Abstract
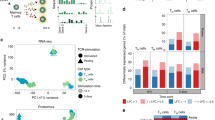
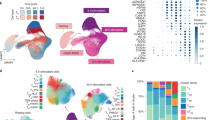
Many pathological processes, including those causing allergies and autoimmune diseases, are associated with the presence of specialized subsets of T helper cells at the site of inflammation1. Understanding the genetic program that controls the functional properties of T helper type 1 (Th1) versus T helper type 2 (Th2) cells may provide insight into the pathophysiology of inflammatory diseases. We compared the gene-expression profiles of human Th1 and Th2 cells using high-density oligonucleotide arrays with the capacity to display transcript levels of 6,000 human genes2. Here we analyse the data sets derived from five independent experiments using statistical algorithms. This approach resulted in the identification of 215 differentially expressed genes, encoding proteins involved in transcriptional regulation, apoptosis, proteolysis, and cell adhesion and migration. A subset of these genes was further upregulated by exposure of differentiated Th1 cells to interleukin-12 (IL-12), as confirmed by kinetic PCR analysis, indicating that IL-12 modulates the effector functions of Th1 cells in the absence of antigenic stimulation. Functional assays and in vivo expression of selected genes have validated the biological relevance of our study. Our results provide new insight into the transcriptional program controlling the functional diversity of subsets of T helper cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Romagnani, S. Lymphokine production by human T cells in disease states. Annu. Rev. Immunol. 12, 227–257 (1994).
Lipshutz, R.J., Fodor, S.P.A., Gingeras, T.R. & Lockhart, D.J. High density synthetic oligonucleotide arrays. Nature Genet. 21, 20–24 (1999).
Abbas, A.K., Murphy, K.M. & Sher, A. Functional diversity of helper T lymphocytes. Nature 383, 787–793 (1996).
O'Garra, A. Cytokines induce the development of functionally heterogenous T helper subsets. Immunity 8, 275–283 (1998).
Gately, M.K. et al. The interleukin-12/interleukin-12-receptor system: role in normal and pathologic immune responses. Annu. Rev. Immunol. 16, 495–521 (1998).
Seder, R.A. & Paul, W.E. Acquisition of lymphokine-producing phenotype by CD4+ T cells. Annu. Rev. Immunol. 12, 635–673 (1994).
Trinchieri, G. Interleukin-12: a proinflammatory cytokine with immunoregulatory functions that bridge innate resistance and antigen-specific adaptive immunity. Annu. Rev. Immunol. 13, 251–276 (1995).
Rogge, L. et al. Selective expression of an interleukin-12 receptor component by human T helper 1 cells. J. Exp. Med. 185, 825–832 (1997).
Rogge, L. et al. The role of Stat4 in species-specific regulation of T helper cell development by type I interferons. J. Immunol. 161, 6567–6574 (1998).
Higuchi, R. & Watson, R. Kinetic PCR analysis using a CCD camera and without using oligonucleotide probes. in PCR Applications (eds Innis, M.A., Gelfand, D.H. & Sninsky, J.J.) 263–284 (Academic, San Diego, 1999).
Zheng, W.-P. & Flavell, R.A. The transcription factor GATA-3 is necessary and sufficient for Th2 cytokine gene expression in CD4 T cells. Cell 89, 587–596 (1997).
Xu, D. et al. Selective expression and functions of interleukin 18 receptor on T helper (Th) type 1 but not Th2 cells. J. Exp. Med. 188, 1485–1492 (1998).
Taki, S. et al. Multistage regulation of Th1-type immune responses by the transcription factor IRF-1. Immunity 6, 673–679 (1997).
Lohoff, M. et al. Interferon regulatory factor-1 is required for a T helper 1 immune response in vivo. Immunity 6, 681–689 (1997).
