Abstract
Gene fission and fusion, the processes by which a single gene is split into two separate genes and two adjacent genes are fused into a single gene, respectively, are among the primary processes that generate new genes1,2,3,4. Despite their seeming reversibility4,5, nothing is known about the mechanism of gene fission. Because the nucleotide sequences of fission genes record little about their origination process, conventional analysis of duplicate genes may not be powerful enough to unravel the underlying mechanism. In a survey for young genes in species of the Drosophila melanogaster subgroup using fluorescence in situ hybridization, we identified a young gene family, monkey king, whose genesis sheds light on the evolutionary process of gene fission. Its members originated 1–2 million years ago as retroposed duplicates and evolved into fission genes that separately encode protein domains from a multidomain ancestor. The mechanism underlying this process is gene duplication with subsequent partial degeneration.
Similar content being viewed by others
Main
To identify young genes in closely related Drosophila species (Fig. 1a), we used fluorescence in situ hybridization FISH analysis of polytene chromosomes with D. melanogaster cDNA probes. We selected the cDNAs that generated additional hybridization signals as candidates for further analysis. In this screening, we identified a gene family with up to three new members in the clade of D. simulans, D. sechellia and D. mauritiana, which have diverged in less than 1 million years6. We named this gene family monkey-king (mkg) after a mythical monkey king in ancient China who could transform his hair into many offspring.
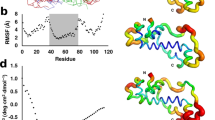
(a) Origination events in the phylogenetic tree of the D. melanogaster species subgroup6,27: D. melanogaster (Dmel), D. sechellia (Dsec), D. simulans (Dsim), D. mauritiana (Dmau), D. teissieri (Dtei), D. yakuba (Dyak), D. santomea (Dsan), D. erecta (Dere) and D. orena (Dore). mya, million years ago. Pink bars indicate mkg-p genes; red, mkg-r genes; black, mkg-r2 genes; and green, mkg-r3 genes. The circular symbol in Dmau\mkg-p indicates a degeneration event in the gene. (b) FISH detection of new genes in D. melanogaster subgroup using digoxigenin-labeled GH05885 cDNA as probe (the D. erecta data is not shown but is available on request). The cytological positions of the parental and new genes are given next to the insets that show the signals of the genes from FISH detection (green). (c) Neighbor-joining tree of members of the mkg family based on Kimura-2-parameter distance generated from the gene sequences. Bootstrap percentages are shown on the branches. Branch lengths are drawn to scale.
Two probes from cDNAs encoding zinc-finger protein–related sequences from the gene CG7163 in D. melanogaster had identical hybridization patterns: three extra signals in D. mauritiana and one extra signal in both D. simulans and D. sechellia in addition to a common signal at cytological site 66C of chromosome 3 in all species (Fig. 1b). The three new signals in D. mauritiana were localized at regions 9A (Dmau\mkg-r2), 9B (Dmau\mkg-r) and 5A (Dmau\mkg-r3) of the X chromosome. The new signal in D. simulans and D. sechellia was located at region 9B. Southern-blot hybridization experiments with digestion of genomic DNAs of the subgroup species by HindIII confirmed the FISH results (Supplementary Fig. 1 online).
We identified and sequenced all four mkg members in a D. mauritiana genomic library. From the sequences of the four copies with their flanking sequences, we observed extensive changes in the parental copy in D. mauritiana (Dmau\mkg-p) compared with D. melanogaster, including a large insertion in exon 2 that contained an inverted repeat of a downstream coding segment (Fig. 2a,b; sequence alignment data stored in GenBank), a 1-bp deletion in exon 3 and numerous substitutions. These changes disrupted the previous reading frame in these exons. The three new genes are intronless, suggesting that retroposition was involved in their origination7. Using flanking copy-specific sequence probes, we mapped their cytological positions in related species by FISH. The specific probe for the D. mauritiana 9B copy (Dmau\mkg-r) hybridized at the same cytological position as the 9B copy in D. simulans and D. sechellia, indicating that they are orthologs. Dmau\mkg-r2 at 9A seems to be derived from a single duplication event of Dmau\mkg-r, as they are similar in both the retrosequence and the immediate flanking sequences (sequence alignment data stored in GenBank). Dmau\mkg-r3 at 5A, which contains an AT-rich repetitive sequence with a repeat unit of ∼105 bp in length, seems to be a new processed retrosequence from the parental gene. This sequence is more closely related to the parental gene and has a poly-A tract and flanking short direct repeats (TATA/TATT) (data not shown). To our knowledge, this is the first example of new genes being repeatedly generated in a genome by retroposition from the same parent gene and then becoming fixed in natural populations within a short evolutionary period (<2 million years).
