Abstract
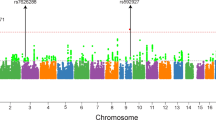
To identify susceptibility loci for schizophrenia, we performed a two-stage genome-wide association study (GWAS) of schizophrenia in the Han Chinese population (GWAS: 746 individuals with schizophrenia and 1,599 healthy controls; validation: 4,027 individuals with schizophrenia and 5,603 healthy controls). We identified two susceptibility loci for schizophrenia at 6p21-p22.1 (rs1233710 in an intron of ZKSCAN4, Pcombined = 4.76 × 10−11, odds ratio (OR) = 0.79; rs1635 in an exon of NKAPL, Pcombined = 6.91 × 10−12, OR = 0.78; rs2142731 in an intron of PGBD1, Pcombined = 5.14 × 10−10, OR = 0.79) and 11p11.2 (rs11038167 near the 5′ UTR of TSPAN18, Pcombined = 1.09 × 10−11, OR = 1.29; rs11038172, Pcombined = 7.21 × 10−10, OR = 1.25; rs835784, Pcombined = 2.73 × 10−11, OR = 1.27). These results add to previous evidence of susceptibility loci for schizophrenia at 6p21-p22.1 in the Han Chinese population. We found that NKAPL and ZKSCAN4 were expressed in postnatal day 0 (P0) mouse brain. These findings may lead to new insights into the pathogenesis of schizophrenia.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Lichtenstein, P. et al. Common genetic determinants of schizophrenia and bipolar disorder in Swedish families: a population-based study. Lancet 373, 234–239 (2009).
Thaker, G.K. & Carpenter, W. Advances in schizophrenia. Nat. Med. 7, 667–671 (2001).
Harrison, P.J. & Owen, M.J. Genes for schizophrenia? Recent findings and their pathophysiological implications. Lancet 361, 417–419 (2003).
Allen, N.C. et al. Systematic meta-analyses and field synopsis of genetic association studies in schizophrenia: the SzGene database. Nat. Genet. 40, 827–834 (2008).
O'Donovan, M.C. et al. Identification of loci associated with schizophrenia by genome-wide association and follow-up. Nat. Genet. 40, 1053–1055 (2008).
International Schizophrenia Consortium et al. Common polygenic variation contributes to risk of schizophrenia and bipolar disorder. Nature 460, 748–752 (2009).
Stefansson, H. et al. Common variants conferring risk of schizophrenia. Nature 460, 744–747 (2009).
Shi, J. et al. Common variants on chromosome 6p22.1 are associated with schizophrenia. Nature 460, 753–757 (2009).
Price, A.L. et al. Principal components analysis corrects for stratification in genome-wide association studies. Nat. Genet. 38, 904–909 (2006).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Pruim, R.J. et al. LocusZoom: regional visualization of genome-wide association scan results. Bioinformatics 26, 2336–2337 (2010).
Higgins, J.P. & Thompson, S.G. Quantifying heterogeneity in a meta-analysis. Stat. Med. 21, 1539–1558 (2002).
Mantel, N. & Haenszel, W. Statistical aspects of the analysis of data from retrospective studies. J. Natl. Cancer Inst. 22, 719–748 (1959).
Gibbs, J.R. et al. Abundant quantitative trait loci exist for DNA methylation and gene expression in human brain. PLoS Genet. 6, e1000952 (2010).
Richards, A.L. et al. Schizophrenia susceptibility alleles are enriched for alleles that affect gene expression in adult human brain. Mol. Psychiatry (in the press).
Pajerowski, A.G. et al. NKAP is a transcriptional repressor of Notch signaling and is required for T cell development. Immunity 30, 696–707 (2009).
Chen, D. et al. Identification of a nuclear protein that promotes NF-κB activation. Biochem. Biophys. Res. Commun. 310, 720–724 (2003).
Li, J. et al. ZNF307, a novel zinc finger gene suppresses p53 and p21 pathway. Biochem. Biophys. Res. Commun. 363, 895–900 (2007).
Bertram, L. & Tanzi, R.E. Genome-wide association studies in Alzheimer's disease. Hum. Mol. Genet. 18, R137–R145 (2009).
Strous, R.D. & Shoenfeld, Y. Schizophrenia, autoimmunity and immune system dysregulation: a comprehensive model updated and revisited. J. Autoimmun. 27, 71–80 (2006).
McGlashan, T.H. & Hoffman, R.E. Schizophrenia as a disorder of developmentally reduced synaptic connectivity. Arch. Gen. Psychiatry 57, 637–648 (2000).
