Abstract
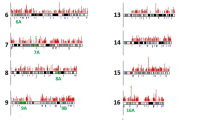
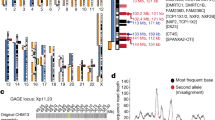
Trinucleotide microsatellites are widespread in the human and other mammalian genomes1. Expansions of unstable trinucleotide repeats have been associated so far with a number of different genetic diseases including fragile X, myotonic dystrophy (DM) and Huntington disease2–5. While ten possible trinucleotides can occur at the DMA level, only CTG and CCG repeats are involved in the disorders described so far. However, the repeat expansion detection (RED) technique6 has identified additional large repeats of ATG, CCT, CTT, and TGG of potentially pathological significance in the human genome7. We now show that conclusive information about the chromosomal localization of long trinucleotide repeats can be achieved in a relatively short time using fluorescence in situ hybridization (FISH) with biotin-labelled trinucleotide polymers. Large CTG expansions (>1 kb) in DM and an unstable (CTG)306 repeat in a patient with schizophrenia were detected by eye through the microscope without electronic enhancement. Digital imaging was used to analyse the chromosomal distribution of long CCA and AGG repeats. Our results suggest that long trinucleotide repeats occur in the normal human genome and that the size of individual repeat loci may be polymorphic.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Stallings, R.L. Distribution of trinucleotide microsatellites in different categories of mammalian genomic sequence: implications for human genetic diseases. Genomics. 21, 116–121 (1994).
Bates, G. & Lehrach, H. Trinucleotide repeat expansions and human genetic disease. BioEssays 16, 277–284 (1994).
Willems, P.J. Dynamic mutations hit double figures. Nature Genet. 8, 213–215 (1994).
Richards, R.I. & Sutherland, G.R. Simple repeat DNA is not replicated simply. Nature Genet. 6, 114–116 (1994).
Sutherland, G.R. & Richards, R.I. Simple tandem DNA repeats and human genetic disease. Proc. Natl. Acad. Sci. USA 92, 3636–3641 (1995).
Schalling, M., Hudson, T.J., Buetow, K.H. & Housman, D.E. Direct detection of novel expanded trinucleotide repeats in the human genome. Nature Genet. 4, 135–139 (1993).
Lindblad, K., Zander, C., Schalling, M. & Hudson, T. Growing triplet repeats. Nature Genet. 7, 124 (1994).
Sirugo, G. & Kidd, K.K. Repeat expansion detection using Ampligase thermostable DNA ligase. Epicentre Forum 2, 1–3 (1995).
Matera, A.G. & Ward, D.C. Oligonucleotide probes for the analysis of specific repetitive DNA sequences by fluorescence in situ hybridization. Hum. Molec. Genet. 1, 535–539 (1992).
Holmquist, G.P. Chromosome bands, their chromatin flavors, and their functional features. Am. J. Hum. Genet. 51, 17–37 (1992).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Haaf, T., Sirugo, G., Kidd, K. et al. Chromosomal localization of long trinucleotide repeats in the human genome by fluorescence in situ hybridization. Nat Genet 12, 183–185 (1996). https://doi.org/10.1038/ng0296-183
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/ng0296-183
This article is cited by
-
Anticipation and CAG•CTG repeat expansion in schizophrenia and bipolar affective disorder
Current Psychiatry Reports (2003)