(a) Three transcripts of D. melanogaster mkg (CG7163). (b) The four members of mkg gene family in D. mauritiana. Different transcripts are shown for mkg-p and Dmau\mkg-r3. (c,d) mkg-r and the parental copies of mkg in D. simulans and D. sechellia. The cytological positions are indicated. Green boxes indicate protein-coding regions, red boxes are untranslated regions, and white boxes show pseudoexon regions. Lines are introns or intergenic regions. Double lines indicate the indigenous flanking sequence at the new loci. Arrows show the inverted repeat in Dmau\mkg-p. Black triangles indicate the repetitive sequence proximal to Dmau\mkg-r3. The retained poly-A tract is indicated in Dmau\mkg-r3. (e) A schematic model of gene fission by duplication and subsequent partial degeneration, based on findings from Dmau\mkg-p and Dmau\mkg-r3.
We constructed a neighbor-joining tree by defining the gene CG7163 in D. melanogaster (Dmel\mkg-p) as the outgroup (Fig. 1c). The tree supports the relationship deduced from molecular features described above; retroposed mkg-r in the three species share a common ancestor, Dmau\mkg-r2 is a duplicate of Dmau\mkg-r, and Dmau\mkg-r3 is a recently retroposed gene from Dmau\mkg-p.
Processed copies of gene duplicates are often called pseudogenes because these retroposed elements usually do not carry promoters and insert randomly in the genome. But retroposition can contribute to evolution by creating new functional genes8,9,10. To study whether these three new copies and the changed parental copy in the mkg family were expressed, we carried out RT-PCR and detected transcripts of Dmau\mkg-r, Dmau\mkg-r3 and Dmau\mkg-p, but not Dmau\mkg-r2 (Fig. 3). mkg-r in D. simulans (Dsim\mkg-r) and D. sechellia (Dsec\mkg-r) was expressed only in adult males, whereas Dmau\mkg-r was detected in both sexes, with stronger signals in females (Fig. 3).
We characterized gene structures of all mkg members by 5′ and 3′ RACE (Fig. 2b–d). Each mkg-r copy in the three species had different fates concerning survival and exaptation as new genes. Dmau\mkg-r retained a transcript similar to that of the original gene CG7163 and is expressed ubiquitously, as detectable by RT-PCR. As the retroposed mkg-r sequence does not contain the parental promoter, the retrosequence was probably fortuitously inserted adjacent to a sequence with basal promoter function. Furthermore, the new promoters evolved rapidly: four new promoters were created in a short evolutionary time. Dsim\mkg-r and Dsec\mkg-r had new transcription initiation sites. Compared with Dmau\mkg-r, Dsim\mkg-r had a 139-bp deletion, a 1-bp deletion and several substitutions in the upstream region, which may be correlated with the new male-specific expression pattern (Fig. 2c). In Dsec\mkg-r, the transcription start point was located in the center of the original exon 2 (Fig. 2d), suggesting that the former coding sequence had become a regulatory region. There were three substitutions within the 100 bp of sequence upstream of the new transcription start site of Dsec\mkg-r, compared with the other mkg-r sequences. The finding that new promoters evolved in these new genes less than 1 million years ago supported the prediction that new promoters could originate and evolve rapidly11.
