Abstract
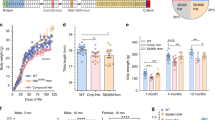
Dystonia musculorum (dt) is a hereditary neurodegenerative disease in mice that leads to a sensory ataxia. We describe cloning of a candidate dt gene, dystonin, that is predominantly expressed in the dorsal root ganglia and other sites of neurodegeneration in dt mice. Dystonin encodes an N–terminal actin binding domain and a C–terminal portion comprised of the hemidesmosomal protein, bullous pemphigoid antigen 1 (bpagl). dt and bpag1 are part of the same transcription unit which is partially deleted in a transgenic strain of mice, Tg4, that harbours an insertional mutation at the dt locus, and in mice that carry a spontaneous dt mutation, dtAlb. We also demonstrate abnormal dystonin transcripts in a second dt mutant, dt24J. We conclude that mutations in the dystonin gene are the primary genetic lesion in dt mice.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Duchen, L.W., Falconer, D.S. & Strich, S.J., A hereditary neuropathy of mice affecting mainly sensory pathways. J. Physiol. 165, 7–9 (1963).
Duchen, L.W. & Strich, S.J. Clinical and pathological studies of an hereditary neuropathy in mice. Brain 87, 367–378 (1964).
Duchen, L.W. Dystonia musculorum — an inherited disease of the nervous system in the mouse. Adv. Neurol. 14, 353–365 (1976).
Hanker, J.S. & Peach, R. Histochemical and ultrastructural studies of primary sensory neurons in mice with dystonia musculorum. I. Acetylcholinesteraseand lysosomal hydrolases. Neuropath. app. Neurobiol. 2, 79–97 (1976).
Janota, I. Ultrastructural studies of an hereditary sensory neuropathy in mice (dystonia musculorum). Brain 95, 529–536 (1972).
Sotelo, C. & Guenet, J.L. Pathologic changes in the CMS of dystonia musculorum mutant mouse: an animal model for human spinocerebellar ataxia. Neuroscience 27, 403–424 (1988).
AI-Ali, S.Y. & Al-Zuhar, A.G.H. Fine structural study of the spinal cord and spinal ganglia in mice afflicted with a hereditary sensory neuropathy, dystonia musculorum. J. submicr. Cytol. Pathol. 21, 737–748 (1989).
Messer, A. & Strominger, N.L. An allele of the mouse mutant dystonia musculorum exhibits lesions in red nucleus and striatum. Neuroscience 5, 543–549 (1980).
Kothary, R. et al. A transgene containing lacZ inserted into the dystonia locus is expressed in neural tube. Nature 335, 435–437 (1988).
Brown, A. et al. The genomic structure of an insertional mutation in the Dystonia musculorum locus. Genomics 20, 371–376 (1994).
Roberds, L. GRAIL seeks out genes buried in DMA sequence. Science 254, 805 (1991).
Frohman, A.M., Dush, M.K. & Martin, G.R. Rapid amplification of full-length cDNAs from rare transcripts: amplification using a single gene-specific oligonucleotide primer. Proc. Natn. Acad. Sci. U.S.A. 85, 8998–9002 (1988).
Loh, E.Y., Elliott, J.F., Cwirla, S., Lanier, L.L. & Davis, M.M. Polymerasechain reaction with single-sided specificity: analysis of T-cell receptor δ-chain. Science 258, 955–964 (1989).
Koenig, M. et al. Complete cloning of the Duchenne muscular dystrophy (DMD) cDNA and preliminary genomic organization of the DMD gene in normal and affected individuals. Cell 50, 509–517 (1987).
Dubreuil, R.R., Byers, T.J., Stewart, C.T. & Kiehart, D.P. A β-spectrin isoform from Drosophila (beta H) is similar in size to vertebrate dystrophin. J. Cell Biol. 111, 1849–1858 (1990).
Witke, W., Schleicher, M., Lottspeich, F. & Noegel, A. Studies on the transcription, translation, and structure of α-actinin in Dictyostelium discoideum. J. Cell Biol. 103, 969–975 (1986).
Noegel, A., Witke, W. & Schleicher, M. Calcium-sensitive non-muscle α-actinin contains EF-hand structures and highly conserved regions. FEBS Lett. 221, 391–396 (1987).
Vandekerckhove, J. & Vancompernolle, K. Structural relationships of actin binding proteins. Curr. Opin. Cell Biol. 4, 36–42 (1992).
Frappier, T., Derancourt, J. & Pradel, L.-A. Actin and neurofilament binding domain of brain spectrin β-subunit. Eur. J. Biochem. 205, 85–91 (1992).
Copeland, N.G. et al. Chromosomal localization of mouse bulbus pemphigoid antigens, BPAG1 and BPAG2: Identification of a new region of homology between mouse and human chromosomes. Genomics. 15, 180–181 (1993).
