Abstract
Impaired axonal transport in motor neurons has been proposed as a mechanism for neuronal degeneration in motor neuron disease. Here we show linkage of a lower motor neuron disease to a region of 4 Mb at chromosome 2p13. Mutation analysis of a gene in this interval that encodes the largest subunit of the axonal transport protein dynactin showed a single base-pair change resulting in an amino-acid substitution that is predicted to distort the folding of dynactin's microtubule-binding domain. Binding assays show decreased binding of the mutant protein to microtubules. Our results show that dysfunction of dynactin-mediated transport can lead to human motor neuron disease.
Similar content being viewed by others
Main
Selective death of motor neurons occurs in a variety of different diseases that affect upper and lower motor neurons. Although a common mechanism has not been identified, defects in axonal transport have long been implicated in this process1. Transgenic mice with mutant superoxide dismutase-1 show deficits in slow axonal transport early in the disease course2, and an early upregulation of the kinesin superfamily motor protein KIF1A was detected in spinal motor neurons3. In a cell model of spinal bulbar muscular atrophy, a motor neuron disease caused by a polyglutamine repeat expansion, cytoplasmic aggregates alter axonal trafficking4. Furthermore, mutations in the gene encoding KIF1Bβ cause an axonal form of a hereditary neuropathy5.
We identified a mutation in the gene encoding the p150 subunit of the transporter protein dynactin (DCTN1) in a family with a slowly progressive, autosomal dominant form of lower motor neuron disease without sensory symptoms (see Supplementary Fig. 1 and Supplementary Note 1 online). Onset of the disease is in early adulthood with breathing difficulty due to vocal fold paralysis, progressive facial weakness and weakness and muscle atrophy in the hands. Weakness and muscle atrophy in the distal lower extremities develop later. A genome-wide screen showed linkage to chromosome 2p13 between flanking markers D2S291 and D2S2114, with a maximum lod score of 4.05 for D2S2109 at recombination fraction θ = 0 (Fig. 1a). Multipoint linkage analysis with the markers D2S2110, D2S2109 and D2S286 gave a maximum lod score of 5.6. The gene DCTN1 is located in this interval (Fig. 1b,c). Mutation analysis in the affected family showed a single base-pair change resulting in an amino-acid substitution of serine for glycine at position 59 (G59S) in all affected family members (Fig. 2a). We did not find this mutation in the unaffected family members or in 200 unaffected control individuals of European descent, and there was no variation in the remaining transcribed sequence of DCTN1 in the affected family.
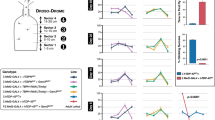
a, Centromeric region of chromosome 2. The centromere is indicated as a black oval. Recombinations with markers D2S291 and D2S2114 defined the critical region of 4 Mb at chromosome 2p13. b, The region of interest contains 44 genes of known and unknown function. Selected candidate genes are included in the figure: early growth response gene (EGR4), dual specific phosphatase (DUSP), dynactin 1 (DCTN1), rhotekin (RTKN) and semaphorin 4F (SEMA4F). The polymorphic markers indicated in black were used for multipoint analysis. c, Exon–intron structure of DCTN1. The exons are shown in red; the first exon is partially translated. The second exon, which contains the mutation resulting in the amino-acid substitution G59S, is indicated by an arrow. See Supplementary Note 1 online for methods.
a, Partial nucleotide sequence of exon 2 from the wild-type (lower panel) and mutated (upper panel) alleles. The heterozygous mutation C→T at nucleotide 957 is indicated by an arrow. b, Conservation of the mutated region across species. The position of the amino-acid change from glycine to serine is indicated in bold. c, Modeling of the CAP-Gly domain of DCTN1. The wild-type structure is shown on the left, and the G59S mutant is modeled on the right. Gly59 is closely packed with Phe88. The steric hindrance introduced by the larger side chain of serine would probably distort the folding13,14,15. d, The G59S mutation reduces the affinity of DCTN1 for microtubules. In the presence of microtubules (MTs), 86 ± 1% (n = 4) of the wild-type (wt) protein pelleted (P) with microtubules and 14% remained soluble (S), whereas only 51 ± 3% (n = 4) of the G59S mutant bound to microtubules (upper panel). The lower panel shows the Coomassie-stained gel. See Supplementary Note 1 online for methods.
The G59S substitution occurs in the highly conserved CAP-Gly motif of the p150Glued subunit of dynactin, a domain that has been shown to bind directly to microtubules (ref. 6; Fig. 2b). Modeling studies using the recently solved crystal structure for an uncharacterized CAP-Gly protein from Caenorhabditis elegans7 suggest that Gly59 is not directly involved in binding to microtubules but rather is buried in the novel protein fold described for CAP-Gly proteins. Modeling of the G59S mutation suggests that the greater size of the serine side chain may cause steric hindrance (Fig. 2c) and distort the folding of the microtubule-binding domain. To test for binding to microtubules, we expressed 35S-labeled wild-type and G59S mutant constructs of DCTN1 spanning residues 1–333. In the presence of microtubules, 86±1% of the wild-type protein sedimented with microtubules, but only 51±1% of the mutant G59S protein bound to microtubules (Fig. 2d).