Hemler, M.E. Tetraspanin proteins mediate cellular penetration, invasion, and fusion events and define a novel type of membrane microdomain. Annu. Rev. Cell Dev. Biol. 19, 397–422 (2003).
Berditchevski, F. & Odintsova, E. Tetraspanins as regulators of protein trafficking. Traffic 8, 89–96 (2007).
Sklar, P. et al. Whole-genome association study of bipolar disorder. Mol. Psychiatry 13, 558–569 (2008).
Chen, J. et al. Genetic structure of the Han Chinese population revealed by genome-wide SNP variation. Am. J. Hum. Genet. 85, 775–785 (2009).
Xu, S. et al. Genomic dissection of population substructure of Han Chinese and its implication in association studies. Am. J. Hum. Genet. 85, 762–774 (2009).
Zhang, F.R. et al. Genomewide association study of leprosy. N. Engl. J. Med. 361, 2609–2618 (2009).
Higgins, J.P. & Thompson, S.G. Quantifying heterogeneity in a meta-analysis. Stat. Med. 21, 1539–1558 (2002).
Higgins, J.P. et al. Measuring inconsistency in meta-analyses. Br. Med. J. 327, 557–560 (2003).
DerSimonian, R. & Laird, N. Meta-analysis in clinical trials. Control. Clin. Trials 7, 177–188 (1986).
Acknowledgements
We thank J. Liu from the Human Genetics, Genome Institute of Singapore for his suggestions for revision of the manuscript. We acknowledge with appreciation all the individuals with schizophrenia and healthy control subjects whose contributions made this work possible. This work was supported by research grants from the National High-Tech Research and Development Program of China (2009AA022702), the National Natural Science Foundation of China (30530290, 30870896, 81071087 and 81071088), the National Basic Research Program of China (2007CB512301, 2011CB707805) and the International Science & Technology Cooperation Program of China (2010DFB30820).
Author information
Authors and Affiliations
Contributions
D.Z., W.H., X.-J.Z., G.-P.Z. and T.L. designed the study. D.Z. and X.-J.Z. revised the manuscript. D.Z. and W.-H.Y. obtained financial support. W.-H.Y., L.-D.S. and L.-F.W. prepared the manuscript. H.-F.W., W.-H.Y. and L.-D.S. supervised the experiments and data analysis. H.-X.Z., W.-Q.L., Y.-L.Z., C.-C.M., B.D., Y.-Q.R., Y.-F.Y., X.-F.H., Y.W., W.D., L.-W.T., Y.-L.T., Q.C., G.-M.X., G.-G.Y., H.Y., Y.-Y.R., T.-L.L., X.H., X.-H.M., Y.W., L.-W.C., C.J., H.-Y.Z., J.Y., W.-F.M., X.-Y.Y., W.-B.M., Q.L., L.K., W.S., C.-Y.P., M.S., F.-D.Y., C.-Y.W., J.-L.Y., K.-Q.L., X.M., L.-J.L., X.Y. and L.-X.L. conducted sample selection and data management, undertook recruitment, collected phenotype data, undertook related data handling and calculation, managed recruitment and obtained biological samples. W.-H.Y., L.-F.W., X.B.Z. and Q.-Z.L. undertook data processing, statistical analysis and bioinformatics investigations. F.-L.T., Z.-H.L., Y.Z. and X.H. performed in situ hybridization and RNAi experiments. All authors critically reviewed the manuscript and approved the final version.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Tables 1–6, Supplementary Figures 1–6 and Supplementary Note (PDF 2238 kb)
Rights and permissions
About this article
Cite this article
Yue, WH., Wang, HF., Sun, LD. et al. Genome-wide association study identifies a susceptibility locus for schizophrenia in Han Chinese at 11p11.2. Nat Genet 43, 1228–1231 (2011). https://doi.org/10.1038/ng.979
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.979
This article is cited by
-
The genetic architecture of schizophrenia: review of large-scale genetic studies
Journal of Human Genetics (2023)
-
Genome-wide Mendelian randomization identifies actionable novel drug targets for psychiatric disorders
Neuropsychopharmacology (2023)
-
Screening properties of trend tests in genetic association studies
Scientific Reports (2023)
-
Involvement of the long intergenic non-coding RNA LINC00461 in schizophrenia
BMC Psychiatry (2022)
-
The schizophrenia-associated missense variant rs13107325 regulates dendritic spine density
Translational Psychiatry (2022)