Brown, A., Lemieux, N., Rossant, J. & Kothary, R. Human homolog of a mouse sequence from the dystonia musculorum locus is on chromosome 6p12. Mamm. Genome 5, 434–437 (1994).
Stanley, J.R., Tanaka, T., Mueller, S., Klaus-Kovtun, V. & Roop, D. Isolation of cDNA for bullous pemphigoid antigen by use of patients' autoantibodies. J. Clin. Invest. 82, 1864–1870 (1988).
Sawamura, D., Li, K.-H., Chu, M.-L. & Uitto, J., Antigen (BPAG1): Amino acid sequences deduced from cloned cDNAs predict biologically important peptide segments and protein domains. J. biol. Chem. 266, 17784–17790 (1991).
Tamai, K. et al. The human 230-kD Bullous pemphigoid antigen gene (BPAG1). Exon-lntron organization and identification of regulatory tissue specific elements in the promoter region. J. Clin. Invest. 92, 814–822 (1993).
Tamai, K., Li, K. & Uitto, J. Identification of a DNA-binding protein (Keratinocyte Transcriptional Protein-1) recognizing a keratinocyte-specific regulatory element in the 230-kDa Bullous pemphigoid antigen gene. J. biol. Chem. 289, 493–502 (1994).
Sawamura, D. et al. Mouse 230-kDa bullous pemphigoid antigen gene : structural and functional characterization of the 5′-flanking region and interspecies conservation of the deduced amino-terminal peptide sequence of the protein. J. invest. Dermatol. 103, 651–655 (1994).
Kaneko, T. & LePage, G.A. Growth characteristics and drug responses of a murine lung carcinoma in vitro and in vivo. Cancer Res. 38, 2084–2090 (1978).
Luna, E.J. & Hitt, A.L. Cytoskeleton-plasma membrane interactions. Science 258, 955–964 (1992).
Palek, J. & Sahr, E. Mutations of the red blood cell membrane proteins: from clinical evaluation to detection of the underlying genetic defect. Blood 80, 308–330 (1992).
Ann, A.H. & Kunkel, L.M. The structural and functional diversity of dystrophin. Nature Genet. 3, 283–291 (1993).
Sealock, R. & Froehner, S.C. Dystrophin-associated proteins and synapse formation: Is α-dystroglycan the agrin receptor? Cell 77, 617–619 (1994).
Gee, S.H., Montanaro, F., Lindenbaum, M.H., Carbonetto, S. Dystroglycan-α, a dystrophin-associated glycoprotein, is a functional agrin receptor. Cell 77, 675–686 (1994).
Campanelli, J.T., Roberds, S.L., Campbell, K.P. & Scheller, R.H. A role for dystrophin-associated glycoproteins and utrophin in agrin-induced AChr clustering. Cell 77, 663–674 (1994).
Hoffman, E.P., Brown, R.H. & Kunkel, L.M. Dystrophin: the protein product of the Duchenne muscular dystrophy locus. Cell 51, 919–928 (1987).
Mutasim, D.F. et al. A pool of bullous pemphigoid antigens isintracellularand associated with the basal cell cytoskeleton-hemidesmosome complex. J. Invest. Dermatol. 84, 47–53 (1985).
Klatte, D.H., Kurpakus, M.A., Grelling, K.A. & Jones, J.C.R. Immunochemical characterization of three components of the hemidesmosome and their expression in cultured epithelial cells. J. Cell Biol. 109, 3377–3390 (1989).
Guo, F. et al. Gene targeting of bpag1 abnormalities in mechanical strength and cell migration in stratified epithelia and neurologic degeneration. Cell 81, 233–243 (1995).
Goldman, J.E. & Yen, S.H. Cytoskeletal protein abnormalities in neurodegenerative diseases. Ann. Neurol. 19, 209–223 (1986).
Campbell, R.M. & Peterson, A.C. An intrinsic neuronal defect operates in dystonia musculorum: a study of dt/dt<>+/+ chimeras. Neuron 9, 693–703 (1992).
Xu, Z., Cork, L.C., Griffin, J.W. & Cleveland, D.W. Increased expression of neurofilament subunit NF-L produces morphological alterations that resemble the pathology of human motor neuron disease. Cell 73, 23–34 (1993).
Coté, F., Collard, J.F. & Julien, J.-P. Progressive neuropathy in transgenic mice expressing the human neurofilament heavy chain: a mouse model of amyotrophic lateral sclerosis. Cell 73, 35–46 (1993).
Lee, M.K., Marszalek, J.R. & Cleveland, D.W. A mutant neurofilament subunit causes massive, selective motor neuron death: implications for the pathogenesis of human motor neuron disease. Neuron 13, 975–988 (1994).
