Abstract
In eukaryotes, Suv39h H3K9 trimethyltransferases are required for pericentric heterochromatin formation and function. In early mouse preimplantation embryos, however, paternal pericentric heterochromatin lacks Suv39h-mediated H3K9me3 and downstream marks. Here we demonstrate Ezh2-independent targeting of maternally provided polycomb repressive complex 1 (PRC1) components to paternal heterochromatin. In Suv39h2 maternally deficient zygotes, PRC1 also associates with maternal heterochromatin lacking H3K9me3, thereby revealing hierarchy between repressive pathways. In Rnf2 maternally deficient zygotes, the PRC1 complex is disrupted, and levels of pericentric major satellite transcripts are increased at the paternal but not the maternal genome. We conclude that in early embryos, Suv39h-mediated H3K9me3 constitutes the dominant maternal transgenerational signal for pericentric heterochromatin formation. In absence of this signal, PRC1 functions as the default repressive back-up mechanism. Parental epigenetic asymmetry, also observed along cleavage chromosomes, is resolved by the end of the 8-cell stage—concurrent with blastomere polarization—marking the end of the maternal-to-embryonic transition.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Change history
16 March 2008
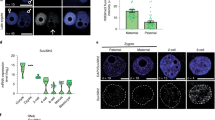
In the version of this article initially published online, the text in green associated with the right-most panel in Figure 1c should read ‘H3K27me3’, not ’H3K9me3’. The error has been corrected for all versions of the article.
References
Surani, M.A., Hayashi, K. & Hajkova, P. Genetic and epigenetic regulators of pluripotency. Cell 128, 747–762 (2007).
Arney, K.L., Bao, S., Bannister, A.J., Kouzarides, T. & Surani, M.A. Histone methylation defines epigenetic asymmetry in the mouse zygote. Int. J. Dev. Biol. 46, 317–320 (2002).
Dean, W. et al. Conservation of methylation reprogramming in mammalian development: aberrant reprogramming in cloned embryos. Proc. Natl. Acad. Sci. USA 98, 13734–13738 (2001).
Govin, J. et al. Pericentric heterochromatin reprogramming by new histone variants during mouse spermiogenesis. J. Cell Biol. 176, 283–294 (2007).
Liu, H., Kim, J.M. & Aoki, F. Regulation of histone H3 lysine 9 methylation in oocytes and early pre-implantation embryos. Development 131, 2269–2280 (2004).
Martin, C. et al. Genome restructuring in mouse embryos during reprogramming and early development. Dev. Biol. 292, 317–332 (2006).
Santos, F., Peters, A.H., Otte, A.P., Reik, W. & Dean, W. Dynamic chromatin modifications characterise the first cell cycle in mouse embryos. Dev. Biol. 280, 225–236 (2005).
van der Heijden, G.W. et al. Transmission of modified nucleosomes from the mouse male germline to the zygote and subsequent remodeling of paternal chromatin. Dev. Biol. 298, 458–469 (2006).
van der Heijden, G.W. et al. Asymmetry in histone H3 variants and lysine methylation between paternal and maternal chromatin of the early mouse zygote. Mech. Dev. 122, 1008–1022 (2005).
Merico, V. et al. Epigenomic differentiation in mouse preimplantation nuclei of biparental, parthenote and cloned embryos. Chromosome Res. 15, 341–360 (2007).
Kishigami, S. et al. Epigenetic abnormalities of the mouse paternal zygotic genome associated with microinsemination of round spermatids. Dev. Biol. 289, 195–205 (2006).
Ekwall, K. et al. Mutations in the fission yeast silencing factors clr4+ and rik1+ disrupt the localisation of the chromo domain protein Swi6p and impair centromere function. J. Cell Sci. 109, 2637–2648 (1996).
Peters, A.H. et al. Loss of the Suv39h histone methyltransferases impairs mammalian heterochromatin and genome stability. Cell 107, 323–337 (2001).
Peters, A.H. & Schubeler, D. Methylation of histones: playing memory with DNA. Curr. Opin. Cell Biol. 17, 230–238 (2005).
Wustmann, G., Szidonya, J., Taubert, H. & Reuter, G. The genetics of position-effect variegation modifying loci in Drosophila melanogaster. Mol. Gen. Genet. 217, 520–527 (1989).
Lachner, M., O'Carroll, D., Rea, S., Mechtler, K. & Jenuwein, T. Methylation of histone H3 lysine 9 creates a binding site for HP1 proteins. Nature 410, 116–120 (2001).
