Abstract
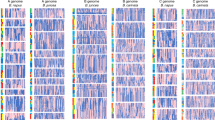
We report the annotation and analysis of the draft genome sequence of Brassica rapa accession Chiifu-401-42, a Chinese cabbage. We modeled 41,174 protein coding genes in the B. rapa genome, which has undergone genome triplication. We used Arabidopsis thaliana as an outgroup for investigating the consequences of genome triplication, such as structural and functional evolution. The extent of gene loss (fractionation) among triplicated genome segments varies, with one of the three copies consistently retaining a disproportionately large fraction of the genes expected to have been present in its ancestor. Variation in the number of members of gene families present in the genome may contribute to the remarkable morphological plasticity of Brassica species. The B. rapa genome sequence provides an important resource for studying the evolution of polyploid genomes and underpins the genetic improvement of Brassica oil and vegetable crops.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Tang, H. et al. Unraveling ancient hexaploidy through multiply-aligned angiosperm gene maps. Genome Res. 18, 1944–1954 (2008).
Johnston, J.S. et al. Evolution of genome size in Brassicaceae. Ann. Bot. 95, 229–235 (2005).
Koch, M.A. & Kiefer, M. Genome evolution among cruciferous plants: a lecture from the comparison of the genetic maps of three diploid species—Capsella rubella, Arabidopsis lyrata subsp Petraea, and A. thaliana. Am. J. Bot. 92, 761–767 (2005).
Yogeeswaran, K. et al. Comparative genome analyses of Arabidopsis spp.: inferring chromosomal rearrangement events in the evolutionary history of A. thaliana. Genome Res. 15, 505–515 (2005).
Bowers, J.E., Chapman, B.A., Rong, J. & Paterson, A.H. Unravelling angiosperm genome evolution by phylogenetic analysis of chromosomal duplication events. Nature 422, 433–438 (2003).
Yang, Y.W., Lai, K.N., Tai, P.Y. & Li, W.H. Rates of nucleotide substitution in angiosperm mitochondrial DNA sequences and dates of divergence between Brassica and other angiosperm lineages. J. Mol. Evol. 48, 597–604 (1999).
Town, C.D. et al. Comparative genomics of Brassica oleracea and Arabidopsis thaliana reveal gene loss, fragmentation, and dispersal after polyploidy. Plant Cell 18, 1348–1359 (2006).
Lysak, M.A., Koch, M.A., Pecinka, A. & Schubert, I. Chromosome triplication found across the tribe Brassiceae. Genome Res. 15, 516–525 (2005).
Labana, K.S. & Gupta, M.L. Importance and origin. in Breeding Oilseed Brassicas (eds. Labana, K.S., Banga, S.S. & Banga, S.K.) 1–20 (Springer-Verlag, Berlin, Germany, 1993).
Nagaharu, U. Genome analysis in Brassica with special reference to the experimental formation of B. napus and peculiar mode of fertilization. Jap. J. Bot. 7, 389–452 (1935).
O'Neill, C.M. & Bancroft, I. Comparative physical mapping of segments of the genome of Brassica oleracea var. alboglabra that are homoeologous to sequenced regions of chromosomes 4 and 5 of Arabidopsis thaliana. Plant J. 23, 233–243 (2000).
Park, J.Y. et al. Physical mapping and microsynteny of Brassica rapa ssp. pekinensis genome corresponding to a 222 kbp gene-rich region of Arabidopsis chromosome 4 and partially duplicated on chromosome 5. Mol. Genet. Genomics 274, 579–588 (2005).
Beilstein, M.A., Nagalingum, N.S., Clements, M.D., Manchester, S.R. & Mathews, S. Dated molecular phylogenies indicate a Miocene origin for Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA 107, 18724–18728 (2010).
Mun, J.H. et al. Genome-wide comparative analysis of the Brassica rapa gene space reveals genome shrinkage and differential loss of duplicated genes after whole genome triplication. Genome Biol. 10, R111 (2009).
Mun, J.H. et al. Sequence and structure of Brassica rapa chromosome A3. Genome Biol. 11, R94 (2010).
Arabidopsis Genome Initiative. Analysis of the genome sequence of the flowering plant Arabidopsis thaliana. Nature 408, 796–815 (2000).
Ming, R. et al. The draft genome of the transgenic tropical fruit tree papaya (Carica papaya Linnaeus). Nature 452, 991–996 (2008).
Jaillon, O. et al. The grapevine genome sequence suggests ancestral hexaploidization in major angiosperm phyla. Nature 449, 463–467 (2007).
Sankoff, D., Zheng, C. & Zhu, Q. The collapse of gene complement following whole genome duplication. BMC Genomics 11, 313 (2010).
Messing, J. et al. Sequence composition and genome organization of maize. Proc. Natl. Acad. Sci. USA 101, 14349–14354 (2004).