Hildebrand, C. Ultrastructural and light microscopic studies of the developing feline spinal cord white matter.II. Cell death and myelin sheath disintegration in the early postnatal period. Acta. Physiol. Scand. (suppl.). 364, 109–144 (1971).
McLeod, J.G. An electrophysiological and pathological study of peripheral nerves in Friedreich's ataxia. J. neurol. Sci. 12, 333–349 (1971).
Ouvrier, R.A., McLeod, J.G. & Conchin, T.E. Friedreich's ataxia. Early detection and progression of peripheral nerve abnormalities. J. neurol. Sci. 55, 137–145 (1982).
McLeod, J.G. & Evans, W.A. Peripheral neuropathy in spinocerebellar degenerations. Muscle & Nerve 4, 51–61 (1981).
Harding, A.E. The Hereditary Ataxias and Related Disorders (Churchill Livingston, New York, 1984).
Koskinen, T. et al. Infantile onset spinocerebellar ataxia with sensory neuropathy: a new inherited disease. J. neurol. Sci. 121, 50–56 (1994).
Koskinen, T., Sainio, K., Rapola, J., Pihko, H. & Paetau, A. Sensory neuropathy in infantile onset spinocerebellar ataxia (IOSCA). Muscle & Nerve 17, 509–515 (1994).
Sawamura, D. et al. Bullous pemphigoid antigen: cDNA cloning and mapping of the gene to the short arm of human chromosome 6. Genomics 8, 722–726 (1990).
Hanauer, A. et al. The Friedreich's ataxia gene is assigned to chromosome 9q13–q21 by mapping of tightly linked markers and shows linkage disequilibrium with D9S15. Am. J. Hum. Genet. 46, 133–137 (1990).
Orr, H.T. et al. Expansion of an unstable trinucleotide CAG repeat in spinocerebellar ataxia type 1. Nature Genet. 4, 221–226 (1993).
Gispert, S. et al. Chromosomal assignment of the second locus for autosomal dominant cerebellar ataxia (SCA2) to chromosome 12q23–24.1. Nature Genet. 4, 295–299 (1993).
Takiyama, Y. et al. The gene for Machado-Joseph disease maps to human chromosome 14q. Nature Genet. 4, 300–304 (1993).
Ranum, L., Schut, L.J., Lundgren, J.K., Orr, H.T. & Livingston, D.M. Spinocerebellar ataxia type 5 in a family descended from the grandparents of President Lincoln maps to chromosome 11. Nature Genet. 8, 280–284 (1994).
Muller, U., Graeber, M.B., Haberhausen, G. & Kohler, A. Molecular basis and diagnosis of neurogenetic disorders. J. neurol. Sci. 124, 119–140 (1994).
Subramony, S.H. Degenerative ataxias. Curr. Opin. Neurol. 7, 316–322 (1994).
Lopes, C.I., Andermann, E. & Rouleau, G.A. Evidence for the existence of a fourth dominantly inherited spinocerebellar ataxia locus. Genomics 21, 270–274 (1994).
Hogan, B., Costantini, F. & Lacy, E. Manipulating the Mouse Embryo. A Laboratory Manual (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, 1986).
Altschul, S.F., Gish, W., Miller, W., Myers, E.W. & Lipman, D.J. Basic local alignment search tool. J. molec. Biol. 215, 403–410 (1990).
Sambrook, J., Fritsch, E.F. & Maniatis, T., Cloning: A Laboratory Manual. (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, 1989).
Tso, J.Y., Sun, X.H., Kao, T.H., Reece, K.S. & Wu, R. Isolation and characterization of rat and human glyceraldehyde-3-phosphate dehydrogenase cDNAs: genomic complexity and molecular evolution of the gene. Nucl. Acids Res. 13, 2485–2502 (1985).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Brown, A., Bernier, G., Mathieu, M. et al. The mouse dystonia musculorum gene is a neural isoform of bullous pemphigoid antigen 1. Nat Genet 10, 301–306 (1995). https://doi.org/10.1038/ng0795-301
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/ng0795-301
This article is cited by
-
Roles of dystonin isoforms in the maintenance of neural, muscle, and cutaneous tissues
Anatomical Science International (2024)
-
Reduced Proliferation of Oligodendrocyte Progenitor Cells in the Postnatal Brain of Dystonia Musculorum Mice
Neurochemical Research (2018)
-
Subepidermal autoimmune bullous diseases: overview, epidemiology, and associations
Immunologic Research (2018)
-
Molecular architecture and function of the hemidesmosome
Cell and Tissue Research (2015)
-
Molecular architecture and function of the hemidesmosome
Cell and Tissue Research (2015)