Abstract
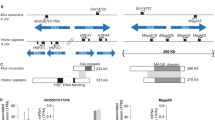
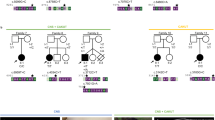
Vertebrates position unpaired organs of the chest and abdomen asymmetrically along the left–right (LR) body axis. Each structure comes to lie non-randomly with respect to the midline in an overall position designated situs solitus, exemplified in humans by placement of the heart, stomach and spleen consistently to the left. Aberrant LR axis development can lead to randomization of individual organ position (situs ambiguus) or to mirror-image reversal of all lateralized structures (situs in versus)1. Previously we mapped a locus for situs abnormalities in humans, HTX1, to Xq26.2 by linkage analysis in a single family (LR1) and by detection of a deletion in an unrelated situs ambiguus male (Family LR2; refs 2,3). From this chromosomal region we have positionally cloned ZIC3, a gene encoding a putative zinc-finger transcription factor. One frameshift, two missense and two nonsense mutations have been identified in familial and sporadic situs ambiguus. The frameshift allele is also associated with situs inversus among some heterozygous females, suggesting that ZIC3 functions in the earliest stages of LR-axis formation. ZIC3, which has not been previously implicated in vertebrate LR-axis development, is the first gene unequivocally associated with human situs abnormalities.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Kosaki, K. & Casey, B. Genetics of human left-right axis malformations. Semin. Cell Dev. Biol. (in the press).
Casey, B., Devoto, M., Jones, K.L. & Ballabio, A. Mapping a gene for familial situs abnormalities to human chromosome Xq24-q27. 1.. Nature Genet. 5, 403–407 (1993).
Ferrero, G.B. et al. A submicroscopic deletion in Xq26 associated with familial situs ambiguussitus ambiguus. Am. J. Hum. Genet. 61, 395–401 (1997).
Larsen, W.J., Human Embryology (Churchill-Livingstone, New York, 1993).
Britz-Cunningham, S.H., Shah, M.M., Zuppan, C.W. & Fletcher, W.H. Mutations of the connexin43 gap-junction gene in patients with heart malformations and defects of laterality. N. Engl. J. Med. 332, 1323–1329 (1995).
Gebbia, M., Towbin, J.A. & Casey, B. Failure to detect connexin43 mutations in 38 cases of sporadic and familial heterotaxy. Circulation 94, 1909–1912 (1996).
Penman-Splitt, M., Tsai, M.Y., Burn, J. & Goodship, J.A. Absence of mutations in the regulatory domain of the gap junction protein connexin 43 in patients with visceroatrial heterotaxy. Heart 77, 369–370 (1997).
Pilia, G. et al. YAC/STS map of 9 Mb of Xq26 at 100-kb resolution, localizing 6 ESTs, 6 genes, and 32 genetic markers. Genomics 34, 55–62 (1996).
Aruga, J. et al. The mouse Zic gene family. J. Biol. Chem. 271, 1043–1047 (1996).
Aruga, J. et al. Identification and characterization of Zic4, a new member ofthe mouse Zic gene family. Gene 172, 291–294 (1996).
Nagai, T. et al. The expression of the mouse Zic7, Zic2, and Zic3 gene suggests an essential role for Zic genes in body pattern formation. Dev.Biol. 182, 299–313 (1997).
Benedyk, M.J., Mullen, J.R. & DiNardo, S. odd-paired: a zinc finger pair-rule protein required for the tirnely activation of engrailed and wingless in Drosophila embryos. Genes Dev. 8, 105–117 (1994).
Cimbora, D.M. & Sakonju, S. Drosophila midgut morphogenesis requires the function of the segmentation gene odd-paired. Dev. Biol. 169, 580–595 (1995).
Levin, M., Johnson, R.L., Stern, C.D., Kuehn, M. & Tabin, C. A molecular pathway determining left-right asymmetry in chick embryogenesis. Cell 82, 803–814 (1995).
Levin, M. Left-right asymmetry in vertebrate embryogenesis. Bioessays 19, 287–296 (1997).
Kozak, M. Structural features in eukaryotic mRNAs that modulate the initiation of translation. j Biol. Chem. 266, 196–197 (1991).
Casey, B. et al. Autosomal dominant transmission of familial laterality defects. Am. J. Med. Genet. 61, 325–328 (1996).
Lowe, L.A. et al. Conserved left-right asymmetry of nodal expression and alterations in murine situs inversus. Nature 381, 158–161 (1996).
Collignon, J., Varlet, I. & Robertson, E.J. Relationship between asymmetric nodal expression and the direction of embryonic turning. Nature 381, 155–158 (1996).
Meno, C. et al. Left-right asymmetric expression of the TGFβ-family member lefty in mouse embryos. Nature 381, 151–155 (1996).
Isaac, A., Sargent, M.G. & Cooke, J. Control of vertebrate left-right asymmetry by a snail-related zinc finger gene. Science 275, 1301–1304 (1997).
Danos, M.C. & Yost, H.J. Linkage of cardiac left-right asymmetry and dorsal-anterior development in Xenopus. Development 121, 1467–1474 (1995).
Danos, M.C. & Yost, H.J. Role of notochord in specification ofthe cardiac left-right axis in zebrafish and Xenopus. Dev. Biol. 177, 96–103 (1996).
Lohr, J.L., Danos, M.C. & Yost, H.J. Left-right asymmetry of a noda/-related gene is regulated by dorsoanterior midline structures during Xenopus development. Development, 124, 1465–1472 (1997).
Hyatt, B.A., Lohr, J.L. & Yost, H.J. Initiation of vertebrate left-right axis formation by maternal Vg1. Nature 384, 62–65 (1996).
Burke, D., Carle, G. & Olson, M. Cloning of large segments of exogenous DNA into yeast by means of artificial chromosome vectors. Science 236, 806–812 (1987).
Wapenaar, M.C. et al. A YAC-based binning strategy for facilitating the rapid assembly of cosmid contigs: 1.6 Mb of overlapping cosmids in Xp22. Hum. Mol. Genet. 3, 1155–1161 (1994).
Orita, M., Suzuki, Y., Sekiya, T. & Haayashi, K. Rapid and sensitive detection of point mutations and DNA polymorphisms using the polymerase chain reaction. Genomics 5, 874–879 (1989).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Gebbia, M., Ferrero, G., Pilia, G. et al. X-linked situs abnormalities result from mutations in ZIC3. Nat Genet 17, 305–308 (1997). https://doi.org/10.1038/ng1197-305
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/ng1197-305