Abstract
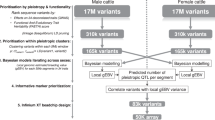
We report mapping of a quantitative trait locus (QTL) with a major effect on bovine stature to a ∼780-kb interval using a Hidden Markov Model–based approach that simultaneously exploits linkage and linkage disequilibrium. We re-sequenced the interval in six sires with known QTL genotype and identified 13 clustered candidate quantitative trait nucleotides (QTNs) out of >9,572 discovered variants. We eliminated five candidate QTNs by studying the phenotypic effect of a recombinant haplotype identified in a breed diversity panel. We show that the QTL influences fetal expression of seven of the nine genes mapping to the ∼780-kb interval. We further show that two of the eight candidate QTNs, mapping to the PLAG1-CHCHD7 intergenic region, influence bidirectional promoter strength and affect binding of nuclear factors. By performing expression QTL analyses, we identified a splice site variant in CHCHD7 and exploited this naturally occurring null allele to exclude CHCHD7 as single causative gene.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Stern, D.L. & Orgogozo, V. Is genetic evolution predictable? Science 323, 746–751 (2009).
Manolio, T.A. et al. Finding the missing heritability of complex diseases. Nature 461, 747–753 (2009).
Schadt, E.E. Molecular networks as sensors and drivers of common human diseases. Nature 461, 218–223 (2009).
Galton, F. Regression towards mediocrity in hereditary stature. J. R. Anthropol. Inst. 5, 329–348 (1885).
Fisher, R.A. The correlation between relatives on the supposition of Mendelian inheritance. Trans. R. Soc. Edinb. 52, 399–433 (1918).
Visscher, P.M. et al. Genome partitioning of genetic variation for height from 11,214 sibling pairs. Am. J. Hum. Genet. 81, 1104–1110 (2007).
Weedon, M.N. & Frayling, T.M. Reaching new heights: insights into the genetics of human stature. Trends Genet. 24, 595–603 (2008).
Lango Allen, H. et al. Hundreds of variants clustered in genomic loci and biological pathways affect human height. Nature 467, 832–838 (2010).
Yang, J. et al. Common SNPs explain a large proportion of the heritability for human height. Nat. Genet. 42, 565–569 (2010).
Sutter, N.B. et al. A single IGF1 allele is a major determinant of small size in dogs. Science 316, 112–115 (2007).
Boyko, A.R. et al. A simple genetic architecture underlies morphological variation in dogs. PLoS Biol. 8, e1000451 (2010).
Nelsen, T.C. et al. Heritabilities and genetic correlations of growth and reproductive measurements in Hereford bulls. J. Anim. Sci. 63, 409–417 (1986).
Northcutt, S.L. & Wilson, D.E. Genetic parameter estimates and expected progeny differences for mature size in Angus cattle. J. Anim. Sci. 71, 1148–1153 (1993).
McClure, M.C. et al. A genome scan for quantitative trait loci influencing carcass, post-natal growth and reproductive traits in commercial Angus cattle. Anim. Genet. 41, 597–607 (2010).
Arias, J.A., Keehan, M., Fisher, P., Coppieters, W. & Spelman, R. A high density linkage map of the bovine genome. BMC Genet. 10, 18 (2009).
Haley, C.S. et al. Mapping quantitative trait loci in crosses between outbred lines using least squares. Genetics 136, 1195–1207 (1994).
Coppieters, W. et al. A rank-based non parametric method to map QTL in outbred half-sib pedigrees: application to milk production in a grand-daughter design. Genetics 149, 1547–1555 (1998).
Visscher, P.M. et al. Confidence intervals in QTL mapping by bootstrapping. Genetics 143, 1013–1020 (1996).
Mizoshita, K. et al. Quantitative trait loci analysis for growth and carcass traits in a half-sib family of purebred Japanese Black (Wagyu) cattle. J. Anim. Sci. 82, 3415–3420 (2004).
Takasuga, A. et al. Identification of bovine QTL for growth and carcass traits in Japanese Black cattle by replication and identical-by-descent mapping. Mamm. Genome 18, 125–136 (2007).
Buchanan, F.C. et al. Single nucleotide polymorphisms in the corticotrophin-releasing hormone and pro-opiomelancortin genes are associated with growth and carcass yield in beef cattle. Anim. Genet. 36, 127–131 (2005).
