Abstract
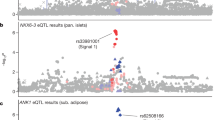
We carried out a genome-wide association study of type-2 diabetes (T2D) in individuals of South Asian ancestry. Our discovery set included 5,561 individuals with T2D (cases) and 14,458 controls drawn from studies in London, Pakistan and Singapore. We identified 20 independent SNPs associated with T2D at P < 10−4 for testing in a replication sample of 13,170 cases and 25,398 controls, also all of South Asian ancestry. In the combined analysis, we identified common genetic variants at six loci (GRB14, ST6GAL1, VPS26A, HMG20A, AP3S2 and HNF4A) newly associated with T2D (P = 4.1 × 10−8 to P = 1.9 × 10−11). SNPs at GRB14 were also associated with insulin sensitivity (P = 5.0 × 10−4), and SNPs at ST6GAL1 and HNF4A were also associated with pancreatic beta-cell function (P = 0.02 and P = 0.001, respectively). Our findings provide additional insight into mechanisms underlying T2D and show the potential for new discovery from genetic association studies in South Asians, a population with increased susceptibility to T2D.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Change history
16 September 2011
In the version of this article initially published online, Elin Grundberg’s name was misspelled as Elin Grunberg, and Xinzhong Li’s name was misspelled as Xinzhing Li. The error has been corrected for the print, PDF and HTML versions of the article.
References
Chambers, J.C. et al. Plasma homocysteine concentrations and risk of coronary heart disease in UK Indian Asian and European men. Lancet 355, 523–527 (2000).
Ramachandran, A., Ma, R.C. & Snehalatha, C. Diabetes in Asia. Lancet 375, 408–418 (2010).
Shaw, J.E., Sicree, R.A. & Zimmet, P.Z. Global estimates of the prevalence of diabetes for 2010 and 2030. Diabetes Res. Clin. Pract. 87, 4–14 (2010).
McCarthy, M.I. Genomics, type 2 diabetes, and obesity. N. Engl. J. Med. 363, 2339–2350 (2010).
Jowett, J.B. et al. Genetic influences on type 2 diabetes and metabolic syndrome related quantitative traits in Mauritius. Twin Res. Hum. Genet. 12, 44–52 (2009).
Chambers, J.C. et al. Genetic variation in SCN10A influences cardiac conduction. Nat. Genet. 42, 149–152 (2010).
Saleheen, D. et al. The Pakistan Risk of Myocardial Infarction Study: a resource for the study of genetic, lifestyle and other determinants of myocardial infarction in South Asia. Eur. J. Epidemiol. 24, 329–338 (2009).
Lavanya, R. et al. Methodology of the Singapore Indian Chinese Cohort (SICC) eye study: quantifying ethnic variations in the epidemiology of eye diseases in Asians. Ophthalmic Epidemiol. 16, 325–336 (2009).
Price, A.L. et al. Principal components analysis corrects for stratification in genome-wide association studies. Nat. Genet. 38, 904–909 (2006).
Sladek, R. et al. A genome-wide association study identifies novel risk loci for type 2 diabetes. Nature 445, 881–885 (2007).
Voight, B.F. et al. Twelve type 2 diabetes susceptibility loci identified through large-scale association analysis. Nat. Genet. 42, 579–589 (2010).
Kahn, S.E., Hull, R.L. & Utzschneider, K.M. Mechanisms linking obesity to insulin resistance and type 2 diabetes. Nature 444, 840–846 (2006).
1000 Genomes Project Consortium. et al. A map of human genome variation from population-scale sequencing. Nature 467, 1061–1073 (2010).
Heid, I.M. et al. Meta-analysis identifies 13 new loci associated with waist-hip ratio and reveals sexual dimorphism in the genetic basis of fat distribution. Nat. Genet. 42, 949–960 (2010).
Chambers, J.C. et al. Common genetic variation near MC4R is associated with waist circumference and insulin resistance. Nat. Genet. 40, 716–718 (2008).
Kooner, J.S. et al. Genome-wide scan identifies variation in MLXIPL associated with plasma triglycerides. Nat. Genet. 40, 149–151 (2008).
