Abstract
Telomere length in humans is partly controlled by a feedback mechanism in which telomere elongation by telomerase is limited by the accumulation of the TRF1 complex at chromosome ends1,2,3,4,5. TRF1 itself can be inhibited by the poly(ADP-ribose) polymerase (PARP) activity of its interacting partner tankyrase 1, which abolishes its DNA binding activity in vitro and removes the TRF1 complex from telomeres in vivo. Here we report that the inhibition of TRF1 by tankyrase is in turn controlled by a second TRF1-interacting factor, TIN2 (ref. 6). Partial knockdown of TIN2 by small hairpin RNA in a telomerase-positive cell line resulted in telomere elongation, which is typical of reduced TRF1 function. Transient inhibition of TIN2 with small interfering RNA led to diminished telomeric TRF1 signals. This effect could be reversed with the PARP inhibitor 3-aminobenzamide and did not occur in cells overexpressing a PARP-dead mutant of tankyrase 1. TIN2 formed a ternary complex with TRF1 and tankyrase 1 and stabilized their interaction, an effect also observed with the PARP-dead mutant of tankyrase 1. In vitro, TIN2 protected TRF1 from poly(ADP-ribosyl)ation by tankyrase 1 without affecting tankyrase 1 automodification. These data identify TIN2 as a PARP modulator in the TRF1 complex and can explain how TIN2 contributes to the regulation of telomere length.
Similar content being viewed by others
Main
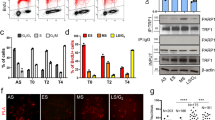
To determine the role of TIN2 in telomere length control, we used small hairpin RNA (shRNA) expressed from a pBabe-based retrovirus7 to create cell populations with diminished TIN2 levels (Fig. 1). Of eight shRNAs, two (shRNA2 and shRNA3) substantially reduced TIN2 levels as determined by quantitative immunoblotting (30% and 67% reduction, respectively; Fig. 1b). We used two shRNAs (shRNA1 and shRNA4) that did not affect TIN2 expression as controls. We determined the effect of lowered TIN2 levels on telomere dynamics using the telomerase-positive HTC75 cell line, a subclone of the human fibrosarcoma line HT1080, which has been used extensively to study telomere length2. Relative to the control (shRNA1), TIN2 shRNA3 and, to a lesser extent, shRNA2 (which has a more modest effect on TIN2 level) caused progressive telomere elongation (Fig. 1c,d). These findings establish TIN2 as a negative regulator of telomere length and are in accordance with a previous study of a TIN2 truncation mutant (TIN2-13)6 that suggested a role for TIN2 in controlling telomere length.
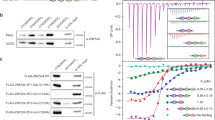
(a) Schematic of the components of the TRF1 telomere length regulation complex. Myb, Myb-related DNA-binding motif; TBD, TRF1-binding domain; TRFH, TRF homology (dimerization) domain; PARP, poly(ADP-ribose) polymerase catalytic domain. The positions of shRNAs and siRNAs that affect TIN2 levels are indicated by asterisks. The positions of the mutations that abolish PARP activity in tankyrase 1-PD are shown below the schematic. (b) Western blot showing TIN2 levels before and after knockdown by shRNAs. HTC75 cells were transduced by retroviruses carrying sequences encoding shRNAs against TIN2, and total cellular proteins were analyzed by immunoblotting using antibodies to TIN2 (864) and to α-tubulin. The TIN2 levels were determined from two independent blots using α-tubulin as a loading control. (c) Genomic blots of telomeric restriction fragments in three HTC75 cell lines infected with the indicated TIN2 shRNA viruses. Genomic DNAs were isolated at the indicated population doublings (PDs), digested with AluI and MboI and analyzed by Southern blotting using a double-stranded TTAGGG repeat probe. (d) Telomere length changes in the shRNA TIN2 knockdown cell lines. shRNA1 and shRNA4 do not affect TIN2 levels.