Kneeland, J. et al. Identification and fine mapping of quantitative trait loci for growth traits on bovine chromosomes 2, 6, 14, 19, 21, and 23 within one commercial line of Bos taurus. J. Anim. Sci. 82, 3405–3414 (2004).
Nkrumah, J.D. et al. Primary genome scan to identify putative quantitative trait loci for feedlot growth rate, feed intake, and feed efficiency of beef cattle. J. Anim. Sci. 85, 3170–3181 (2007).
Mizoshita, K. et al. Identification of a 1.1-Mb region for a carcass weight QTL on bovine Chromosome 14. Mamm. Genome 16, 532–537 (2005).
Druet, T. & Georges, M. An efficient haplotype-based approach for combined linkage and linkage disequilibrium QTL mapping using Hidden Markov Models. Genetics 184, 789–798 (2010).
Matukumalli, L.K. et al. Development and characterization of a high density SNP genotyping assay for cattle. PLoS ONE 4, e5350 (2009).
Lynch, M. & Walsh, B. Genetics and Analysis of Quantitative Traits. (Sinauer, Sunderland, Massachusetts, USA, 1997).
Gudbjartsson, D.F. et al. Many sequence variants affecting diversity of adult human height. Nat. Genet. 40, 609–615 (2008).
Lettre, G. et al. Identification of ten loci associated with height highlights new biological pathways in human growth. Nat. Genet. 40, 584–591 (2008).
Soranzo, N. et al. Meta-analysis of genome-wide scans for human adult stature identifies novel loci and associations with measures of skeletal frame size. PLoS Genet. 5, e1000445 (2009).
Cho, Y.S. et al. A large-scale genome-wide association study of Asian populations uncovers genetic factors influencing eight quantitative traits. Nat. Genet. 41, 527–534 (2009).
Kim, J.J. et al. Identification of 15 loci influencing height in a Korean population. J. Hum. Genet. 55, 27–31 (2010).
Okada, Y. et al. A genome-wide association study in 19,633 Japanese subjects identified LHX3-QSOX2 and IGF1 as adult height loci. Hum. Mol. Genet. 19, 2303–2312 (2010).
Ge, B. et al. Survey of allelic expression using EST mining. Genome Res. 15, 1584–1591 (2005).
Trinklein, N.D. et al. An abundance of bidirectional promoters in the human genome. Genome Res. 14, 62–66 (2004).
McGowan, K.A. et al. Ribosomal mutations cause p53-mediated dark skin and pleiotropic effects. Nat. Genet. 40, 963–970 (2008).
Sagata, N. What does Mos do in occytes and somatic cells? Bioessays 19, 13–21 (1997).
Van Dyck, F. et al. PLAG1, the prototype of the PLAG gene family: versatility in tumour development. Int. J. Oncol. 30, 765–774 (2007).
Voz, M.L. et al. Microarray screening for target genes of the proto-oncogene PLAG1. Oncogene 23, 179–191 (2004).
Hensen, K. et al. Targeted disruption of the murine Plag1 proto-oncogene causes growth retardation and reduced fertility. Dev. Growth Differ. 46, 459–470 (2004).
Ross, S.A., McCaffrey, P.J., Drager, U.C. & De Luca, L.M. Retinoids in embryonal development. Physiol. Rev. 80, 1021–1054 (2000).
Kieffer, B.L. et al. Exploring the opioid system by gene knock-out. Prog. Neurobiol. 66, 285–306 (2002).
Georges, M. Mapping, fine-mapping and molecular dissection of Quantitative Trait Loci in domestic animals. Annu. Rev. Genomics Hum. Genet. 8, 131–162 (2007).
Mackay, T.F. Quantitative trait loci in Drosophila. Nat. Rev. Genet. 2, 11–20 (2001).
Service, P.M. How good are quantitative complementation tests? Sci. SAGE KE 12, pe13 (2004).
Steinmetz, L.M. et al. Dissecting the architecture of a quantitative trait locus in yeast. Nature 416, 326–330 (2002).
McGregor, A.P. et al. Morphological evolution through multiple cis-regulatory mutations at a single gene. Nature 448, 587–590 (2007).
Van Laere, A.S. et al. A regulatory mutation in IGF2 causes a major QTL effect on muscle growth in the pig. Nature 425, 832–836 (2003).