Grewal, S.I. & Jia, S. Heterochromatin revisited. Nat. Rev. Genet. 8, 35–46 (2007).
Lehnertz, B. et al. Suv39h-mediated histone H3 lysine 9 methylation directs DNA methylation to major satellite repeats at pericentric heterochromatin. Curr. Biol. 13, 1192–1200 (2003).
Schotta, G. et al. A silencing pathway to induce H3–K9 and H4–K20 trimethylation at constitutive heterochromatin. Genes Dev. 18, 1251–1262 (2004).
Ringrose, L. & Paro, R. Epigenetic regulation of cellular memory by the Polycomb and Trithorax group proteins. Annu. Rev. Genet. 38, 413–443 (2004).
Schwartz, Y.B. & Pirrotta, V. Polycomb silencing mechanisms and the management of genomic programmes. Nat. Rev. Genet. 8, 9–22 (2007).
Levine, S.S., King, I.F. & Kingston, R.E. Division of labor in polycomb group repression. Trends Biochem. Sci. 29, 478–485 (2004).
de Napoles, M. et al. Polycomb group proteins Ring1A/B link ubiquitylation of histone H2A to heritable gene silencing and X inactivation. Dev. Cell 7, 663–676 (2004).
Wang, H. et al. Role of histone H2A ubiquitination in Polycomb silencing. Nature 431, 873–878 (2004).
Sparmann, A. & van Lohuizen, M. Polycomb silencers control cell fate, development and cancer. Nat. Rev. Cancer 6, 846–856 (2006).
Boyer, L.A. et al. Polycomb complexes repress developmental regulators in murine embryonic stem cells. Nature 441, 349–353 (2006).
Bernstein, E. et al. Mouse polycomb proteins bind differentially to methylated histone H3 and RNA and are enriched in facultative heterochromatin. Mol. Cell. Biol. 26, 2560–2569 (2006).
Fujimura, Y. et al. Distinct roles of Polycomb group gene products in transcriptionally repressed and active domains of Hoxb8. Development 133, 2371–2381 (2006).
Guenatri, M., Bailly, D., Maison, C. & Almouzni, G. Mouse centric and pericentric satellite repeats form distinct functional heterochromatin. J. Cell Biol. 166, 493–505 (2004).
Probst, A.V., Santos, F., Reik, W., Almouzni, G. & Dean, W. Structural differences in centromeric heterochromatin are spatially reconciled on fertilisation in the mouse zygote. Chromosoma 116, 403–415 (2007).
Peters, A.H. et al. Partitioning and plasticity of repressive histone methylation states in mammalian chromatin. Mol. Cell 12, 1577–1589 (2003).
Mayer, W., Smith, A., Fundele, R. & Haaf, T. Spatial separation of parental genomes in preimplantation mouse embryos. J. Cell Biol. 148, 629–634 (2000).
O'Carroll, D. et al. The polycomb-group gene Ezh2 is required for early mouse development. Mol. Cell. Biol. 21, 4330–4336 (2001).
Fischle, W. et al. Regulation of HP1-chromatin binding by histone H3 methylation and phosphorylation. Nature 438, 1116–1122 (2005).
Leeb, M. & Wutz, A. Ring1B is crucial for the regulation of developmental control genes and PRC1 proteins but not X inactivation in embryonic cells. J. Cell Biol. 178, 219–229 (2007).
Baxter, J. et al. Histone hypomethylation is an indicator of epigenetic plasticity in quiescent lymphocytes. EMBO J. 23, 4462–4472 (2004).
Voncken, J.W. et al. Rnf2 (Ring1b) deficiency causes gastrulation arrest and cell cycle inhibition. Proc. Natl. Acad. Sci. USA 100, 2468–2473 (2003).
Lu, J. & Gilbert, D.M. Proliferation-dependent and cell cycle regulated transcription of mouse pericentric heterochromatin. J. Cell Biol. 179, 411–421 (2007).
Aoki, F., Worrad, D.M. & Schultz, R.M. Regulation of transcriptional activity during the first and second cell cycles in the preimplantation mouse embryo. Dev. Biol. 181, 296–307 (1997).
Rudolph, T. et al. Heterochromatin formation in Drosophila is initiated through active removal of H3K4 methylation by the LSD1 homolog SU(VAR)3–3. Mol. Cell 26, 103–115 (2007).