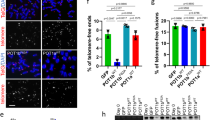
Because the effect of diminished TIN2 levels on telomere length resembles that of TRF1 inhibition2, we examined the effect of TIN2 depletion on TRF1. Compared with the shRNA experiments, transient transfection of TIN2 small interfering RNA (siRNA) into HTC75 or HeLa cells lowered the TIN2 levels further, allowing better analysis of the effect of TIN2 depletion on TRF1 (Fig. 2). Immunofluorescence analysis showed that a large fraction of the cells treated with TIN2 siRNA lacked the typical punctate nuclear staining of TIN2 on telomeres (Fig. 2a). This depletion of TIN2 led to a concomitant reduction in the TRF1 signal at telomeres (Fig. 2a), even though genomic blotting indicated that there was no detectable loss of the duplex telomeric TTAGGG repeat array to which TRF1 binds (data not shown). Eventually, these low TIN2 levels seemed to inhibit cell growth (data not shown), a phenotype that might be related to the lethal phenotype of TRF1 gene targeting in mice8,9. The loss of the telomeric TRF1 signals in TIN2-depleted cells was accompanied by a reduction in the amount of TRF1 associated with chromatin, which can be released by treating nuclei with 420 mM KCl (ref. 10). In both HeLa cells and HTC75 cells, the abundance of chromatin-associated TRF1 was diminished on depletion of TIN2 (Fig. 2b). Furthermore, there was a slight reduction in total TRF1 protein levels when TIN2 was knocked down (Fig. 2b). By contrast, TIN2 siRNA had little or no effect on the chromatin association of TRF2 and hRap1 or the total amount of these proteins (Fig. 2b). A second TIN2 siRNA oligo set had the same effects on TRF1 immunofluorescence signals and protein levels (data not shown); by contrast, a control siRNA directed to green fluorescent protein (GFP) did not affect TRF1 (Fig. 2a,b).
(a) Immunofluorescence analysis showed loss of telomeric TRF1 signal after TIN2 siRNA. HeLaI.2.11 cells were transfected with siRNAs targeting GFP or TIN2, as indicated, and analyzed by indirect immunofluorescence. Primary antibodies used were rabbit polyclonal antibody to TIN2 (865; green) and mouse polyclonal antibody to TRF1 (red). DNA was stained with DAPI (blue) and merged with green and red channels. (b) Western blot showing the reduction of TRF1 and TIN2 protein levels after TIN2 knockdown by siRNA. HeLa1.2.11 and HTC75 cells transfected with GFP or TIN2 siRNA were extracted with buffer containing 150 mM KCl and then with buffer containing 420 mM KCl. The pellets were sonicated in Laemmli buffer. Whole-cell extracts (WCE) were reconstituted by mixing the three fractions proportionally. Antibodies used in western blotting were rabbit polyclonal antibody to TIN2 (864), rabbit polyclonal antibody to TRF1 (371), rabbit polyclonal antibody to TRF2 (647), rabbit polyclonal antibody to hRap1 (765) and mouse monoclonal antibody to γ-tubulin (GTU88). (c) 3-aminobenzamide blocked loss of telomeric TRF1 after knockdown of TIN2 by siRNA. HeLa cells were transfected with TIN2 siRNA in the presence of either DMSO (carrier control; top panels) or 5 mM 3-aminobenzamide (bottom panels). Immunofluorescence was analyzed as in a. (d) Tankyrase-1-PD overexpression rescued loss of TRF1 from telomeres after TIN2 depletion by siRNA. HeLa1.2.11 cells expressing retroviral FN-tankyrase 1-PD were transfected with TIN2 siRNA and immunofluorescence was analyzed as in a.