Schnable, J.C., Springer, N.M. & Freeling, M. Differentiation of the maize subgenomes by genome dominance and both ancient and ongoing gene loss. Proc. Natl. Acad. Sci. USA 108, 4069–4074 (2011).
Thomas, B.C., Pedersen, B. & Freeling, M. Following tetraploidy in an Arabidopsis ancestor, genes were removed preferentially from one homeolog leaving clusters enriched in dose-sensitive genes. Genome Res. 16, 934–946 (2006).
Woodhouse, M.R. et al. Following tetraploidy in maize, a short deletion mechanism removed genes preferentially from one of the two homologs. PLoS Biol. 8, e1000409 (2010).
Wang, X., Tang, H., Bowers, J.E. & Paterson, A.H. Comparative inference of illegitimate recombination between rice and sorghum duplicated genes produced by polyploidization. Genome Res. 19, 1026–1032 (2009).
Wang, X.Y., Tang, H.B. & Paterson, A.H. Seventy million years of concerted evolution of a homoeologous chromosome pair, in parallel, in major poaceae lineages. Plant Cell 23, 27–37 (2011).
Birchler, J.A. & Veitia, R.A. The gene balance hypothesis: from classical genetics to modern genomics. Plant Cell 19, 395–402 (2007).
Ha, M., Kim, E.D. & Chen, Z.J. Duplicate genes increase expression diversity in closely related species and allopolyploids. Proc. Natl. Acad. Sci. USA 106, 2295–2300 (2009).
Paterson, A.H., Lan, T.H., Amasino, R., Osborn, T.C. & Quiros, C. Brassica genomics: a complement to, and early beneficiary of, the Arabidopsis sequence. Genome Biol. 2, R1011 (2001).
Teale, W.D., Paponov, I.A. & Palme, K. Auxin in action: signalling, transport and the control of plant growth and development. Nat. Rev. Mol. Cell Biol. 7, 847–859 (2006).
Theologis, A. et al. Sequence and analysis of chromosome 1 of the plant Arabidopsis thaliana. Nature 408, 816–820 (2000).
Vanneste, S. & Friml, J. Auxin: a trigger for change in plant development. Cell 136, 1005–1016 (2009).
Reeves, P.A. & Olmstead, R.G. Evolution of the TCP gene family in Asteridae: cladistic and network approaches to understanding regulatory gene family diversification and its impact on morphological evolution. Mol. Biol. Evol. 20, 1997–2009 (2003).
Michaels, S.D. & Amasino, R.M. FLOWERING LOCUS C encodes a novel MADS domain protein that acts as a repressor of flowering. Plant Cell 11, 949–956 (1999).
Levy, Y.Y., Mesnage, S., Mylne, J.S., Gendall, A.R. & Dean, C. Multiple roles of Arabidopsis VRN1 in vernalization and flowering time control. Science 297, 243–246 (2002).
Günl, M., Liew, E.F., David, K. & Putterill, J. Analysis of a post-translational steroid induction system for GIGANTEA in Arabidopsis. BMC Plant Biol. 9, 141 (2009).
Li, D. et al. A repressor complex governs the integration of flowering signals in Arabidopsis. Dev. Cell 15, 110–120 (2008).
Ledger, S., Strayer, C., Ashton, F., Kay, S.A. & Putterill, J. Analysis of the function of two circadian-regulated CONSTANS-LIKE genes. Plant J. 26, 15–22 (2001).
Paterson, A.H. et al. The Sorghum bicolor genome and the diversification of grasses. Nature 457, 551–556 (2009).
Li, R. et al. De novo assembly of human genomes with massively parallel short read sequencing. Genome Res. 20, 265–272 (2010).
Li, R. et al. The sequence and de novo assembly of the giant panda genome. Nature 463, 311–317 (2010).
Delcher, A.L., Phillippy, A., Carlton, J. & Salzberg, S.L. Fast algorithms for large-scale genome alignment and comparison. Nucleic Acids Res. 30, 2478–2483 (2002).
Parkin, I.A. et al. Segmental structure of the Brassica napus genome based on comparative analysis with Arabidopsis thaliana. Genetics 171, 765–781 (2005).
Birney, E., Clamp, M. & Durbin, R. GeneWise and Genomewise. Genome Res. 14, 988–995 (2004).
Haas, B.J. et al. Improving the Arabidopsis genome annotation using maximal transcript alignment assemblies. Nucleic Acids Res. 31, 5654–5666 (2003).
Elsik, C.G. et al. Creating a honey bee consensus gene set. Genome Biol. 8, R13 (2007).
Li, L., Stoeckert, C.J. & Roos, D.S. OrthoMCL: identification of ortholog groups for eukaryotic genomes. Genome Res. 13, 2178–2189 (2003).
Tamura, K., Dudley, J., Nei, M. & Kumar, S. MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Mol. Biol. Evol. 24, 1596–1599 (2007).
