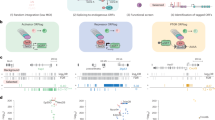
Abstract
To verify the genome annotation and to create a resource to functionally characterize the proteome, we attempted to Gateway-clone all predicted protein-encoding open reading frames (ORFs), or the 'ORFeome,' of Caenorhabditis elegans. We successfully cloned approximately 12,000 ORFs (ORFeome 1.1), of which roughly 4,000 correspond to genes that are untouched by any cDNA or expressed-sequence tag (EST). More than 50% of predicted genes needed corrections in their intron-exon structures. Notably, approximately 11,000 C. elegans proteins can now be expressed under many conditions and characterized using various high-throughput strategies, including large-scale interactome mapping. We suggest that similar ORFeome projects will be valuable for other organisms, including humans.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Mardis, E., McPherson, J., Martienssen, R., Wilson, R.K. & McCombie, W.R. What is finished, and why does it matter. Genome Res. 12, 669–671 (2002).
Vidal, M. A biological atlas of functional maps. Cell 104, 333–339 (2001).
Ideker, T., Galitski, T. & Hood, L. A new approach to decoding life: systems biology. Annu. Rev. Genomics Hum. Genet. 2, 343–372 (2001).
Kapranov, P. et al. Large-scale transcriptional activity in human chromosomes 21 and 22. Science 296, 916–919 (2002).
The International Human Genome Sequencing Consortium. Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001).
Reboul, J. et al. Open-reading-frame sequence tags (OSTs) support the existence of at least 17,300 genes in C. elegans. Nat. Genet. 27, 332–336 (2001).
Blandin, G. et al. Genomic exploration of the hemiascomycetous yeasts: 4. The genome of Saccharomyces cerevisiae revisited. FEBS Lett. 487, 31–36 (2000).
Oshiro, G. et al. Parallel identification of new genes in Saccharomyces cerevisiae. Genome Res. 12, 1210–1220 (2002).
Zhu, H. et al. Global analysis of protein activities using proteome chips. Science 293, 2101–2105 (2001).
MacBeath, G. & Schreiber, S.L. Printing proteins as microarrays for high-throughput function determination. Science 289, 1760–1763 (2000).
Ziauddin, J. & Sabatini, D.M. Microarrays of cells expressing defined cDNAs. Nature 411, 107–110 (2001).
Gera, J.F., Hazbun, T.R. & Fields, S. Array-based methods for identifying protein–protein and protein–nucleic acid interactions. Methods Enzymol. 350, 499–512 (2002).
Mammalian Gene Collection (MGC) Program Team. Generation and initial analysis of more than 15,000 full-length human and mouse cDNA sequences. Proc. Natl. Acad. Sci. USA 99, 16899–16903 (2002).
The FANTOM Consortium and the RIKEN Genome Exploration Research Group Phase I & II Team. Analysis of the mouse transcriptome based on functional annotation of 60,770 full-length cDNAs. Nature 420, 563–573 (2002).
Seki, M. et al. Functional annotation of a full-length Arabidopsis cDNA collection. Science 296, 141–145 (2002).
Stein, L., Sternberg, P., Durbin, R., Thierry-Mieg, J. & Spieth, J. WormBase: network access to the genome and biology of Caenorhabditis elegans. Nucleic Acids Res. 29, 82–86 (2001).
Walhout, A.J. et al. Gateway recombinational cloning: application to the cloning of large numbers of open reading frames or ORFeomes. Methods Enzymol. 328, 575–592 (2000).
Walhout, A.J. et al. Protein interaction mapping in C. elegans using proteins involved in vulval development. Science 287, 116–122 (2000).
Hartley, J.L., Temple, G.F. & Brasch, M.A. DNA cloning using in vitro site-specific recombination. Genome Res. 10, 1788–1795 (2000).
The C. elegans Sequencing Consortium. Genome sequence of the nematode C. elegans: a platform for investigating biology. Science 282, 2012–2018 (1998).
Morin, X., Daneman, R., Zavortink, M. & Chia, W. A protein trap strategy to detect GFP-tagged proteins expressed from their endogenous loci in Drosophila. Proc. Natl. Acad. Sci. USA 98, 15050–15055 (2001).
Harrison, P.M., Echols, N. & Gerstein, M.B. Digging for dead genes: an analysis of the characteristics of the pseudogene population in the Caenorhabditis elegans genome. Nucleic Acids Res. 29, 818–830 (2001).
Vaglio, P. et al. WorfDB: the C. elegans ORFeome Database. Nucleic Acids Res. 31, 237–240 (2003).
Hazbun, T.R. & Fields, S. Networking proteins in yeast. Proc. Natl. Acad. Sci. USA 98, 4277–4278 (2001).
Davy, A. et al. A protein–protein interaction map of the Caenorhabditis elegans 26S proteasome. EMBO Rep. 2, 821–828 (2001).
Boulton, S.J. et al. Combined functional genomic maps of the C. elegans DNA damage response. Science 295, 127–131 (2002).
Kinoshita, N., Minshull, J. & Kirschner, M.W. The identification of two novel ligands of the FGF receptor by a yeast screening method and their activity in Xenopus development. Cell 83, 621–630 (1995).
Braun, P. et al. Proteome-scale purification of human proteins from bacteria. Proc. Natl. Acad. Sci. USA 99, 2654–2659 (2002).
Hammarstrom, M., Hellgren, N., van Den Berg, S., Berglund, H. & Hard, T. Rapid screening for improved solubility of small human proteins produced as fusion proteins in Escherichia coli. Protein Sci. 11, 313–321 (2002).
Hillier, L. & Green, P. OSP: a computer program for choosing PCR and DNA sequencing primers. PCR Methods Appl. 1, 124–128 (1991).
Acknowledgements
We thank the C. elegans Sequencing Consortium for the genome sequence; the participants of the annual ORFeome meeting for their input and numerous suggestions; the members of M.V.'s laboratory for their input and help; C. McCowan for administrative assistance; B. Sobhian, A.-S. Nicot, N. Tzellas and the GenomeVision Service sequencing staff at Genome Therapeutics for technical assistance; and P. Braun for the protein expression plasmids. This work was supported by grants from the National Cancer Institute, the National Human Genome Research Institute, the National Institute of General Medical Sciences and the Merck Genome Research Institute awarded to M.V.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
T.M. has financial interests in Open Biosystems, one of the companies responsible for the distribution of the C. elegans ORFeome version 1.1.
Supplementary information
Rights and permissions
About this article
Cite this article
Reboul, J., Vaglio, P., Rual, JF. et al. C. elegans ORFeome version 1.1: experimental verification of the genome annotation and resource for proteome-scale protein expression. Nat Genet 34, 35–41 (2003). https://doi.org/10.1038/ng1140
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng1140
This article is cited by
-
Sulfur amino acid supplementation displays therapeutic potential in a C. elegans model of Duchenne muscular dystrophy
Communications Biology (2022)
-
DAF-16/FOXO requires Protein Phosphatase 4 to initiate transcription of stress resistance and longevity promoting genes
Nature Communications (2020)
-
Biosynthetic Pathway Analysis for Improving the Cordycepin and Cordycepic Acid Production in Hirsutella sinensis
Applied Biochemistry and Biotechnology (2016)
-
Transcriptome sequencing and analysis of the entomopathogenic fungus Hirsutella sinensis isolated from Ophiocordyceps sinensis
BMC Genomics (2015)
-
Soybean transcription factor ORFeome associated with drought resistance: a valuable resource to accelerate research on abiotic stress resistance
BMC Genomics (2015)