Abstract
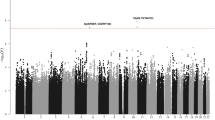
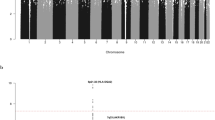
To identify susceptibility loci for classical Hodgkin's lymphoma (cHL), we conducted a genome-wide association study of 589 individuals with cHL (cases) and 5,199 controls with validation in four independent samples totaling 2,057 cases and 3,416 controls. We identified three new susceptibility loci at 2p16.1 (rs1432295, REL, odds ratio (OR) = 1.22, combined P = 1.91 × 10−8), 8q24.21 (rs2019960, PVT1, OR = 1.33, combined P = 1.26 × 10−13) and 10p14 (rs501764, GATA3, OR = 1.25, combined P = 7.05 × 10−8). Furthermore, we confirmed the role of the major histocompatibility complex in disease etiology by revealing a strong human leukocyte antigen (HLA) association (rs6903608, OR = 1.70, combined P = 2.84 × 10−50). These data provide new insight into the pathogenesis of cHL.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Kuppers, R. The biology of Hodgkin's lymphoma. Nat. Rev. Cancer 9, 15–27 (2009).
Swerdlow, A.J. Epidemiology of Hodgkin's disease and non-Hodgkin's lymphoma. Eur. J. Nucl. Med. Mol. Imaging 30 Suppl 1, S3–S12 (2003).
Smith, A. et al. The Haematological Malignancy Research Network (HMRN): a new information strategy for population based epidemiology and health service research. Br. J. Haematol. 148, 739–753 (2010).
Kutok, J.L. & Wang, F. Spectrum of Epstein-Barr virus–associated diseases. Annu. Rev. Pathol. 1, 375–404 (2006).
Goldin, L.R. et al. Familial aggregation of Hodgkin lymphoma and related tumors. Cancer 100, 1902–1908 (2004).
Amiel, J. Study of the leukocyte phenotypes in Hodgkin's disease. in Histocompatibility Testing (ed, Teraski, P.I.) (Munksgaard, Copenhagen, Denmark, 1967).
Klitz, W., Aldrich, C.L., Fildes, N., Horning, S.J. & Begovich, A.B. Localization of predisposition to Hodgkin disease in the HLA class II region. Am. J. Hum. Genet. 54, 497–505 (1994).
Oza, A.M. et al. A clinical and epidemiological study of human leukocyte antigen-DPB alleles in Hodgkin's disease. Cancer Res. 54, 5101–5105 (1994).
Hjalgrim, H. et al. HLA-A alleles and infectious mononucleosis suggest a critical role for cytotoxic T-cell response in EBV-related Hodgkin lymphoma. Proc. Natl. Acad. Sci. USA 107, 6400–6405 (2010).
Niens, M. et al. HLA-A*02 is associated with a reduced risk and HLA-A*01 with an increased risk of developing EBV+ Hodgkin lymphoma. Blood 110, 3310–3315 (2007).
Risch, N. Assessing the role of HLA-linked and unlinked determinants of disease. Am. J. Hum. Genet. 40, 1–14 (1987).
Goldin, L.R. et al. A genome screen of families at high risk for Hodgkin lymphoma: evidence for a susceptibility gene on chromosome 4. J. Med. Genet. 42, 595–601 (2005).
Mack, T.M. et al. Concordance for Hodgkin's disease in identical twins suggesting genetic susceptibility to the young-adult form of the disease. N. Engl. J. Med. 332, 413–418 (1995).
Power, C. & Elliott, J. Cohort profile: 1958 British birth cohort (National Child Development Study). Int. J. Epidemiol. 35, 34–41 (2006).
Clayton, D.G. et al. Population structure, differential bias and genomic control in a large-scale, case-control association study. Nat. Genet. 37, 1243–1246 (2005).
Price, A.L. et al. Principal components analysis corrects for stratification in genome-wide association studies. Nat. Genet. 38, 904–909 (2006).
Wellcome Trust Case-Control Consortium. Genome-wide association study of 14,000 cases of seven common diseases and 3,000 shared controls. Nature 447, 661–678 (2007).
Cabannes, E., Khan, G., Aillet, F., Jarrett, R.F. & Hay, R.T. Mutations in the IkBa gene in Hodgkin's disease suggest a tumour suppressor role for IkappaBalpha. Oncogene 18, 3063–3070 (1999).
Emmerich, F. et al. Overexpression of IκBα without inhibition of NF-κB activity and mutations in the IκBα gene in Reed-Sternberg cells. Blood 94, 3129–3134 (1999).