Coronary Artery Disease (C4D) Genetics Consortium. A genome-wide association study in Europeans and South Asians identifies five new loci for coronary artery disease. Nat. Genet. 43, 339–344 (2011).
Dufresne, A.M. & Smith, R.J. The adapter protein GRB10 is an endogenous negative regulator of insulin-like growth factor signaling. Endocrinology 146, 4399–4409 (2005).
Depetris, R.S., Wu, J. & Hubbard, S.R. Structural and functional studies of the Ras-associating and pleckstrin-homology domains of Grb10 and Grb14. Nat. Struct. Mol. Biol. 16, 833–839 (2009).
Holt, L.J. et al. Dual ablation of Grb10 and Grb14 in mice reveals their combined role in regulation of insulin signaling and glucose homeostasis. Mol. Endocrinol. 23, 1406–1414 (2009).
Woodard-Grice, A.V., McBrayer, A.C., Wakefield, J.K., Zhuo, Y. & Bellis, S.L. Proteolytic shedding of ST6Gal-I by BACE1 regulates the glycosylation and function of α4β1 integrins. J. Biol. Chem. 283, 26364–26373 (2008).
Maeda, N. et al. Diet-induced insulin resistance in mice lacking adiponectin/ACRP30. Nat. Med. 8, 731–737 (2002).
Siitonen, N. et al. Association of ADIPOQ gene variants with body weight, type 2 diabetes and serum adiponectin concentrations: the Finnish Diabetes Prevention Study. BMC Med. Genet. 12, 5 (2011).
Seaman, M.N., Marcusson, E.G., Cereghino, J.L. & Emr, S.D. Endosome to Golgi retrieval of the vacuolar protein sorting receptor, Vps10p, requires the function of the VPS29, VPS30, and VPS35 gene products. J. Cell Biol. 137, 79–92 (1997).
Seaman, M.N., Harbour, M.E., Tattersall, D., Read, E. & Bright, N. Membrane recruitment of the cargo-selective retromer subcomplex is catalysed by the small GTPase Rab7 and inhibited by the Rab-GAP TBC1D5. J. Cell Sci. 122, 2371–2382 (2009).
Kim, E. et al. Identification of novel retromer complexes in the mouse testis. Biochem. Biophys. Res. Commun. 375, 16–21 (2008).
Dell'Angelica, E.C. et al. AP-3: an adaptor-like protein complex with ubiquitous expression. EMBO J. 16, 917–928 (1997).
Brasaemle, D.L. et al. Perilipin A increases triacylglycerol storage by decreasing the rate of triacylglycerol hydrolysis. J. Biol. Chem. 275, 38486–38493 (2000).
Qi, L. et al. Genetic variation at the perilipin (PLIN) locus is associated with obesity-related phenotypes in White women. Clin. Genet. 66, 299–310 (2004).
Beller, M. et al. PERILIPIN-dependent control of lipid droplet structure and fat storage in Drosophila. Cell Metab. 12, 521–532 (2010).
Sumoy, L. et al. HMG20A and HMG20B map to human chromosomes 15q24 and 19p13.3 and constitute a distinct class of HMG-box genes with ubiquitous expression. Cytogenet. Cell Genet. 88, 62–67 (2000).
Artegiani, B. et al. The interaction with HMG20a/b proteins suggests a potential role for β-dystrobrevin in neuronal differentiation. J. Biol. Chem. 285, 24740–24750 (2010).
Battle, M.A. et al. Hepatocyte nuclear factor 4α orchestrates expression of cell adhesion proteins during the epithelial transformation of the developing liver. Proc. Natl. Acad. Sci. USA 103, 8419–8424 (2006).
Harries, L.W., Brown, J.E. & Gloyn, A.L. Species-specific differences in the expression of the HNF1A, HNF1B and HNF4A genes. PLoS ONE 4, e7855 (2009).
Yamagata, K. et al. Mutations in the hepatocyte nuclear factor-4α gene in maturity-onset diabetes of the young (MODY1). Nature 384, 458–460 (1996).