The dynactin complex is required for dynein-mediated retrograde transport of vesicles and organelles along microtubules. It provides a link between specific cargos, the microtubule and cytoplasmic dynein during vesicle transport8. Overexpression of dynamitin, the p50 subunit of dynactin, disrupts the complex and has recently been found to produce a late-onset, progressive motor neuron disease in transgenic mice9. Mutations in the Drosophila melanogaster homolog for DCTN1 (refs. 10,11) and in the gene encoding cytoplasmic dynein heavy chain12 produce a rough-eye phenotype with severe disruption in the organization of the retina and in the retinal axonal projections. The substantial but limited reduction in the affinity of G59S for microtubules is consistent with the late onset and relatively mild disease progression observed in this kindred. A mutation that more severely inhibits the ability of dynactin to bind to microtubules would probably result in a more pronounced degeneration of motor neurons, as is predicted by the transgenic mouse model in which dynamitin is overexpressed9.
Identification of the mutation in DCTN1 in the family reported here shows that dysfunction of a protein involved in retrograde transport can cause human motor neuron disease. Primary motor neurons may be particularly vulnerable to defects in axonal transport because of their unusual size and metabolic activity and their dependence on trophic factors delivered from the periphery. Further studies may indicate whether this motor neuron disease is caused by impaired transportation of one or more specific factors or by a more general effect. Other members of the dynactin–dynein complex may now be considered candidates for disruption in other hereditary and sporadic disorders with motor neuron degeneration.
Note: Supplementary information is available on the Nature Genetics website.
References
Griffin, J.W. & Watson, D.F. Ann. Neurol. 23, 3–13 (1988).
Williamson, T.L. & Cleveland, D.W. Nat. Neurosci. 2, 50–56 (1999).
Dupuis, L. et al. Neurobiol. Dis. 7, 274–285 (2000).
Piccioni, F. et al. FASEB J. 16, 1418–1420 (2002).
Zhao, C. et al. Cell 105, 587–597 (2001).
Waterman-Storer, C.M., Karki, S. & Holzbaur, E.L. Proc. Natl. Acad. Sci. USA 92, 1634–1638 (1995).
Li, S. et al. J. Biol. Chem. 277, 48596–48601 (2002).
Waterman-Storer, C.M. et al. Proc. Natl. Acad. Sci. USA 94, 12180–12185 (1997).
LaMonte, B.H. et al. Neuron 34, 715–727 (2002).
Waterman-Storer, C.M. & Holzbaur, E.L. J. Biol. Chem. 271, 1153–1159 (1996).
Harte, P.J. & Kankel, D.R. Dev. Biol. 99, 88–102 (1983).
Gepner, J. et al. Genetics 142, 865–878 (1996).
Guex, N. & Peitsch, M.C. Electrophoresis 18, 2714–2723 (1997).
Peitsch, M.C. Biochem. Soc. Trans. 24, 274–279 (1996).
Peitsch, M.C. Biotechnology 13, 658–660 (1995).
Acknowledgements
We thank the participating family members for their cooperation throughout the study and Y. Zhang, A. Singleton, D. Hernandez, J.W. Nagle and Y. Goldman for their help. This work was supported in part by grants from the Amyotrophic Lateral Sclerosis Association and the US National Institutes of Health to E.L.F.H. R.H.B. received support from the US National Institutes of Health (National Institute of Neurologic Disorders and Stroke and National Institute on Aging), Amyotrophic Lateral Sclerosis Association, Project ALS, the Muscular Dystrophy Association, the Angel Fund for ALS research and the Al Athel ALS Research program.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Puls, I., Jonnakuty, C., LaMonte, B. et al. Mutant dynactin in motor neuron disease. Nat Genet 33, 455–456 (2003). https://doi.org/10.1038/ng1123
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng1123
This article is cited by
-
A novel DCTN1 mutation causing perry syndrome leads to abnormal splicing of mRNA: genetic and functional analyses
Acta Neurologica Belgica (2024)
-
Genetics of amyotrophic lateral sclerosis: seeking therapeutic targets in the era of gene therapy
Journal of Human Genetics (2023)
-
Clinical genetics of Charcot–Marie–Tooth disease
Journal of Human Genetics (2023)
-
Deficiency of Perry syndrome-associated p150Glued in midbrain dopaminergic neurons leads to progressive neurodegeneration and endoplasmic reticulum abnormalities
npj Parkinson's Disease (2023)
-
Developmental demands contribute to early neuromuscular degeneration in CMT2D mice
Cell Death & Disease (2020)