Abstract
We report a high-quality draft genome sequence of the domesticated apple (Malus × domestica). We show that a relatively recent (>50 million years ago) genome-wide duplication (GWD) has resulted in the transition from nine ancestral chromosomes to 17 chromosomes in the Pyreae. Traces of older GWDs partly support the monophyly of the ancestral paleohexaploidy of eudicots. Phylogenetic reconstruction of Pyreae and the genus Malus, relative to major Rosaceae taxa, identified the progenitor of the cultivated apple as M. sieversii. Expansion of gene families reported to be involved in fruit development may explain formation of the pome, a Pyreae-specific false fruit that develops by proliferation of the basal part of the sepals, the receptacle. In apple, a subclade of MADS-box genes, normally involved in flower and fruit development, is expanded to include 15 members, as are other gene families involved in Rosaceae-specific metabolism, such as transport and assimilation of sorbitol.
Similar content being viewed by others
Main
The domesticated apple (Malus × domestica Borkh., family Rosaceae, tribe Pyreae) is the main fruit crop of temperate regions of the world. Here we describe a high-quality draft genome sequence of the diploid apple cultivar 'Golden Delicious'. Domesticated apple genotypes are all highly heterozygous, imposing technical challenges in genome sequencing and assembly1 while allowing identification of a very large set of SNPs2.
Rosaceae belong to the rosids, which include one-third of all flowering plants3. Whereas the haploid (x) chromosome numbers of most Rosaceae are 7, 8 or 9, Pyreae have a distinctive x = 17. Pyreae have long been considered an example of allopolyploidization between species related to extant Spiraeoideae (x = 9) and Amygdaleoideae (x = 8), although a within-lineage polyploidization event has also been hypothesized4.
In addition, we examine the genetic variability in Rosaceae and related taxa, comparing Pyreae species, Rosaceae tribes and two rosid families. Gene content and order of the assembled chromosomes indicate that both recent and old GWDs have occurred. We provide a model describing the evolution of the Pyreae genome, including Malus, and offer insights into the origin of the domesticated apple.
Results
Sequencing, assembling and anchoring the apple genome
Sequencing and assembly of the 'Golden Delicious' apple genome followed the whole-genome shotgun approach. Of the 16.9-fold genome coverage, 26% was provided by Sanger dye primer sequencing of paired reads, and the remaining 74% was from 454 sequencing by synthesis of paired and unpaired reads (Supplementary Table 1 and Supplementary Note). An iterative assembly approach, previously used to assemble the highly heterozygous grape genome1, produced 122,146 contigs, 103,076 of which were assembled into 1,629 metacontigs (Table 1, Supplementary Fig. 1 and Supplementary Note). The total contig length (603.9 Mb) covers about 81.3% of the apple genome (Table 1 and Supplementary Note). Anchoring of metacontigs (598.3 Mbp, or 71.2% of genome) was based on the high-quality genetic map with 1,643 markers (Supplementary Table 2 and Supplementary Note). In total, 17 linkage groups, or chromosomes, were reconstructed. In the genome, repetitive elements correspond to 500.7 Mb (67%; Supplementary Note). The unassembled part of the genome is 98% repetitive (138.4 Mb), and the estimated genome size is 742.3 Mb (Table 1 and Supplementary Note). We compared repetitive elements among ten plant species (Supplementary Tables 3–6). Information on relevant genes and genome parameters is provided in Tables 1 and 2, Supplementary Figures 2–5 and Supplementary Tables 7–19. Comparing gene families among ten sequenced plant species revealed apple-specific subclades of genes encoding MADS-box transcription factors and overrepresented sorbitol-related genes, which may contribute to specific aspects of apple development and carbohydrate metabolism (Table 2 and Supplementary Table 7, and see Discussion). The 71.2% of the genomic sequences that were anchored represent the gene-rich part of the genome, which covers as many as 90.2% of the genes assigned to the chromosomes. The distribution of transposable elements and predicted genes along the linkage groups is reported in Supplementary Figure 6. The total number of genes predicted for the apple genome (57,386, including some genes that may be present only in one of the two chromosomes of a pair) is the highest reported among plants so far (Supplementary Note).
Genome-wide duplications and the origin of the Pyreae
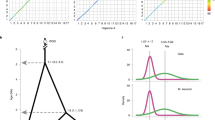
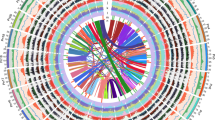
Pairwise comparison of 17 apple chromosomes highlighted strong collinearity between large segments of chromosomes 3 and 11, 5 and 10, 9 and 17, and 13 and 16, and between shorter segments of chromosomes 1 and 7, 2 and 7, 2 and 15, 4 and 12, 12 and 14, 6 and 14, and 8 and 15 (Fig. 1a). The distribution of synonymous substitution rates (KS)—an indication of the relative age of duplication, based on the number of synonymous substitutions in the coding sequences—peaked around 0.2 for recently duplicated genes (Fig. 1b), indicating that a (recent) GWD has shaped the genome of the domesticated apple.