Nezer, C. et al. Haplotype sharing refines the location of an imprinted QTL with major effect on muscle mass to a 250-kb chromosome segment containing the porcine IGF2 gene. Genetics 165, 277–285 (2003).
Bodmer, W. & Bonilla, C. Common and rare variants in multifactorial susceptibility to common diseases. Nat. Genet. 40, 695–701 (2008).
Long, A.D., Lyman, R.F., Langley, C.H. & Mackay, T.F. Genetic interactions between naturally occurring alleles at QTL and mutant alleles at candidate loci affecting bristle number in D. melanogaster. Genetics 144, 1497–1510 (1996).
Yalcin, B. et al. Genetic dissection of behavioral QTL shows that Rgs2 modulates anxiety in mice. Nat. Genet. 36, 1197–1202 (2004).
1000 Genomes Project Consortium. et al. A map of human genome variation from population-scale sequencing. Nature 467, 1061–1073 (2010).
Coppieters, W. et al. A QTL with major effect on milk yield and composition maps to bovine chromosome 14. Mamm. Genome 9, 540–544 (1998).
Churchill, G.A. & Doerge, R.W. Empirical threshold values for quantitative trait mapping. Genetics 138, 963–971 (1994).
Misztal, I. et al. BLUPF90 and related programs (BGF90). In: 7th World Congress on Genetics Applied to Livestock Production (Montpelier, 19–23 August 2002).
Vandesompele, J. et al. Accurate normalization of real-time quantitative RT-PCR data by geometric averaging multiple internal control genes. Genome Biol. 3, research0034 (2002).
Acknowledgements
This work was funded by Livestock Improvement Corporation (LIC; Hamilton, New Zealand) and by grants from the Unit of Animal Genomics, the University of Liège, the Communauté Française de Belgique (ARC Biomod) and the Belgian Science Policy Organisation (SSTC Genefunc PAI). T.D. is Research Associate of the Fond National de Recherche Scientifique. We are grateful for the support of the GIGA-R sequencing core facility.
Author information
Authors and Affiliations
Contributions
J.A.C.A., B.L.H., M.D.K. and R.J.S. designed and performed line-cross QTL mapping in the F2 population. L.K., L.L., N.C., B.G. and W.C. developed additional BTA14 markers, genotyped the F2 population and performed half-sibling QTL mapping. T.D., F.F. and W.C. performed combined linkage and LD QTL fine mapping. L.K. and W.C. performed high throughput resequencing and analysis of the 780-kb confidence interval. L.L. performed sequence finishing of the 780-kb interval. L.K., N.C. and W.C. performed haplotype analysis in the breed diversity panel. S.R.D. collected fetal samples. L.K., H.T. and L.L. checked the integrity of the open reading frames. H.T., L.L., M.D.L. and M.G. performed quantitative RT-PCR experiments. H.T. performed the allelic imbalance tests. H.T. performed the reporter assays. L.K. and H.T. performed the EMSA. S.R.D., M.D.K. and R.J.S. generated and performed initial analysis of the transcriptome data. T.D., D.B. and W.C. performed eQTL analyses. L.L. analyzed the effect of the CHCHD7 splice site variant. W.C. and T.D. performed the QCA. M.G. designed experiments, analyzed data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–12, Supplementary Tables 1–3 and Supplementary Note. (PDF 6262 kb)
Supplementary Table 4
Expression data for eQTL analysis (XLS 190 kb)
Supplementary Table 5
Pedigree file for eQTL analysis (XLS 247 kb)
Rights and permissions
About this article
Cite this article
Karim, L., Takeda, H., Lin, L. et al. Variants modulating the expression of a chromosome domain encompassing PLAG1 influence bovine stature. Nat Genet 43, 405–413 (2011). https://doi.org/10.1038/ng.814
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.814
This article is cited by
-
Sequence level genome-wide associations for bull production and fertility traits in tropically adapted bulls
BMC Genomics (2023)
-
Sequence-based GWAS meta-analyses for beef production traits
Genetics Selection Evolution (2023)
-
Sequenced-based GWAS for linear classification traits in Belgian Blue beef cattle reveals new coding variants in genes regulating body size in mammals
Genetics Selection Evolution (2023)
-
The molecular evolution of genes previously associated with large sizes reveals possible pathways to cetacean gigantism
Scientific Reports (2023)
-
Genome-wide association study for growth traits in Blanco Orejinegro and Romosinuano cattle
Tropical Animal Health and Production (2023)