Schmitz, R. et al. TNFAIP3 (A20) is a tumor suppressor gene in Hodgkin lymphoma and primary mediastinal B cell lymphoma. J. Exp. Med. 206, 981–989 (2009).
Barth, T.F. et al. Gains of 2p involving the REL locus correlate with nuclear c-Rel protein accumulation in neoplastic cells of classical Hodgkin lymphoma. Blood 101, 3681–3686 (2003).
Joos, S. et al. Hodgkin's lymphoma cell lines are characterized by frequent aberrations on chromosomes 2p and 9p including REL and JAK2. Int. J. Cancer 103, 489–495 (2003).
Martin-Subero, J.I. et al. Recurrent involvement of the REL and BCL11A loci in classical Hodgkin lymphoma. Blood 99, 1474–1477 (2002).
Villeneuve, L., Rassart, E., Jolicoeur, P., Graham, M. & Adams, J.M. Proviral integration site Mis-1 in rat thymomas corresponds to the pvt-1 translocation breakpoint in murine plasmacytomas. Mol. Cell. Biol. 6, 1834–1837 (1986).
Beck-Engeser, G.B. et al. Pvt1-encoded microRNAs in oncogenesis. Retrovirology 5, 4 (2008).
Bakkus, M.H., Brakel-van Peer, K.M., Michiels, J.J., van 't Veer, M.B. & Benner, R. Amplification of the c-myc and the pvt-like region in human multiple myeloma. Oncogene 5, 1359–1364 (1990).
Huppi, K., Siwarski, D., Skurla, R., Klinman, D. & Mushinski, J.F. Pvt-1 transcripts are found in normal tissues and are altered by reciprocal(6;15) translocations in mouse plasmacytomas. Proc. Natl. Acad. Sci. USA 87, 6964–6968 (1990).
Storlazzi, C.T. et al. Identification of a commonly amplified 4.3 Mb region with overexpression of C8FW, but not MYC in MYC-containing double minutes in myeloid malignancies. Hum. Mol. Genet. 13, 1479–1485 (2004).
Crowther-Swanepoel, D. et al. Common variants at 2q37.3, 8q24.21, 15q21.3 and 16q24.1 influence chronic lymphocytic leukemia risk. Nat. Genet. 42, 132–136 (2010).
Gudmundsson, J. et al. Genome-wide association study identifies a second prostate cancer susceptibility variant at 8q24. Nat. Genet. 39, 631–637 (2007).
Easton, D.F. et al. Genome-wide association study identifies novel breast cancer susceptibility loci. Nature 447, 1087–1093 (2007).
Tomlinson, I. et al. A genome-wide association scan of tag SNPs identifies a susceptibility variant for colorectal cancer at 8q24.21. Nat. Genet. 39, 984–988 (2007).
Yeager, M. et al. Genome-wide association study of prostate cancer identifies a second risk locus at 8q24. Nat. Genet. 39, 645–649 (2007).
Amundadottir, L.T. et al. A common variant associated with prostate cancer in European and African populations. Nat. Genet. 38, 652–658 (2006).
Kiemeney, L.A. et al. Sequence variant on 8q24 confers susceptibility to urinary bladder cancer. Nat. Genet. 40, 1307–1312 (2008).
Tuupanen, S. et al. The common colorectal cancer predisposition SNP rs6983267 at chromosome 8q24 confers potential to enhanced Wnt signaling. Nat. Genet. 41, 885–890 (2009).
Pomerantz, M.M. et al. The 8q24 cancer risk variant rs6983267 shows long-range interaction with MYC in colorectal cancer. Nat. Genet. 41, 882–884 (2009).
Faumont, N. et al. c-Myc and Rel/NF-κB are the two master transcriptional systems activated in the latency III program of Epstein-Barr virus-immortalized B cells. J. Virol. 83, 5014–5027 (2009).
Ho, I.C., Tai, T.S. & Pai, S.Y. GATA3 and the T-cell lineage: essential functions before and after T-helper-2-cell differentiation. Nat. Rev. Immunol. 9, 125–135 (2009).
Li, J., Qian, C.N. & Zeng, Y.X. Regulatory T cells and EBV associated malignancies. Int. Immunopharmacol. 9, 590–592 (2009).
Atayar, C. et al. Expression of the T-cell transcription factors, GATA-3 and T-bet, in the neoplastic cells of Hodgkin lymphomas. Am. J. Pathol. 166, 127–134 (2005).
La Rosa, F.A., Pierce, J.W. & Sonenshein, G.E. Differential regulation of the c-myc oncogene promoter by the NF-κB rel family of transcription factors. Mol. Cell. Biol. 14, 1039–1044 (1994).
Maurice, D., Hooper, J., Lang, G. & Weston, K. c-Myb regulates lineage choice in developing thymocytes via its target gene Gata3. EMBO J. 26, 3629–3640 (2007).