The effects of TIN2 depletion on TRF1 and telomere length resemble the phenotypes of forced overexpression of tankyrase 1 or 2 (refs. 11,12), two nearly identical PARPs that bind to TRF1 and each other11,12,13,14. The tankyrases have been implicated in controlling telomere length and in numerous other cellular processes15,16. Nuclear overexpression of tankyrases leads to reduced telomeric TRF1 signals, diminished TRF1 protein levels and telomere elongation11,17. Although these effects are thought to involve the modification of TRF1, ADP-ribosylated TRF1 has not been detected in vivo, possibly owing to its rapid degradation18. To test whether the effect of TIN2 depletion was also mediated by tankyrase activity, we used 3-aminobenzamide, a general PARP inhibitor that blocks both automodification of tankyrases and ADP-ribosylation of TRF1 (ref. 12). Treatment with 3-aminobenzamide did not affect RNAi depletion of TIN2, leading to the same reduction of TIN2 levels as in cells treated with dimethylsulfoxide (DMSO; data not shown). 3-aminobenzamide abolished the effect of TIN2 siRNA on the telomeric TRF1 signals, however, whereas control cells again showed reduced TRF1 signals on depletion of TIN2 (Fig. 2c). Nuclear overexpression of an inactive form of tankyrase 1, FN-tankyrase 1-PD, also reverted the TIN2 siRNA effect on TRF1. This form of tankyrase 1 was mutated at positions His1164 and Glu1291 in the PARP domain (see Fig. 1a), abolishing all PARP activity11,19. This mutant protein contains an N-terminal FLAG tag and a nuclear localization signal (NLS, indicated by FN-), ensuring that it accumulates in the nucleus rather than at one of the many other subcellular locations of tankyrases15,16. Consistent with the 3-aminobenzamide result, TRF1 persisted on telomeres in cells expressing FN-tankyrase 1-PD, even when TIN2 levels were severely reduced (Fig. 2d).
Because these data suggested that TIN2 affects the interplay between tankyrase 1 and TRF1, we investigated whether these factors form a ternary complex. Coimmunoprecipitation experiments showed that endogenous TRF1 interacted with TIN2 (Fig. 3a), in agreement with previous reports based on overexpressed proteins6. The TIN2 and TRF1 immunoprecipitates also contained endogenous tankyrase 1 (Fig. 3a), suggesting that these three proteins could form a ternary complex in vivo. To examine this possibility more closely, we carried out coimmunoprecipitations of transfected tagged versions of tankyrase 1 (FN-tankyrase 1), TIN2 and TRF1. TRF1 complexes were precipitated from transiently transfected 293T cells using antibodies against the epitopes used to tag tankyrase 1 and TIN2 (Fig. 3b). The coimmunoprecipitation of tankyrase 1 and TIN2 depended on TRF1, suggesting that TIN2 does not bind tankyrase 1 directly and that TRF1 bridges their interaction. Notably, the interaction between TRF1 and TIN2 was enhanced by cotransfection of tankyrase 1 (Fig. 3b), indicative of mutually reinforcing interactions.
(a) Coimmunoprecipitation of endogenous TRF1 and TIN2 with tankyrase 1. TRF1 complexes precipitated from HeLa cells using antibodies to TRF1 (371) or to TIN2 (864) were analyzed by western blotting. Protein G–conjugated Sepharose beads were used as negative control. (b) Coimmunoprecipitations from transfected 293T cells. FLAG-tagged tankyrase 1, HA-tagged TIN2 and TRF1 were transiently transfected into 293T cells in the combinations shown. Whole-cell extracts (input) were immunoprecipitated (IP) using an antibody to FLAG (M2) or to HA (12CA5). (c) Experimental strategy of sequential coimmunoprecipitation and elution of the ternary TRF1 complex. Two equal aliquots volume were subjected to the entire sequential immunoprecipitation, one of which contained a blocking peptide (FLAG) during the primary immunoprecipitation using antibody to FLAG. WCE, whole-cell extracts; IP, immunoprecipitation. (d) Results of the sequential immunoprecipitation (IP). Protein eluant from the primary immunoprecipitation and the two secondary immunoprecipitations were analyzed using antibodies to tankyrase 1 (465), TRF1 (371) or TIN2 (864). (e) TRF1–tankyrase 1 complex is stabilized by TIN2. 293T cells were transfected with fixed amounts of TRF1 and FN-tankyrase 1 plasmids and increasing amounts of TIN2 plasmid (as indicated). Immunoprecipitations (IP) were done in duplicate using antibody to FLAG (M2) for FN-tankyrase 1 and immunoblotted. (f) PARP-dead alleles of tankyrase 1 coimmunoprecipitated more efficiently with TRF1. 293T cells were cotransfected with plasmids encoding TRF1 and tankyrase 1 constructs as indicated. Complexes were immunoprecipitated (IP) using antibody to FLAG for FLAG-tagged tankyrase 1 proteins with either functional NLS (FN-tankyrase1 alleles) or nonfunctional NLS (FN*-tankyrase 1 alleles). Antibodies to FLAG or MYC were used for western blotting. Left columns (Input) show the protein levels in 293T cell lysates.