Teo, Y.Y. et al. Singapore Genome Variation Project: a haplotype map of three Southeast Asian populations. Genome Res. 19, 2154–2162 (2009).
Chidambaram, M., Radha, V. & Mohan, V. Replication of recently described type 2 diabetes gene variants in a South Indian population. Metabolism 59, 1760–1766 (2010).
Jafar, T.H. et al. Community-based interventions to promote blood pressure control in a developing country: a cluster randomized trial. Ann. Intern. Med. 151, 593–601 (2009).
Bellary, S. et al. Enhanced diabetes care to patients of South Asian ethnic origin (the United Kingdom Asian Diabetes Study): a cluster randomised controlled trial. Lancet 371, 1769–1776 (2008).
Rees, S.D. et al. An FTO variant is associated with type 2 diabetes in South Asian populations after accounting for body mass index and waist circumference. Diabet. Med. 28, 673–680 (2011).
Söderberg, S. et al. High incidence of type 2 diabetes and increasing conversion rates from impaired fasting glucose and impaired glucose tolerance to diabetes in Mauritius. J. Intern. Med. 256, 37–47 (2004).
Takeuchi, F. et al. Common variants at the GCK, GCKR, G6PC2-ABCB11 and MTNR1B loci are associated with fasting glucose in two Asian populations. Diabetologia 53, 299–308 (2010).
Sanghera, D.K. et al. The Khatri Sikh Diabetes Study (SDS): study design, methodology, sample collection, and initial results. Hum. Biol. 78, 43–63 (2006).
Katulanda, P. et al. Prevalence and projections of diabetes and pre-diabetes in adults in Sri Lanka–Sri Lanka Diabetes, Cardiovascular Study (SLDCS). Diabet. Med. 25, 1062–1069 (2008).
Marchini, J., Howie, B., Myers, S., McVean, G. & Donnelly, P. A new multipoint method for genome-wide association studies by imputation of genotypes. Nat. Genet. 39, 906–913 (2007).
Reich, D., Thangaraj, K., Patterson, N., Price, A.L. & Singh, L. Reconstructing Indian population history. Nature 461, 489–494 (2009).
Bacanu, S.A., Devlin, B. & Roeder, K. The power of genomic control. Am. J. Hum. Genet. 66, 1933–1944 (2000).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 26, 589–595 (2010).
Li, R. et al. SNP detection for massively parallel whole-genome resequencing. Genome Res. 19, 1124–1132 (2009).
Ong, R.T. & Teo, Y.Y. varLD: a program for quantifying variation in linkage disequilibrium patterns between populations. Bioinformatics 26, 1269–1270 (2010).
Myers, A.J. et al. A survey of genetic human cortical gene expression. Nat. Genet. 39, 1494–1499 (2007).
Webster, J.A. et al. Genetic control of human brain transcript expression in Alzheimer disease. Am. J. Hum. Genet. 84, 445–458 (2009).
Dixon, A.L. et al. A genome-wide association study of global gene expression. Nat. Genet. 39, 1202–1207 (2007).
Ge, B. et al. Global patterns of cis variation in human cells revealed by high-density allelic expression analysis. Nat. Genet. 41, 1216–1222 (2009).
Dimas, A.S. et al. Common regulatory variation impacts gene expression in a cell type-dependent manner. Science 325, 1246–1250 (2009).
Schadt, E.E. et al. Mapping the genetic architecture of gene expression in human liver. PLoS Biol. 6, e107 (2008).
Nica, A.C. et al. The architecture of gene regulatory variation across multiple human tissues: the MuTHER study. PLoS Genet. 7, e1002003 (2011).