We further investigated functionality of the mkg family. The ratio of Ka (nonsynonymous substitution rate) to Ks (synonymous substitution rate) is a simple and useful measurement of functional constraint on a protein-coding gene12. The Ka/Ks ratio should be 1 for pseudogenes, <1 for genes subject to functional constraint and >1 for genes subject to strong positive darwinian selection13. But because new genes have rapid changes in protein sequences4,8, they may have more similar values of Ka and Ks than their parental genes, bringing the Ka/Ks ratio closer to 1. Therefore, the Ka/Ks ratio of a new gene may not be a powerful measure of functionality. Considering this peculiarity of new gene origination, we developed four independent lines of functionality analysis.
First, we found that all Ka/Ks ratios between expressed offspring copies, between expressed offspring and parental copies and between parental copies are <1 (Table 1). Under the null hypothesis that Ka/Ks ratios for pseudogenes would be randomly distributed around 1, the probability that all mkg members would have Ka/Ks ratios <1 is low (P < 0.0156). Second, a within-species variation analysis showed that all gene members, except one with only one polymorphism (Dsec\mkg-r), have fewer replacement polymorphisms than synonymous changes (Table 2). Third, comparison of mutation patterns in coding and noncoding regions showed functional constraints among mkg-r genes. Although numerous deletions occur in the 3′ untranslated regions of mkg-r genes and different deletion patterns are associated with different species, no deletion was found in the coding regions (protein sequence data stored in GenBank), suggestive of a functional constraint on the coding regions of all mkg members. Fourth, these analyses showed that almost all members of the mkg family are expressed with tissue-specific patterns. Taken together, these results suggest that all mkg members are functional.
Further analysis of these new gene members identified a mechanism for gene fission: duplication was followed by subsequent partial degeneration to form complementary functions between Dmau\mkg-p and Dmau\mkg-r3 (Fig. 2e). By BLAST search in the SWISS-PROT database, we found that the 659–amino acid product of the parental gene in D. melanogaster contains two domains (protein sequence data stored in GenBank). The first segment of 50 amino acids is homologous to the KRAB domain of zinc-finger protein 267 (ref. 14 and protein sequence data stored in GenBank), and the region of amino acids 100–400 is homologous to a group of proteins that contain a poly(A)-binding domain15. At first glimpse, the parental copy in D. mauritiana (Dmau\mkg-p) seems to be a pseudogene because of extensive disruptions of the open reading frame in exons 2 and 3 (Fig. 1a). But RACE and RT-PCR experiments showed that this disrupted region had become an intron sequence and is spliced out together with the original intron 2 (Fig. 2b; sequence alignment data stored in GenBank) and that the parental copy in D. mauritiana actually encodes a protein starting from amino acid residue 290 of the ancestral protein, containing the poly(A) binding region (protein sequence data stored in GenBank).
We detected three short transcripts of Dmau\mkg-r3 (Fig. 2b). All of them contain intact coding sequence for the KRAB domain of parental zinc-finger proteins and are polyadenylated at 5′ premature positions. Dmau\mkg-r3 is expressed as ubiquitously as the parental copy (Fig. 3), probably providing a KRAB-domain function that has been lost in Dmau\mkg-p. It is conceivable that, in its early stage, this new duplicate was redundant with the parental gene and that subsequent complementary degenerations resulted in the current compensatory pattern. The result was that the ancestral mkg-p gene in D. melanogaster, D. yakuba and D. teissieri was split into two loci in D. mauritiana, a typical case of gene fission.
The idea that these new genes were subject to functional partitions and evolution was also supported by evolutionary analysis. A relative rate test13,16 for these genes using Dmel\mkg-p as the outgroup showed that evolution rates were significantly higher in the mkg-r and mkg-r2 genes than in the parental genes (Fig. 4), except for Dmau\mkg-r3, which has a small number of changes with a limited statistical power (data not shown). Dmau\mkg-p was not included in the comparison because of its evolved gene structure. This suggests possible functional divergence of mkg-r and mkg-r2 from the parental genes, although population genetic tests16,17,18,19,20 did not detect recent selection on these genes in D. mauritiana and D. sechellia.