We formally established the occurrence of a TIN2–TRF1–tankyrase 1 triple complex by a sequential coimmunoprecipitation and epitope elution protocol (Fig. 3c,d). We first bound protein complexes formed in 293T cells expressing FN-tankyrase 1, MYC-TRF1 and hemagglutinin (HA)-TIN2 to beads conjugated with antibody to FLAG, resulting in coimmunoprecipitation of all three proteins. We carried out a parallel immunoprecipitation experiment including the FLAG peptide, which showed that the recovery of all three proteins depended on the interaction of FN-tankyrase 1 with the FLAG antibody. To confirm the existence of a complex containing tankyrase 1 associated with both TRF1 and TIN2, we eluted the FLAG immunoprecipitate with the FLAG peptide and immunoprecipitated the eluate a second time with an antibody to either MYC or HA (Fig. 3c,d). All three proteins were present in the two secondary immunoprecipitates (Fig. 3d), indicating that TRF1, TIN2 and tankyrase 1 could form a stable ternary complex in vivo. Because tankyrase 1 is much more difficult to detect on telomeres than TRF1 (refs. 12,20) and seems to be a minor component in immunoprecipitates of the endogenous TRF1 (Fig. 3a), tankyrase 1 is probably much less abundant on telomeres than TRF1 and TIN2.
Coimmunoprecipitation experiments showed that the interaction between TRF1 and tankyrase 1 was affected by TIN2. As the amount of TIN2 increased, more TRF1 could be brought down with tankyrase 1 (Fig. 3e). We observed a similarly improved interaction between TRF1 and tankyrase 1 when we carried out coimmunoprecipitations between the tankyrase 1-PD mutant and TRF1 in the absence of TIN2 overexpression (Fig. 3f). The two-point mutations in the PARP domain of tankyrase1-PD probably do not have a direct effect on TRF1 binding, as TRF1 interacts with the ankyrin-repeat domain12. These findings suggest that the PARP activity of tankyrase 1 diminishes its interaction with TRF1 and that TIN2 reverses this effect.
Because TIN2 seemed to counteract the effect of tankyrase 1 on TRF1, we examined the effect of TIN2 on the in vitro activities of this enzyme. Tankyrase 1 poly(ADP-ribosyl)ates TRF1 and itself in vitro, and this reaction can be monitored through the incorporation of labeled ADP-ribose derived from the 32P-β-NAD+ precursor12 (Fig. 4). Using relevant proteins purified from baculovirus-infected insect cells, we observed that TIN2 was not a substrate for tankyrase 1, even when TRF1 was present. But TIN2 had an obvious inhibitory effect on the ability of tankyrase 1 to modify TRF1. Quantitative analysis indicated that the addition of TIN2 (at 2:1 molar ratio of TIN2:TRF1) reduced the modification of TRF1 by a factor of 6.5 (± 0.1; n = 3). We observed this inhibition with two independent TIN2 isolates, and it seemed to be highly specific, as hRap1 and a TIN2 truncation mutant (TIN2-13; Fig. 1a) did not affect TRF1 modification (Fig. 4). The lack of suppression of TRF1 modification by TIN2-13 is particularly informative because this protein retains the ability to bind to TRF1 (ref. 6; data not shown). Although TIN2 inhibited the modification of TRF1, it did not block the automodification activity of tankyrase 1 (∼10% reduction in automodification; inhibited by a factor of 0.9 ± 0.1, n = 3), further illustrating the specificity of the effect. We conclude that TIN2 can specifically block tankyrase 1 from modifying TRF1 in vitro.
The autoradiograph (top) of the tankyrase PARP assay. TRF1 protein (2 μg) was premixed with increasing amount of TIN2, TIN2-13 or hRap1 before addition of tankyrase 1 and 32P-β-NAD. The highest molar ratios of individual factor to TRF1 (the last lane in each group) are 2 (for TIN2) and 4 (for TIN2-13 and hRap1). Products from each reaction were loaded onto two separate SDS gels (8% and 10%) for autoradiography (top) and Coomassie blue staining (bottom).