Acknowledgements
This work was primarily funded by the Chinese Ministry of Science and Technology, Ministry of Agriculture, Ministry of Finance, the National Natural Science Foundation of China. Other funding sources included: Core Research Budget of the Non-profit Governmental Research Institution; the European Union 7th Framework Project; funds from Shenzhen Municipal Government of China; the Danish Natural Science Research Council; National Academy of Agricultural Science and the Next-Generation Biogreen21 Program, Rural Development Administration, Korea; the Technology Development Program for Agriculture and Forestry, Ministry for Food, Agriculture, Forestry and Fisheries, Korea; United Kingdom's Biotechnology and Biological Sciences Research Council; Institute National de la Recherche Agronomique, France; Japanese Kazusa DNA Research Institute Foundation; National Science Foundation, USA; Bielefeld University, Germany; the Australian Research Council; the Australian Grains Research and Development Corporation; Agriculture and Agri-Food Canada; and the National Research Council of Canada's Plant Biotechnology Institute. See the Supplementary Note for a full list of support and acknowledgments.
Author information
Authors and Affiliations
Consortia
Contributions
Principal investigators: Xiaowu Wang, J. Wu, S.L., Y.B., J.-H.M. and I.B. DNA and transcriptome sequencing: Bo Wang (group leader), Xiaowu Wang (group leader), B.C. (group leader), Jun Wang (BGI), K.W., J. Wu, S.L., W.H., B.-S.P., I.B., D.E., I.A.P.P., J.-H.M., H.A., Bernd Weisshaar, Shusei Sato, H.H., S.T., A.G.S., Y. Lim, G.B., J.B., C.L., C.G., J.P., S.-J.K., J.A.K., M.T., F.F., E.S., M.G.L., C.K., K.H., Y.N., P.J.B. and C.D. Sequence assembly: Junyi Wang (group leader), Jun Wang (BGI), D.M., Y. Li, X.X., Bo Liu, Silong Sun, Z.Z., Z.L., Binghang Liu, Q.C., Shu Zhang, Y.B., Zhiwen Wang, X.Z., C.S., J.Y. and J.J. Anchoring to linkage maps: J. Wu (group leader), W.H. (group leader), G.J.K., Y. Lim, B.-S.P., I.B., J.B., D.E., Yan Wang, Bo Liu, Silong Sun, Jun Wang (Rothamsted), I.A.P.P., J. Meng, Hui Wang, J.D., Y. Liao, Y.B., Haiping Wang, M.J., J.-S.K., S.-R.C., N.R. and A.H. Annotation: Y.B. (group leader), S.L. (group leader), R.L., W.F., Q.H., F.C., Bo Liu, D.E., J. Min, Jianwen Li, C.P., H.Z., Shunmou Huang, B.C., J.J., H.B., G.L., N.D. and M.T. Stabilizing the genome of a polyploidy dicotyledonous species: F.C. (group leader), Sanwen Huang (group leader), Y.B., Xiaowu Wang, B. Li, S.C., Y.Y., J.X. and C.T. Comparative genomics: Xiaowu Wang (group leader), J.C.P. (group leader), Xiyin Wang (group leader), I.B., F.C., H.T., G.C., H.G., T.-H.L., Jinpeng Wang and Zhenyi Wang. Retention of genes duplicated by polyploidy: M.F. (group leader), A.H.P. (group leader), F.C., H.T., Bo Liu, Silong Sun, L.F., Z.X., M.Z., Jingping Li, H.J. and X.T. Characteristics of a crop genome: J. Wu (group leader), X.L. (group leader), R.S., Hanzhong Wang, Y.D., Xiaowu Wang, Hui Wang, J.D., D.S., Y.Q., Shujiang Zhang, F.L., L.W. and Yupeng Wang.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Note, Supplementary Tables 1–21 and Supplementary Figures 1–25. (PDF 3166 kb)
Rights and permissions
About this article
Cite this article
The Brassica rapa Genome Sequencing Project Consortium., Wang, X., Wang, H. et al. The genome of the mesopolyploid crop species Brassica rapa. Nat Genet 43, 1035–1039 (2011). https://doi.org/10.1038/ng.919
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.919
This article is cited by
-
Efficient transformation of the isolated microspores of Chinese cabbage (Brassica rapa L. ssp. pekinensis) by particle bombardment
Plant Methods (2024)
-
Genome dosage alteration caused by chromosome pyramiding and shuffling effects on karyotypic heterogeneity, reproductive diversity, and phenotypic variation in Zea–Tripsacum allopolyploids
Theoretical and Applied Genetics (2024)
-
Mechanisms of salinity tolerance and their possible application in the breeding of vegetables
BMC Plant Biology (2023)
-
Identification, evolution and expression analyses of the whole genome-wide PEBP gene family in Brassica napus L.
BMC Genomic Data (2023)
-
Genome-wide identification of the geranylgeranyl pyrophosphate synthase (GGPS) gene family involved in chlorophyll synthesis in cotton
BMC Genomics (2023)