Although gene fission is a common process in prokaryotes2 and has been reported in eukaryotes4,21, the mechanism involved was unknown. The process by which mkg originated identified the first such mechanism: duplication followed by complementary partial degeneration. This also provides a mechanism to generate new introns in the degenerate region of a previously intronless gene by creating new splice signals22 (Fig. 2b). The molecular origins of the mkg family are reminiscent of the multifunctional gene model23 that proposes specification of protein functions in duplicate copies and the later subfunctionalization model24 in which gene duplicates are maintained by a complementary expression pattern. But the mkg family shows that two or more distinct genes can be derived from different domains of an ancestral protein through a fission process whose mechanism differs entirely from that of its inverse, gene fusion3,4,5,25.
Methods
FISH analysis of polytene chromosomes.
To generalize mechanisms of new gene origination, we searched for young genes by using D. melanogaster cDNAs from the Drosophila Gene Collection (Research Genetics, Invitrogen) for in situ hybridization on the polytene chromosomes of species in the D. melanogaster subgroup26. There are nine species in the D. melanogaster subgroup: D. melanogaster, D. simulans, D. mauritiana, D. sechellia, D. teissieri, D. yakuba, D. santomea27, D. erecta and D. orena (Fig. 1a). We amplified the cDNA inserts and labeled them by PCR using the vector primers (T7 and PM001). We labeled the probes with digoxigenin or biotin (Roche Molecular Biochemicals). By comparing hybridization signals in the polytene chromosomes of these species, we could detect new homologs that were duplicated to new cytological sites by retroposition or other possible processes.
Southern-blot hybridization.
We extracted genomic DNAs of D. melanogaster, D. simulans, D. mauritiana, D. sechellia, D. teissieri, D. yakuba, D. erecta and D. orena using the Puregene DNA isolation kit (Gentra Systems). We digested DNAs with HindIII, separated them on an agarose gel and transferred them to a nylon membrane (Roche Molecular Biochemicals) by Southern blotting. We hybridized the GH05885 probes to the membrane to confirm the copy numbers in different species as detected by FISH experiments.
Screening genomic DNA library and sequencing positive clones.
To obtain sequences of all the mkg copies in D. mauritiana, we screened a λ phage genomic library of D. mauritiana (constructed and provided by C.-T. Ting, University of Chicago). All four copies, including the parental one, were identified and sequenced.
Characterization of gene structures and expression patterns.
We used the RACE (rapid amplification of cDNA ends) assay and RT-PCR to detect possible transcripts of each copies. The gene structure of each copy was deduced by comparing the obtained cDNA and genomic DNA sequences. We examined expression patterns in adult females, adult males, second and third instar larvae or pupae. We carried out 5′ RACE using the FirstChoice RLM-RACE kit (Ambion). For 3′ RACE, we used adapter-linked oligo dT primers (Life Technologies) to synthesize first-strand cDNA.
Polymorphism data and statistical analyses.
We generated polymorphism data of genes using population samples of D. simulans, D. mauritiana and D. sechellia. The worldwide D. simulans sample contained 11 strains. We used 13 D. mauritiana strains: lines 72, 75, 105, 197, 207, g23, g35, g62, g74, g122, g130, g193 and G122. We used eight D. sechellia strains: lines 4, 15, 21, 22, 24, 25, 81 and 034. These strains were provided by J. Coyne, S.-C. Tsaur and M.-L. Wu (University of Chicago). We extracted total DNA from a single male of each strain using Puregene DNA extraction kit (Gentra Systems). We did not collect polymorphism data for Dmau\mkg-r because it is difficult to specifically amplify this gene in many D. mauritiana strains. In mkg-r2, all 12 alleles showed significantly stronger preference for synonymous polymorphism, whereas one allele (w136) contained a stop codon, probably a transient mutant, and was excluded from analysis. Alleles of mkg-r2 also had a preference for synonymous substitution (Ks) over nonsynonymous substitution (Ka) in divergence analysis in all comparisons, except for one with mkg-r3. Thus, the sequence analyses at two levels of variation, polymorphism and divergence, suggest that mkg-r2 is subject to functional constraint.