These data identify TIN2 as a negative regulator of telomere length and also suggest a mechanism for this regulation. Through its ability to protect TRF1 from inactivation by tankyrase 1, TIN2 contributes to the accumulation of the TRF1 complex on telomeres. The TRF1 complex negatively regulates telomerase by loading POT1 on the single-stranded part of the telomere20. Inhibition of telomerase at the telomere terminus of over-elongated telomeres is therefore governed by the ability of the TRF1 complex to recruit POT1. We propose that this pathway requires the presence of TIN2 in the TRF1 complex, thereby preventing premature inactivation of TRF1 by tankyrase 1. In agreement with this possibility, longer telomeres contain more TRF1, TIN2 and POT1 (ref. 20).
These findings also suggest that TIN2 could facilitate the stable association of tankyrase 1 with the TRF1 complex. Although tankyrase 1 is a minor component of this complex, it is pertinent to ask what its function might be. One possibility, raised previously13,14, is that tankyrase 1 is necessary for the controlled dismantling of the telomeric complex, for example in S phase. The temporary removal of the TRF1 complex may be necessary for replication fork progression and telomerase-mediated telomere elongation. According to this view, TIN2 could be a crucial control point for these events. Inactivation of TIN2 at individual telomeres could immediately unleash the resident tankyrase 1, resulting in removal of the TRF1 complex. A second function for tankyrase 1 at telomeres, perhaps independent of its role in telomere length regulation, cannot be excluded, however. TRF1 is essential in mouse cells8,9, which is not easily explained by its role in telomere length regulation. Perhaps the death of Trf1-deficient mouse cells is related to an as yet unknown function of tankyrases at telomeres. Resolution of these issues requires experiments in which the association of tankyrases with telomeres is specifically inhibited.
The finding that telomeres contain a specific PARP modulator of tankyrase raises the prospect of finding similar regulatory factors at other sites of tankyrase function. In addition, PARP modulators akin to TIN2 may act on other PARPs, including PARP-1 and PARP-2.
Methods
TIN2 shRNAs and telomere length analysis.
We created eight shRNA retroviral constructs to target TIN2 mRNA using a PCR-based strategy7 to clone it into pENTR and a GATEWAY-modified pBabe vector with puromycin-selection marker. We produced retroviruses from amphotrophic Phoenix cells and used them to transduce HTC75 human fibrosarcoma cells2. Two constructs, shRNA2 (targeting nucleotides 344–371) and shRNA3 (targeting nucleotides 702–729; sequences available on request) substantially reduced TIN2 level in pooled HTC75 cells on western blots, as quantified by densitometry using AlphaImager 2200 program (Alpha Innotech). We used α-tubulin to normalize TIN2 signals. We used four HTC75 cell lines (pooled populations of infected cells) in telomere length analysis: shRNA2, shRNA3 and two negative control lines, shRNA1 and shRNA4, that did not cause TIN2 reduction. We split cells every 3 d and seeded 106 cells in a 15-cm dish for each passage. The growth rates of the cell lines were not significantly different. We isolated genomic DNA, digested it with AluI and MboI, separated it on 0.7% agarose gel and transferred it to Hybond membranes for hybridization using an 800-bp telomeric DNA probe from pSP73Sty11 labeled by Klenow fragment and α-32P-dCTP (ref. 2). We exposed the blot to PhosphoImager screen and quantified telomeric DNA signals using ImageQuan2. We calculated the rates of telomere elongation by linear regression.
TIN2 siRNA.
We synthesized two pairs of double-stranded siRNAs to target human TIN2 mRNA at nucleotides 303–323 and 740–759 (sequences available on request). Both pairs had similar effects on TRF1; the data presented in Figure 2 are from the 5′ pair (nucleotides 303–323). We transfected HeLa cells using oligofectamine (Invitrogen) and a protocol supplied by the manufacturer. We inoculated 1.5–2.0 × 105 cells per well in six-well plates. After 16–24 h, we subjected the cells to two sequential transfections, separated by a 24-h interval. We processed cells 48 h after the initial transfection for immunofluorescence or western blotting. In experiments using 3-aminobenzamide, we added 5 mM 3-aminobenzamide in DMSO or DMSO without 3-aminobenzamide to the medium during the 48-h transfection period, except for two 4-h periods in which cells were exposed to the transfection mixture in the absence of serum.
Transfection and immunoprecipitation.
