored as absent in the denominator (also referred to as the base experiment).
RT-PCR analysis.
We isolated total RNA from the testis of 1-week-old Zfp145+/+ and Zfp145−/− mice and used it to synthesize first-strand cDNA using a Superscript III first-strand cDNA synthesis kit and random hexamer priming (Invitrogen). We then used cDNA for PCR using Platinum PCR Supermix (Invitrogen) and appropriate primers. Primer sequences were designed in different exons of the target genes to prevent productive PCR from genomic DNA contamination (sequences are available on request). We isolated PCR products from a range of different cycle numbers (25, 30 and 35 cycles) to ensure reactions were analyzed during the exponential phase. We separated PCR products on 1.5% agarose gels. We carried out quantitative RT-PCR directly from testis total RNA extracts with a LightCycler (Roche) and SYBR-Green RT-PCR kit (Qiagen) according to manufacturers' instructions. We normalized semiquantitative and quantitative RT-PCR reactions against levels of the transcript for a housekeeping gene (Hprt).
GEO accession number.
Note: Supplementary information is available on the Nature Genetics website.
Accession codes
References
McLaren, A. Germ and somatic cell lineages in the developing gonad. Mol. Cell. Endocrinol. 163, 3–9 (2000).
de Rooij, D.G. Proliferation and differentiation of spermatogonial stem cells. Reproduction 121, 347–354 (2001).
Costoya, J.A. & Pandolfi P.P. The role of promyelocytic leukemia zinc finger and promyelocytic leukemia in leukemogenesis and development. Curr. Opin. Hematol. 8, 212–217 (2001).
Barna, M. et al. Plzf mediates transcriptional repression of HoxD gene expression through chromatin remodeling. Dev. Cell. 3, 499–510 (2002).
Cook, M. et al. Expression of the zinc-finger gene PLZF at rhombomere boundaries in the vertebrate hindbrain. Proc. Natl. Acad. Sci. USA 92, 2249–2253 (1995).
Barna, M., Hawe, N., Niswander, L. & Pandolfi, P.P. Plzf regulates limb and axial skeletal patterning. Nat. Genet. 25, 166–172 (2000).
Shaknovich, R. et al. The promyelocytic leukemia zinc finger protein affects myeloid cell growth, differentiation, and apoptosis. Mol. Cell. Biol. 18, 5533–5545 (1998).
Coulombre, J.L. & Russell, E.S. Analysis of the pleiotropism at the W locus in the mouse: The effects of W and Wv substitution upon postnatal development of germ cells. J. Exp. Zool. 126, 277–296 (1954).
Beumer, T.L., Roepers-Gajadien, H.L., Gademan, I.S., Kal, H.B. & de Rooij, D.G. Involvement of the D-type cyclins in germ cell proliferation and differentiation in the mouse. Biol. Reprod. 63, 1893–1898 (2000).
Wrobel, K.H., Bickel, D. & Kujat, R. Immunohistochemical study of seminiferous epithelium in adult bovine testis using monoclonal antibodies against Ki-67 protein and proliferating cell nuclear antigen (PCNA). Cell Tissue Res. 283, 191–201 (1996).
Beumer, T.L. et al. Regulatory role of p27kip1 in the mouse and human testis. Endocrinology. 140, 1834–1840 (1999).
Narula, A. et al. Smad4 overexpression causes germ cell ablation and leydig cell hyperplasia in transgenic mice. Am. J. Pathol. 161, 1723–1734 (2002).
Handel, M.A., Cobb, J. & Eaker, S. What are the spermatocyte's requirements for successful meiotic division? J. Exp. Zool. 285, 243–250 (1999).
Gown, A.M. & Willingham, M.C. Improved detection of apoptotic cells in archival paraffin sections: immunohistochemistry using antibodies to cleaved caspase 3. J. Histochem. Cytochem. 50, 449–454 (2002).
Brinster, R.L. & Zimmermann, J.W. Spermatogenesis following male germ-cell transplantation. Proc. Natl. Acad. Sci. USA 91, 11298–11302 (1994).
Brinster, R.L. & Avarbock, M.R. Germline transmission of donor haplotype following spermatogonial transplantation. Proc. Natl. Acad. Sci. USA 91, 11303–11307 (1994).
Orwig, K.E., Shinohara, T., Avarbock, M.R. & Brinster, R.L. Functional analysis of stem cells in the adult rat testis. Biol. Reprod. 66, 944–949 (2002).