We calculated Ka, Ks and two statistics that describe within-species variation, π (nucleotide diversity) and Watterson's θ, with DNAsp 3.5 (ref. 16) and K-estimator28. We also carried out Tajima's D test, the McDonald-Kreitman test, Fu-Li test and Fay-Wu test17,18,19,20 with DNAsp 3.5. We created a neighbor-joining tree of genes and carried out a relative rate test using Mega 2.0 (ref. 29).
We used sign tests30 to test the null hypothesis that new genes and parental genes are pseudogenes. If they were pseudogenes, some new genes should have Ka/Ks ratios <1 and some others should have Ka/Ks ratios >1. Under the simple assumption that the probability that the Ka/Ks ratio for a pseudogene is <1 is equal to the probability that the ratio is >1, the binomial distribution on the assumption of p = q = 0.5 can be used to calculate the probability of the null hypothesis, p = n!/(m!k!).(1/2)n, where n = m + k, m = the number of the ratios >1 and k = the number of the ratios <1. For a conservative test, we used the number of independent comparison (n = 6, considering that Dmel\mkg-p, Dsim\mkg-p, Dsec\mkg-p, Dsim\mkg-r, Dsec\mkg-r and Dmau\mkg-r contain two domains whereas Dmau\mkg-p and Dmau\mkg-r3 contain single different domains) when computing this probability.
GenBank accession number.
mkg genes, AY562976–AY562984; sequence alignment data, AY572491–AY572499.
Note: Supplementary information is available on the Nature Genetics website.
References
Enright, A.J., Iliopoulos, I., Kyrpides, N.C. & Ouzounis, C.A. Protein interaction maps for complete genomes based on gene fusion events. Nature 402, 86– 90 (1999).
Snel, B., Bork, P. & Huynen, M. Genome evolution - gene fusion versus fission. Trends Genet. 16, 9– 11 (2000).
Nurminsky, D.I., Nurminskaya, M.V., De Aguiar, D. & Hartl, D.L. Selective sweep of a newly evolved sperm-specific gene in Drosophila. Nature 396, 572– 575 (1998).
Long, M., Betrán, E., Thornton, K. & Wang, W. Origin of new genes: Glimpse from young and old. Nat. Rev. Genet. 4, 865– 875 (2003).
Thomson, T.M. et al. Fusion of the human gene for the polyubiquitination coeffector UEV1 with Kua, a newly identified gene. Genome Res. 10, 1743– 1756 (2000).
Powell, J.R. Progress and Prospects in Evolutionary Biology, the Drosophila Model (Oxford University Press, New York, 1997).
Rogers, J.H. The origin and evolution of retroposons. Int. Rev. Cytol. 93, 187– 279 (1985).
Brosius, J. Retroposition—seeds of evolution. Science 251, 753 (1991).
Long, M. & Langley, C.H. Natural selection and the origin of jingwei, a chimeric processed functional gene in Drosophila . Science 260, 91– 95 (1993).
Wang, W., Brunet, F.G., Nevo, E. & Long, M. Origin of Sphinx, a young chimeric RNA gene in Drosophila melanogaster . Proc. Natl. Acad. Sci. USA 99, 4448– 4453 (2002).
Stone, J.R. & Wray, G.A. Rapid evolution of cis-regulatory sequences via local point mutations. Mol. Biol. Evol. 18, 1764– 1770 (2001).
Nekrutenko, A., Makova, K.D. & Li, W.-H. The K(A)/K(S) ratio test for assessing the protein-coding potential of genomic regions: an empirical and simulation study. Genome Res. 12, 198– 202 (2002).
Li, W.-H. Molecular Evolution (Sinaur Associates, Sunderland, Massachusetts, 1997).
Abrink, M., Aveskogh, M. & Hellman, L. Isolation of cDNA clones for 42 different Kruppel-related zinc finger proteins expressed in the human monoblast cell line U-937. DNA Cell. Biol. 14, 125– 136 (1995).