We plated human 293T cells (5–6 × 106) and transfected them 20–24 h later by the calcium-phosphate coprecipitation method using 10–20 μg of plasmid DNA per 10-cm dish. We changed the medium after 12 h and collected cells 24–30 h after transfection. For immunoprecipitations, we dislodged 293T cells from the dish by flushing with cold phosphate-buffered saline (PBS), collected them by centrifugation and lysed them in ice-cold buffer (50 mM Tris-HCl (pH 7.4), 1 mM EDTA, 400 mM NaCl, 1% Triton X-100, 0.1% SDS, 1 mM dithiothreitol (DTT), 1 mM phenylmethylsulfonyl fluoride, 1 μg ml−1 of aprotinin, 10 μg ml−1 of pepstatin and 1 μg ml−1 of leupeptin). After 10 min on ice, we added an equal volume of ice-cold water and thoroughly mixed. After centrifugation in a microcentrifuge (14,000 r.p.m. for 10 min) we collected the supernatants and used them for immunoprecipitation. We prepared lysates from one 10-cm dish and mixed them with affinity-purified rabbit polyclonal antibody (371 and 864; 0.5–1 μg) or mouse monoclonal antibody (M2, 5 μg; 12CA5, 0.5 μg; 9E10, 1 μg) for each immunoprecipitation (at 4 °C for 5–6 h, nutating). During the final hour, we added 30 μl (settled volume) of protein G–Sepharose beads (preblocked overnight with 10% bovine serum albumin in PBS) to each tube. We washed the beads three times with 1:1 diluted lysis buffer, eluted proteins with Laemmli loading buffer and analyzed them by SDS-PAGE. For immunoprecipitations of the endogenous TRF1 complex from HeLa cells, we precleared lysates containing 5 mg of protein for 30 min at 4 °C by incubating them with 30 μl of settled protein G–Sepharose beads (unblocked). After removing protein G beads by microcentrifugation, we used 0.5 μg of purified rabbit antibodies against either TRF1 or TIN2 for each immunoprecipitation, as above.
For sequential immunoprecipitations, we transfected ten 10-cm plates of 293T cells with pLPC-FN-tankyrase 1 (8 μg per plate), pcDNA-HA-TIN2 (5 μg per plate) and pRc.CMV-MYC-TRF1 (5 μg per plate). We combined cells 24 h after transfection and lysed them in 5 ml as described above. We divided the lysate into two equal aliquots and added FLAG peptide (Sigma) to a final concentration of 100 μg ml−1 to one aliquot. We added anti-FLAG (M2)–sepharose beads (250 μl settled volume; Sigma) to each aliquot and rotated them for 6 h at 4 °C. We washed the beads four times and subjected them to four sequential affinity elutions, each using 250 μl of buffer containing 200 μg ml−1 of FLAG peptide (Sigma). We combined the eluates, divided them into two equal aliquots and incubated the aliquots for 6 h at 4 °C with 6 μg of antibody to MYC (9E10) or 8 μg of antibody to HA (12CA5). We eluted the precipitates with Laemmli buffer and analyzed them by western blotting.
In vitro PARP assay for tankyrase 1.
We carried out in vitro tankyrase 1 assays as described12 with slight modifications. We incubated 1–2 μg of proteins purified from baculovirus-infected insect cells with 32P-β-NAD+ (1.3 μM) at 25 °C for 30 min. We stopped the reactions by adding ice-cold trichloroacetic acid to 20%. After 10 min on ice, we collected proteins by microcentrifugation (10 min at 14,000 r.p.m. at 4 °C). We rinsed pellets gently with ice-cold 5% trichloroacetic acid and dissolved them in sample-loading buffer (1 M Tris-base, 12% SDS, 0.2 M DTT and 0.1% bromophenol blue). We separated the samples by SDS-PAGE, dried them and analyzed them by autoradiography. For experiments in which TIN2 was used to inhibit the tankyrase1 activity on TRF1, we premixed TIN2 (and other proteins as negative controls) with TRF1 for 30 min on ice before adding tankyrase 1 and 32P-β-NAD+.
Indirect immunofluorescence.