Yeyati, P.L. et al. Leukemia translocation protein PLZF inhibits cell growth and expression of cyclin A. Oncogene 18, 925–934 (1999).
McConnell, M.J. et al. Growth suppression by Acute Promyelocytic Leukemia-Associated protein PLZF is mediated by repression of c-myc expression. Mol. Cell. Biol. 23, 9375–9388 (2003).
Venables, J.P. & Eperon, I. The roles of RNA-binding proteins in spermatogenesis and male infertility. Curr. Opin. Genet. 9, 346–354 (1999).
Andersson, S. Molecular genetics of androgenic 17 beta-hydroxysteroid dehydrogenases. J. Steroid. Biochem. Mol. Biol. 55, 533–534 (1995).
Scott, K.L. & Plon, S.E. Loss of Sin3/Rpd3 histone deacetylase restores the DNA damage response in checkpoint-deficient strains of Saccharomyces cerevisiae. Mol. Cell. Biol. 23, 4522–4531 (2003).
Inoue, N. et al. New gene family defined by MORC, a nuclear protein required for mouse spermatogenesis. Hum. Mol. Genet. 8, 1201–1207 (1999).
Cunningham, D.B., Segretain, D., Arnaud, D., Rogner, U.C. & Avner, P. The mouse Tsx gene is expressed in Sertoli cells of the adult testis and transiently in premeiotic germ cells during puberty. Dev. Biol. 204, 345–360 (1998).
Labbaye, C. et al. PLZF induces megakaryocytic development, activates Tpo receptor expression and interacts with GATA1 protein. Oncogene 21, 6669–6679 (2002).
Buaas, F.W. et al. ZFP145 is required in adult male germ cells for stem cell self-renewal. Nat. Genet. advance online publication, 23 May 2004 (doi:10.1038/ng1366).
Cooke, H.J. & Saunders, P.T. Mouse models of male infertility. Nat. Rev. Genet. 3, 790–801 (2002).
Manova, K., Nocka, K., Besmer, P. & Bachvarova, R.F. Gonadal expression of c-kit encoded at the W locus of the mouse. Development. 110, 1057–1069 (1990).
Shinohara, T., Orwig, K.E., Avarbock, M.R. & Brinster, R.L. Spermatogonial stem cell enrichment by multiparameter selection of mouse testis cells. Proc. Natl. Acad. Sci. USA 97, 8346–8351 (2000).
Rajasekhar, V.K. et al. Oncogenic Ras and Akt signaling contribute to glioblastoma formation by differential recruitment of existing mRNAs to polysomes. Mol. Cell. 12, 889–901 (2003).
Acknowledgements
We thank A. Viale for advice and discussion regarding microarray analysis; the Molecular Cytology, Flow-Cytometry, Transgenic and Genomic Core facilities, particularly C. Farrell and R. Lester, for technical assistance; E. Pikarsky and X. Wang for providing material; and the members of the laboratory of M.A.D. -B. and P.E. Cohen for discussion. This work is supported by National Center for Research Resources grant, grants from the US National Institutes of Health (to K.E.O. and M.S. and to D.J.W.) and a grant from the National Cancer Institute (to P.P.P.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Costoya, J., Hobbs, R., Barna, M. et al. Essential role of Plzf in maintenance of spermatogonial stem cells. Nat Genet 36, 653–659 (2004). https://doi.org/10.1038/ng1367
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng1367
This article is cited by
-
Postnatal development of mouse spermatogonial stem cells as determined by immunophenotype, regenerative capacity, and long-term culture-initiating ability: a model for practical applications
Scientific Reports (2024)
-
Hyaluronic acid/silicon nanoparticle scaffold induces proliferation and differentiation of mouse spermatogonial stem cells transplanted to epididymal adipose tissue
Cell and Tissue Banking (2024)
-
Bud31-mediated alternative splicing is required for spermatogonial stem cell self-renewal and differentiation
Cell Death & Differentiation (2023)
-
Induction of promyelocytic leukemia zinc finger protein by miR-200c-3p restores sensitivity to anti-androgen therapy in androgen-refractory prostate cancer and inhibits the cancer progression via down-regulation of integrin α3β4
Cellular Oncology (2023)
-
E-cadherin maintains the undifferentiated state of mouse spermatogonial progenitor cells via β-catenin
Cell & Bioscience (2022)