Acknowledgements
We would like to thank the many colleagues who contributed to collection and phenotypic characterization of the clinical samples as well as genotyping and analysis of the GWAS data. We would also like to acknowledge those individuals who agreed to participate in these studies. Major funding for the work described in this paper comes from Wellcome Trust awards (070854/Z/03/Z, 080747/Z/06/Z, 083270/Z/07/Z, 084723/Z/08/Z); Chennai Wellingdon Corporate Foundation; Diabetes UK (07/0003512); National Institute for Health Research Comprehensive Biomedical Research Centre at Imperial College Healthcare NHS Trust; British Heart Foundation (SP/04/002); Medical Research Council (G0700931); National Institute for Health Research (RP-PG-0407-10371); US National Institutes of Health (DK-25446); KAKENHI (Grant-in-Aid for Scientific Research from the Ministry of Education, Culture, Sports, Science and Technology of Japan; National Center for Global Health and Medicine; US National Institutes of Health (KO1TW006087); National Institute of Diabetes and Digestive and Kidney Diseases (R01DK082766); A*STAR Biomedical Research Council (05/1/21/19/425); Biomedical Research Council Singapore (09/1/35/19/616, 08/1/35/19/550); National Medical Research Council Singapore (NMRC/STaR/0003/2008, 1174/2008); National Science Foundation of Sri Lanka; and Oxford NIHR Biomedical Research Centre. A full list of acknowledgments is provided in the Supplementary Note.
Author information
Authors and Affiliations
Consortia
Contributions
Manuscript preparation: J.S.K., D.S., X.S., J. Sehmi, W.Z., P.E., Y.Y.T., M.I.M., J.D., E.S.T. and J.C.C. wrote the manuscript. All authors read and provided critical comment on the manuscript. Data collection and analysis in the participating studies: COBRA study: T.J., M.I. and T.M.F. Chennai Urban Rural Epidemiology Study: V.R., M. Chidambaram, S.L. and V.M. Diabetes Genetics in Pakistan and UK Asian Diabetes Studies: S.D.R., A.B., Z.I.H., A.S.S., A.H.B. and M.A.K. London Life Sciences Population Study: J.S.K., W.Z., J. Sehmi, X.L., D.D., G.R.A., J. Scott, M. Caulfield, P. Froguel, P.E., M.I.M. and J.C.C. Mauritius study: J.B.M.J., S.K., M.M.K. and P.Z.Z. Pakistan Risk of Myocardial Infarction Study: D.S., P. Frossard, R.Y., A.R., M. Samuel, N.S., P.D. and J.D. Ragama Health Study: N.K., F.T., A.R.W. and J.M.P. Singapore Consortium of Cohort Studies: K.-S.C., W.-Y.L., C.-C.K., J. Liu and E.S.T. Sikh Diabetes Study: L.F.B. and D.K.S. Singapore Indian Eye Study: X.S., C.S., T.A., T.Y.W., M. Seielstad, Y.Y.T. and E.S.T. Sri Lankan Diabetes Study: N.H., I.P., D.R.M., P.K. and M.I.M. Sequencing of T2D loci: M.Y., F.Z., J. Liang, X.L., J.S.K. and J.C.C. Association results among Europeans in DIAGRAM: A.P.M. and M.I.M. eQTL analyses in MuTHER: A.S.D., E.G., Å.K.H., A.C.N., K.S.S. and M.I.M.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
A list of members is provided in the Supplementary Note.
A list of members is provided in the Supplementary Note.
Supplementary information
Supplementary Text and Figures
Supplementary Note, Supplementary Tables 1–15 and Supplementary Figures 1–8. (PDF 2904 kb)
Rights and permissions
About this article
Cite this article
Kooner, J., Saleheen, D., Sim, X. et al. Genome-wide association study in individuals of South Asian ancestry identifies six new type 2 diabetes susceptibility loci. Nat Genet 43, 984–989 (2011). https://doi.org/10.1038/ng.921
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.921
This article is cited by
-
South Asia: The Missing Diverse in Diversity
Behavior Genetics (2024)
-
Organ and cell-specific biomarkers of Long-COVID identified with targeted proteomics and machine learning
Molecular Medicine (2023)
-
CaMK1D signalling in AgRP neurons promotes ghrelin-mediated food intake
Nature Metabolism (2023)
-
Utility of genetic risk scores in type 1 diabetes
Diabetologia (2023)
-
Exploration of the Shared Hub Genes and Biological Mechanism in Osteoporosis and Type 2 Diabetes Mellitus based on Machine Learning
Biochemical Genetics (2023)