Coccia, E.M. et al. IL-12 induces expression of interferon regulatory factor-1 via signal transducer and activator of transcription-4 in human T helper type 1 cells. J. Biol. Chem. 274, 6698–6703 (1999).
Austrup, F. et al. P- and E-selectin mediate recruitment of T-helper-1 but not T-helper-2 cells into inflamed tissues. Nature 385, 81–83 (1997).
Butcher, E.C. & Picker, L.J. Leukocyte homing and homeostasis. Science 272, 60–66 (1996).
Maly, P. et al. The α(1,3) fucosyltransferase Fuc-TVII controls leukocyte trafficking through an essential role in L-, E-, and P-selectin ligand biosynthesis. Cell 86, 643–653 (1996).
Sallusto, F., Lanzavecchia, A. & Mackay, C.R. Chemokines and chemokine receptors in T-cell priming and Th1/Th2-mediated responses. Immunol. Today 19, 568–574 (1998).
Van Damme, J. et al. The role of CD26/DPP IV in chemokine processing. Chem. Immunol. 72, 42–56 (1999).
Colantonio, L. et al. Upregulation of integrin α6/β1 and chemokine receptor CCR1 by interleukin-12 promotes the migration of human type 1 helper T cells. Blood 94, 2981–2989 (1999).
Berlin, C. et al. α4β7 integrin mediates lymphocyte binding to the mucosal vascular addressin MAdCAM-1. Cell 74, 185–195 (1993).
Simon, A.K., Seipelt, E. & Sieper, J. Divergent T-cell cytokine patterns in inflammatory arthritis. Proc. Natl Acad. Sci. USA 91, 8562–8566 (1994).
Dolhain, R.J.E.M., van der Heiden, A.N., ter Haar, N.T., Breedveld, F.C. & Miltenburg, A.M.M. Shift toward T lymphocytes with a T helper 1 cytokine-secretion profile in the joints of patients with rheumatoid arthritis. Arthritis Rheum. 39, 1961–1969 (1996).
Lockhart, D.J. et al. Expression monitoring by hybridization to high-density oligonucleotide arrays. Nature Biotechnol. 14, 1675–1680 (1996).
Fambrough, D., McClure, K., Kazlauskas, A. & Lander, E.S. Diverse signaling pathways activated by growth factor receptors induce broadly overlapping, rather than independent, sets of genes. Cell 97, 727–741 (1999).
Verzeletti, S. et al. Herpes simplex virus thymidine kinase gene transfer for controlled graft-versus-host disease and graft-versus-leukemia: clinical follow-up and improved new vectors. Hum. Gene Ther. 9, 2243–2251 (1998).
Coligan, J.E., Kruisbeek, A.M., Margulies, D.H., Shevach, E.M. & Strober, W. Current Protocols in Immunology (Greene & Wiley, New York, 1991).
Eisen, M.B., Spellman, P.T., Brown, P.O. & Botstein, D. Cluster analysis and display of genome-wide expression patterns. Proc. Natl Acad. Sci. USA 95, 14863–14868 (1998).
Acknowledgements
We thank P. Vigano for cord blood samples, and J. Sninsky and L. Adorini for helpful discussions and critical reading of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Rights and permissions
About this article
Cite this article
Rogge, L., Bianchi, E., Biffi, M. et al. Transcript imaging of the development of human T helper cells using oligonucleotide arrays. Nat Genet 25, 96–101 (2000). https://doi.org/10.1038/75671
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/75671
This article is cited by
-
NFAT primes the human RORC locus for RORγt expression in CD4+ T cells
Nature Communications (2019)
-
CD26 and Asthma: a Comprehensive Review
Clinical Reviews in Allergy & Immunology (2019)
-
Identification of global regulators of T-helper cell lineage specification
Genome Medicine (2015)
-
A first look at TH cell transcriptomes
Nature Reviews Immunology (2015)
-
Multiple correlations of mRNA expression and protein abundance in human cytokine profile
Molecular Biology Reports (2014)