Papaemmanuil, E. et al. Loci on 7p12.2, 10q21.2 and 14q11.2 are associated with risk of childhood acute lymphoblastic leukemia. Nat. Genet. 41, 1006–1010 (2009).
Jundt, F. et al. Loss of PU.1 expression is associated with defective immunoglobulin transcription in Hodgkin and Reed-Sternberg cells of classical Hodgkin disease. Blood 99, 3060–3062 (2002).
Hjalgrim, H. & Engels, E.A. Infectious aetiology of Hodgkin and non-Hodgkin lymphomas: a review of the epidemiological evidence. J. Intern. Med. 264, 537–548 (2008).
Stranger, B.E. et al. Genome-wide associations of gene expression variation in humans. PLoS Genet. 1, e78 (2005).
Stranger, B.E. et al. Relative impact of nucleotide and copy number variation on gene expression phenotypes. Science 315, 848–853 (2007).
Alexander, F.E. et al. An epidemiologic study of index and family infectious mononucleosis and adult Hodgkin's disease (HD): evidence for a specific association with EBV+ve HD in young adults. Int. J. Cancer 107, 298–302 (2003).
Jarrett, R.F. et al. The Scotland and Newcastle epidemiological study of Hodgkin's disease: impact of histopathological review and EBV status on incidence estimates. J. Clin. Pathol. 56, 811–816 (2003).
Willett, E.V., O'Connor, S., Smith, A.G. & Roman, E. Does smoking or alcohol modify the risk of Epstein-Barr virus-positive or -negative Hodgkin lymphoma? Epidemiology 18, 130–136 (2007).
Penegar, S. et al. National study of colorectal cancer genetics. Br. J. Cancer 97, 1305–1309 (2007).
Hjalgrim, H. et al. Infectious mononucleosis, childhood social environment, and risk of Hodgkin lymphoma. Cancer Res. 67, 2382–2388 (2007).
Smedby, K.E. et al. Ultraviolet radiation exposure and risk of malignant lymphomas. J. Natl. Cancer Inst. 97, 199–209 (2005).
Sorensen, K.M. et al. Whole genome amplification on DNA from filter paper blood spot samples: an evaluation of selected systems. Genet. Test. 11, 65–71 (2007).
van Leeuwen, F.E. et al. Roles of radiation dose, chemotherapy, and hormonal factors in breast cancer following Hodgkin's disease. J. Natl. Cancer Inst. 95, 971–980 (2003).
Broeks, A. et al. Increased risk of breast cancer following irradiation for Hodgkin's disease is not a result of ATM germline mutations. Int. J. Radiat. Biol. 76, 693–698 (2000).
De Bruin, M.L. et al. Breast cancer risk in female survivors of Hodgkin's lymphoma: lower risk after smaller radiation volumes. J. Clin. Oncol. 27, 4239–4246 (2009).
Lake, A. et al. Mutations of NFKBIA, encoding IκBα, are a recurrent finding in classical Hodgkin lymphoma but are not a unifying feature of non-EBV-associated cases. Int. J. Cancer 125, 1334–1342 (2009).
Pettiti, D. Meta-analysis Decision Analysis and Cost-Effectiveness Analysis (Oxford University Press, Oxford, New York, 1994).
Higgins, J.P. & Thompson, S.G. Quantifying heterogeneity in a meta-analysis. Stat. Med. 21, 1539–1558 (2002).
Myers, S., Bottolo, L., Freeman, C., McVean, G. & Donnelly, P. A fine-scale map of recombination rates and hotspots across the human genome. Science 310, 321–324 (2005).
Gabriel, S.B. et al. The structure of haplotype blocks in the human genome. Science 296, 2225–2229 (2002).
Matys, V. et al. TRANSFAC and its module TRANSCompel: transcriptional gene regulation in eukaryotes. Nucleic Acids Res. 34, D108–D110 (2006).
Ferretti, V. et al. PReMod: a database of genome-wide mammalian cis-regulatory module predictions. Nucleic Acids Res. 35, D122–D126 (2007).
Hallikas, O. et al. Genome-wide prediction of mammalian enhancers based on analysis of transcription-factor binding affinity. Cell 124, 47–59 (2006).
Cuzick, J. Wilcoxon-type test for trend. Stat. Med. 4, 87–90 (1985).