The FlyBase Consortium. The FlyBase database of the Drosophila genome projects and community literature. Nucleic Acids Res. 31, 172– 175 (2003).
Rozas, J. & Rozas, R. DnaSP version 3: an integrated program for molecular population genetics and molecular evolution analysis. Bioinformatics 15, 174– 175 (1999).
McDonald, J.H. & Kreitman, M. Adaptative protein evolution at the Adh locus in Drosophila . Nature 351, 652– 654 (1991).
Tajima, F. Statistical method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics 123, 585– 595 (1989).
Fay, J.C. & Wu, C.-I. Hitchhiking under positive Darwinian selection. Genetics 155, 1405– 1413 (2000).
Fu, Y.-H. & Li, W.-H. Statistical tests of neutrality of mutations. Genetics 133, 6934– 6945 (1993).
Altschmied, J. et al. Subfunctionalization of duplicate mitf genes associated with differential degeneration of alternative exons in fish. Genetics, 161, 259– 67 (2002).
Brosius, J. Many G-protein-coupled receptors are encoded by retrogenes. Trends Genet. 15, 304– 305 (1999).
Hughes, A.L. The evolution of functionally novel proteins after gene duplication. Proc. R. Soc. Lond. B 256, 119– 124 (1994).
Force, A. et al. Preservation of duplicate genes by complementary, degenerative mutations. Genetics 151, 1531– 1545 (1999).
Finta, C. & Zaphiropoulos, P.G. The human cytochrome P450 3A locus: Gene evolution by capture of downstream exons. Gene 260, 13– 26 (2000).
Wang, W., Zhang, J., Alvarez, C., Llopart, A. & Long, M. The origin of the jingwei gene and the complex modular structure of its parental gene, yellow emperor, in D. melanogaster . Mol. Biol. Evol. 17, 1294– 1301 (2000).
Lachaise, D. et al. Evolutionary novelties in islands: Drosophila santomea, a new melanogaster sister species from Sao Tome. Proc. R. Soc. Lond. B Biol. Sci. 267, 1487– 1495 (2000).
Comeron, J.M. K-estimator: calculation of the number of nucleotide substitutions per site and the confidence intervals. Bioinformatics 15, 763– 764 (1999).
Kumar, S., Tamura, K., Jakobsen, I.B. & Nei, M. MEGA2: Molecular Evolutionary Genetics Analysis software (Arizona State University, Tempe, Arizona, 2001).
Sokal, R.R. & Rohlf, F.J. Biometry 3rd edn. (Freeman, New York, 2000).
Acknowledgements
We thank J. Spofford and T. Martin for critical reading of the manuscript; the members of M.L.'s laboratory for discussions; J. Coyne, S.-C. Tsaur and M.-L. Wu for fly strains; and C.-T. Ting for the genomic library of D. mauritiana. This work was supported by a Packard Fellowship in Science and Engineering, a National Science Foundation CAREER award and a grant from the US National Institutes of Health to M.L, and a CAS-Max Planck Society Fellowship, a CAS key project grant and a National Science Foundation of China award to W.W.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Wang, W., Yu, H. & Long, M. Duplication-degeneration as a mechanism of gene fission and the origin of new genes in Drosophila species. Nat Genet 36, 523–527 (2004). https://doi.org/10.1038/ng1338
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng1338
This article is cited by
-
On the Unknown Proteins of Eukaryotic Proteomes
Journal of Molecular Evolution (2023)
-
Gene fusion as an important mechanism to generate new genes in the genus Oryza
Genome Biology (2022)
-
Identification of polycistronic transcriptional units and non-canonical introns in green algal chloroplasts based on long-read RNA sequencing data
BMC Genomics (2021)
-
Rapid evolutionary dynamics of an expanding family of meiotic drive factors and their hpRNA suppressors
Nature Ecology & Evolution (2021)
-
Evolutionary patterns of chimeric retrogenes in Oryza species
Scientific Reports (2019)