We grew cells glass coverslips, fixed them for 10 min at room temperature with PBS containing 2% paraformaldehyde and permeabilized them for 10 min in PBS plus 0.5% Nonidet P-40. We preblocked cells for at least 30 min with PBS plus 0.2% coldwater fish gelatin and 0.5% BSA before incubating them with primary antibody (2 h at room temperature or overnight at 4 °C). We used purified rabbit polyclonal antibody to TIN2 (865) and crude mouse polyclonal antiserum to TRF1 as primary antibodies and used rhodamine- or fluorescein-conjugated donkey antibodies to rabbit or to mouse (Jackson Laboratory) as secondary antibodies at dilutions recommended by the manufacturer. We did bleedthrough controls by leaving out one of the primary antibodies.
Differential KCl extraction of chromatin.
We treated cells with trypsin, washed them twice with medium containing 10% bovine calf serum and once with ice-cold PBS and resuspended them in ∼10 times the pellet volume of buffer C-150 (20 mM HEPES buffer (pH 7.9), 25% glycerol, 5 mM MgCl, 0.2% Nonidet P-40, 1 mM DTT, 1 mM phenylmethylsulfonyl fluoride, 1 μg ml−1 of aprotinin, 10 μg ml−1 of peptatin, 1 μg ml−1 of leupeptin and 150 mM KCl). After 15 min on ice, we collected supernatants (the 150 mM KCl fractions) containing soluble cytoplasmic and nucleoplasmic proteins by centrifugation at 3,000 r.p.m. for 5 min at 4 °C in a microcentrifuge. We suspended the pellets in buffer C-420 (as above but containing 420 mM KCl), incubated them on ice for 15 min and centrifuged them for 10 min at 14,000 r.p.m. at 4 °C (the 420 mM KCl fraction, containing chromatin-bound proteins). We sonicated the final pellets in Laemmli loading buffer.
Tankyrase 1 and TIN2 reagents.
All tankyrase 1 alleles used were tagged at the N terminus with a FLAG sequence followed by either a functional NLS (PKKKRKVE), or its nonfunctional counterpart (PKQKRKVE). To create alleles with inactive PARP domain (PARP-dead alleles, or tankyrase 1-PD), we changed His1164 and Glu1291 in the PARP homology domain to alanine by PCR-based site-directed mutagenesis. His1164 and Glu1291 align perfectly with two conserved amino acids in PARP-1 from different species, and mutating each of the two residues in human PARP-1 reduced the catalytic activity by factors of ∼100 and ∼500, respectively19. FN-tankyrase 1 consisted of FLAG, a functional NLS and wild-type tankyrase 1. FN-tankyrase 1-PD consisted of FLAG, a functional NLS and PARP-dead tankyrase 1. FN*-tankyrase 1 consisted of FLAG, a nonfunctional NLS and wild-type tankyrase 1. FN*-tankyrase 1-PD consisted of FLAG, a nonfunctional NLS and PARP-dead tankyrase 1. cDNA and baculoviral constructs encoding full-length TIN2 were gifts from J. Campisi (Lawrence Berkeley National Laboratory). We generated TIN2-13 (amino acids 196–354) by PCR amplification and subcloned it into mammalian expression vectors or a baculoviral expression construct (pFastBacHTa). We confirmed fidelity of the PCR products by sequencing. We generated antibodies 864 and 865 by immunizing rabbits with full length baculovirus-derived TIN2 and affinity-purification.
References
Chong, L. et al. A human telomeric protein. Science 270, 1663–1667 (1995).
van Steensel, B. & de Lange, T. Control of telomere length by the human telomeric protein TRF1. Nature 385, 740–743 (1997).
Smogorzewska, A. et al. Control of human telomere length by TRF1 and TRF2. Mol. Cell. Biol. 20, 1659–1668 (2000).
Ancelin, K. et al. Targeting assay to study the cis functions of human telomeric proteins: evidence for inhibition of telomerase by TRF1 and for activation of telomere degradation by TRF2. Mol. Cell. Biol. 22, 3474–3487 (2002).
Smogorzewska, A. & de Lange, T. Regulation of telomerase by telomeric proteins. Annu. Rev. Biochem. 73, 177–208 (2004).
Kim, S.H., Kaminker, P. & Campisi, J. TIN2, a new regulator of telomere length in human cells. Nat. Genet. 23, 405–412 (1999).
Hemann, M.T. et al. An epi-allelic series of p53 hypomorphs created by stable RNAi produces distinct tumor phenotype in vivo. Nat. Genet. 33, 396–400 (2003).