Acknowledgements
Leukemia Lymphoma Research (UK) and Cancer Research UK (C1298/A8362 supported by the Bobby Moore Fund) provided principal funding for the study. We acknowledge National Health Service (NHS) funding to the National Institute for Health Research (NIHR) Biomedical Research Centre. This study made use of control genotyping data generated by the Wellcome Trust Case Control Consortium. We acknowledge use of genotype data from the British 1958 Birth Cohort DNA collection, which was funded by the Medical Research Council grant G0000934 and the Wellcome Trust grant 068545/Z/02. A full list of the investigators who contributed to the generation of the data is available from http://www.wtccc.org.uk. Funding for the project was provided by the Wellcome Trust under awards 076113 and 085475. At the Institute of Cancer Research, sample and data acquisition was supported by Breakthrough Breast Cancer and the European Union, and we acknowledge NHS funding to the NIHR Biomedical Research Centre. We are grateful to the patients and their clinicians who participated in this collection (Supplementary Note). Work at the Leukaemia Research Fund (LRF) Virus Centre was funded by Leukaemia and Lymphoma Research. Sample and data acquisition for the UK replication series was also supported by the Kay Kendall Leukaemia Fund. The Epidemiology and Genetics Lymphoma Case-Control Study (ELCCS) was funded by Leukaemia and Lymphoma Research. Grant support to the German Study Group was through Deutsche Krebshilfe and the EU, grant HEALTH-F4-2007-200767. The SCALE study is supported by the Lundbeck Foundation grant R19 A2364, the Danish Cancer Research Foundation grant 41-08 and the Danish Cancer Society grant DP 08155. At the Department of Pathology and Medical Biology, University of Groningen, sample and data acquisition was supported by two grants from the Dutch Cancer Society (RUG 200-2315 and RUG 2009-4313). The Netherlands Cancer Institute (NKI) study was supported by the Dutch Cancer Society (grants No. NKI 98-1833, NKI 04-3068, NKI 08-3994) and the EU 6th framework programme (project no. 012926). We thank A. Kesminiene for coordinating the EU-framework project, M. Schaapveld and A. Eggermond for data management and L. Braaf and I. Mikolajewska for lab assistance. We are indebted to the patients and physicians who participated in this collection (Supplementary Note).
Author information
Authors and Affiliations
Contributions
R.S.H. designed the study and obtained financial support. R.S.H. drafted the manuscript with contributions from P.B., V.E.-M., Y.M. and S.E.D. Y.M. and V.E.-M. performed statistical and bioinformatic analyses. P.B. performed sample coordination and laboratory analyses. B.O., Amy Lloyd and J.V. performed genotyping. A.J.S., A.A. and R.C. provided samples and data from a study conducted at the Institute of Cancer Research. E.R. initiated ELCCS. T.L., M.T. and E.R. managed and prepared Epidemiology and Genetics Lymphoma Case-Control Study samples. R.F.J. designed and conducted studies contributing to the UK replication series, and R.F.J., L.S., Annette Lake and Dorothy Montgomery prepared samples and collated data. F.E.v.L. designed the Dutch NKI study and obtained financial support. N.S.R. and M.d.B. were involved in identification and inclusion of Dutch cases, study design, review board approval and clinical implementation. A.B. coordinated collection and preparation of the NKI samples. A.F., K.H., A.E., E.P.v.S. and K.S.R. provided samples and data from German cases and controls. A.D., I.M.N. and A.v.d.B. collected samples and data from cHL cases ascertained through Groningen University. R.H., H.W., T.v.W. and R.v.E. performed ascertainment and collection of control samples from The Netherlands. H.H., M.M., K.R., L.P.R., K.E.S., H.-O.A., B.G., Daniel Molin, S.H.-D., K.M.S. and E.T.C. provided samples and data from the SCALE study in Denmark and Sweden. S.H.-D. analyzed samples and provided data from Danish cHL cases. All authors contributed to the final paper. R.F.J. and H.H. contributed equally to the paper and should be considered to have equal positional status in the author list.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5, Supplementary Tables 1–8 and Supplementary Note (PDF 10623 kb)
Rights and permissions
About this article
Cite this article
Enciso-Mora, V., Broderick, P., Ma, Y. et al. A genome-wide association study of Hodgkin's lymphoma identifies new susceptibility loci at 2p16.1 (REL), 8q24.21 and 10p14 (GATA3). Nat Genet 42, 1126–1130 (2010). https://doi.org/10.1038/ng.696
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.696
This article is cited by
-
Investigating the tissue specificity and prognostic impact of cis-regulatory cancer risk variants
Human Genetics (2023)
-
Oncofetal reprogramming in tumour development and progression
Nature Reviews Cancer (2022)
-
Distinct germline genetic susceptibility profiles identified for common non-Hodgkin lymphoma subtypes
Leukemia (2022)
-
Linear and circular PVT1 in hematological malignancies and immune response: two faces of the same coin
Molecular Cancer (2020)
-
c-Rel is a myeloid checkpoint for cancer immunotherapy
Nature Cancer (2020)