Sun, F. et al. Nuclear reprogramming: the zygotic transcription program is established through an “erase-and-rebuild” strategy. Cell Res. 17, 117–134 (2007).
Yoshida, N., Brahmajosyula, M., Shoji, S., Amanai, M. & Perry, A.C. Epigenetic discrimination by mouse metaphase II oocytes mediates asymmetric chromatin remodeling independently of meiotic exit. Dev. Biol. 301, 464–477 (2007).
Schoeftner, S. et al. Recruitment of PRC1 function at the initiation of X inactivation independent of PRC2 and silencing. EMBO J. 25, 3110–3122 (2006).
Umlauf, D. et al. Imprinting along the Kcnq1 domain on mouse chromosome 7 involves repressive histone methylation and recruitment of Polycomb group complexes. Nat. Genet. 36, 1296–1300 (2004).
Maison, C. et al. Higher-order structure in pericentric heterochromatin involves a distinct pattern of histone modification and an RNA component. Nat. Genet. 30, 329–334 (2002).
Chong, S. et al. Modifiers of epigenetic reprogramming show paternal effects in the mouse. Nat. Genet. 39, 614–622 (2007).
Johnson, M.H. Manipulation of early mammalian development: what does it tell us about cell lineages? Dev Biol (N Y 1985) 4, 279–96 (1986).
Okamoto, I., Otte, A.P., Allis, C.D., Reinberg, D. & Heard, E. Epigenetic dynamics of imprinted X inactivation during early mouse development. Science 303, 644–649 (2004).
Blewitt, M.E., Vickaryous, N.K., Paldi, A., Koseki, H. & Whitelaw, E. Dynamic reprogramming of DNA methylation at an epigenetically sensitive allele in mice. PLoS Genet. 2, e49 (2006).
Acknowledgements
We thank M. Vidal (Centro de Investigaciones Biológicas, Spain) and T. Jenuwein (Research Institute of Molecular Pathology, Austria) for providing antisera. Moreover, we are grateful to T. Jenuwein and B. Knowles (The Jackson Laboratory, USA) for providing Suv39h2 deficient and Zp3-cre transgenic mice, respectively. We acknowledge excellent assistance by Friedrich Miescher Institute (FMI) colleagues P. Schwarb and J. Rietdorf (microscopy and imaging facility), B. Heller-Stilb and J.-F. Spetz (animal facility), S. Bichet (histology) and M. Stadler (bioinformatics). We thank members of the Peters laboratory for fruitful discussions and P. de Boer, D. Schübeler, S. Gasser and P. Hublitz for valuable comments on the manuscript. Research at the Friedrich Miescher Institute is supported by the Novartis Research Foundation. M.P. acknowledges the Boehringer Ingelheim Fonds for her PhD fellowship. Research in the Peters laboratory is supported by the EU NoE network 'The Epigenome' (LSHG-CT-2004-503433).
Author information
Authors and Affiliations
Contributions
M.P. and A.H.F.M.P. conceived and designed the experiments. M.P., R.T., U.B. and C.K. performed the experiments. M.P., R.T., U.B. and A.H.F.M.P. analyzed the data. A.P.O. provided antibodies. E.B. and M.v.L. provided conditionally deficient Rnf2 mice. X.M. and S.H.O. provided conditionally deficient Ezh2 mice. K.I. and H.K. provided Rnf2–YFP knock-in mice. M.P. and A.H.F.M.P. wrote the manuscript.
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7, Supplementary Methods (PDF 2679 kb)
Rights and permissions
About this article
Cite this article
Puschendorf, M., Terranova, R., Boutsma, E. et al. PRC1 and Suv39h specify parental asymmetry at constitutive heterochromatin in early mouse embryos. Nat Genet 40, 411–420 (2008). https://doi.org/10.1038/ng.99
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.99
This article is cited by
-
Loss of H3K9 trimethylation alters chromosome compaction and transcription factor retention during mitosis
Nature Structural & Molecular Biology (2023)
-
LSM1-mediated Major Satellite RNA decay is required for nonequilibrium histone H3.3 incorporation into parental pronuclei
Nature Communications (2023)
-
Regulation, functions and transmission of bivalent chromatin during mammalian development
Nature Reviews Molecular Cell Biology (2023)
-
Unreprogrammed H3K9me3 prevents minor zygotic genome activation and lineage commitment in SCNT embryos
Nature Communications (2023)
-
Transient Polycomb activity represses developmental genes in growing oocytes
Clinical Epigenetics (2022)