Karlseder, J. et al. Targeted deletion reveals an essential function for the telomere length regulator Trf1. Mol. Cell. Biol. 23, 6533–6541 (2003).
Iwano, T., Tachibana, M., Reth, M. & Shinkai, Y. Importance of TRF1 for Functional Telomere Structure. J. Biol. Chem. 279, 1442–1448 (2003).
Zhong, Z., Shiue, L., Kaplan, S. & de Lange, T. A mammalian factor that binds telomeric TTAGGG repeats in vitro. Mol. Cell. Biol. 12, 4834–4843 (1992).
Cook, B.D., Dynek, J.N., Chang, W., Shostak, G. & Smith, S. Role for the related poly(ADP-Ribose) polymerases tankyrase 1 and 2 at human telomeres. Mol. Cell. Biol. 22, 332–342 (2002).
Smith, S., Giriat, I., Schmitt, A. & de Lange, T. Tankyrase, a poly(ADP-ribose) polymerase at human telomeres. Science 282, 1484–1487 (1998).
Seimiya, H. & Smith, S. The telomeric poly(ADP-ribose) polymerase, tankyrase 1, contains multiple binding sites for telomeric repeat binding factor 1 (TRF1) and a novel acceptor, 182-kDa tankyrase-binding protein (TAB182). J. Biol. Chem. 277, 14116–14126 (2002).
De Rycker, M., Venkatesan, R.N., Wei, C. & Price, C.M. Vertebrate tankyrase domain structure and sterile alpha motif (SAM)-mediated multimerization. Biochem. J. 372, 87–96 (2003).
Lyon, R.J. et al. Identification of a novel human tankyrase through its interaction with the adaptor protein Grb14. J. Biol. Chem. 276, 17172–17180 (2001).
Chi, N.W. & Lodish, H.F. Tankyrase is a Golgi-associated mitogen-activated protein kinase substrate that interacts with IRAP in GLUT4 vesicles. J. Biol. Chem. 275, 38437–38444 (2000).
Smith, S. & de Lange, T. Tankyrase promotes telomere elongation in human cells. Curr. Biol. 10, 1299–1302 (2000).
Chang, W., Dynek, J.N. & Smith, S. TRF1 is degraded by ubiquitin-mediated proteolysis after release from telomeres. Genes Dev. 17, 1328–1333 (2003).
Marsischky, G.T., Wilson, B.A. & Collier, R.J. Role of glutamic acid 988 of human poly-ADP-ribose polymerase in polymer formation. Evidence for active site similarities to the ADP-ribosylating toxins. J. Biol. Chem. 270, 3247–3254 (1995).
Loayza, D. & De Lange, T. POT1 as a terminal transducer of TRF1 telomere length control. Nature 424, 1013–1018 (2003).
Acknowledgements
We thank J. Campisi and S. Kim for the human TIN2 cDNA and baculovirus, P. Cortes for advice on 293T transfection and sequential immunoprecipitations, G. Hannon for the retrovirus-based shRNA expression system, A. Himelblau and D. Hockemeyer for help in generating the antisera to TIN2 and K. Hoke and other members of T.d.L.'s laboratory for discussion and comments on this manuscript. This work was supported by grants from the US National Cancer Institute and Burroughs Welcome to T.d.L., by a Career Development Award from the US National Cancer Institute to J.Z.Y. and by a grant from the Translational & Integrative Medicine Research Fund from Sloan-Kettering Cancer Institute to J.Z.Y.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Ye, JS., de Lange, T. TIN2 is a tankyrase 1 PARP modulator in the TRF1 telomere length control complex. Nat Genet 36, 618–623 (2004). https://doi.org/10.1038/ng1360
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng1360
This article is cited by
-
Targeting telomeres: advances in telomere maintenance mechanism-specific cancer therapies
Nature Reviews Cancer (2022)
-
Functions of ADP-ribose transferases in the maintenance of telomere integrity
Cellular and Molecular Life Sciences (2022)
-
Poly(ADP-ribose) polymerase inhibition: past, present and future
Nature Reviews Drug Discovery (2020)
-
Structural and functional analyses of the mammalian TIN2-TPP1-TRF2 telomeric complex
Cell Research (2017)
-
82-kDa choline acetyltransferase and SATB1 localize to β-amyloid induced matrix attachment regions
Scientific Reports (2016)