(a) Alignment of apple chromosomes shown by pairwise dot plots based on gene homology. Strong collinearity of members of chromosome doublets, or of large chromosome segments, indicates a recent GWD (red dots and bars in a and b, respectively). Unrelated chromosomes 7 and 13 were compared as a negative control. (b) Reconstruction of the relationships among apple chromosomes based on the most recent and the older GWD. The model derives from data in a for the recent GWD and from data in Supplementary Figure 6b for the oldest GWD. The chromosomes ends represented at bottom right corners in a are marked in black in b. Red bars, regions of synteny that support the recent GWD. Size of chromosomes is proportional to their DNA content in megabases. Segments of chromosomes 1, 5, 6, 8, 10, 13, 14 and 15 have no syntenic counterparts. Chromosome segments predicted to be the outcome of the older duplication are highlighted with blue, green and orange. Chromosomes 1, 2, 7, 8 and 15 do not show obvious signs of the older duplications, although they may contain short blocks of genes that reveal old paleopolyploid events. Inset graphs show that Ks from the comparisons between paralogous genes has a peak at 0.2 when the recent duplication is considered, and between 1.4 and 1.6 for the older paleopolyploid events. (c) Distributions of protein similarities for duplicated genes in duplicated segments compared with grape (red), poplar (green) and apple (blue).
Dating of this GWD (Supplementary Note) was based on the construction of penalized likelihood trees, as described previously5. Given a node of grape to rosids fixed at 115 million years ago (Mya), the GWD has been dated to between 30 and 45 Mya5. If similar rates of protein evolution are assumed for apple and poplar (Fig. 1c), the recent apple GWD may be as old as that of poplar, about 60 to 65 Mya6.
Remnants of older large-scale gene duplications or GWDs were also evident (Supplementary Fig. 7a,b). Genes in these duplicated regions had average KS values around 1.6, as expected for paleoduplication events (Fig. 1b). Most remnants of these older duplications are found between chromosomes 5 and 10 and chromosomes 3 and 11, between chromosomes 3 and 11 and chromosomes 4 and 12, and between chromosomes 6 and 14, 13 and 16, and 9 and 17 (Fig. 1a,b). Chromosomes 1, 2, 7, 8 and 15 seem relatively devoid of older duplicated blocks; however, short blocks of genes showing old polyploidy events were found on all chromosomes. One region in the apple genome with an approximate size of 4 to 7 Mbp seems to be clearly present in six copies (regions in blue, Fig. 1a,b). Remapping those to the ancestral state reveals a triplicate structure among parts of chromosomes 9 and 17, 6 and 14 and 13 and 16. Notably, we found that these regions are collinear with chromosomes 1, 14 and 17 of grape (Fig. 2), which have been demonstrated to be homologous because of an ancient hexaploidy7. Additional chromosomal fragments that we found to be duplicated in apple (green and yellow bars in Fig. 1b) can also be interpreted as remains of a paleohexaploid state of the eudicot progenitor on the basis of dot-plot comparisons among other grape and apple chromosomes (Supplementary Fig. 8a,b). This provides further evidence for a paleohexaploid state shared by most eudicots8,9.
Dot plots are based on gene homology. The apple chromosomes are those with the segment triplication deriving from an old GWD (shown in blue in Fig. 1b). Grape chromosomes 1, 14 and 17 constitute a triplet having the same ancestor in common7. Chromosome segments with homologous genes common both to grape and apple (16 of a total of 18 comparisons) are indicated by gray boxes connected with dashed lines. Green, red and blue dots indicate increasing Ks values, in that order. Perpendicular lines on the x and y axes mark the middle of each chromosome. Green grid separates chromosomes.
The chromosome homologies derived from the recent GWD allow inference of the cytological events that have led to the number and composition of the extant apple chromosomes, starting from a putative nine-chromosome ancestor (Fig. 3). Each doublet of the eight apple chromosomes (3-11, 5-10, 9-17 and 13-16) is derived principally from one ancestor, although minor interchromosomal rearrangements have occurred (Supplementary Fig. 9a–k). Chromosomes 4, 6, 12 and 14 originate from duplications of the ancient chromosomes V and VI, followed by a translocation and a deletion event. Similar events have generated chromosomes 1, 2, 7, 8 and 15 from chromosomes VII, VIII and IX. Chromosome 15 could have been produced from the translocation of an entire copy of chromosome IX into the centromeric region of chromosome VIII, following a model of dysploidy (reduction of chromosome number) common in cereals10. The second copy of ancient chromosome VIII has evolved into the extant chromosome 8. A conservative estimate of the number of large chromosome rearrangements since the divergence of the Pyreae subtribe, corresponding to the recent chromosome duplication, includes one chromosome fusion (extant chromosome 15), three translocations (involving extant chromosomes 1, 2 and 14), six deletions defined by telomeres that are not currently duplicated (chromosomes 4, 6, 8, 10, 11 and 13), one intrachromosome deletion (within chromosome 7, according to the chromosome 1–chromosome 7 comparison) and a deletion of a centromere (from ancient chromosome IX).
A GWD followed by a parsimony model of chromosome rearrangements is postulated. Shared colors indicate homology between extant chromosomes. White fragments of chromosomes indicate lack of a duplicated counterpart. The white-hatched portions of chromosomes 5 and 10 indicate partial homology (see also Supplementary Fig. 9). Black marks at chromosome ends correspond to those in Figure 1b.
Molecular distances, taxonomy and phylogeny of Rosaceae
Available Rosaceae molecular data allow intrafamily comparisons of apple with pear and of a consensus of apple and pear with peach. Further comparisons with grape—a species basal to rosids but belonging to the Vitaceae, a strictly different, although related, family—introduce the possibility of comparing interfamily molecular distances. DNA sequences used in this molecular phylogeny consist of those from EST databases and, for apple, the genomic data as described in detail in the Supplementary Note. Data from a three-way sequence alignment between predicted gene space in apple (∼84 Mb) and experimentally derived EST data from pear (∼14.9 Mb) and peach (∼18 Mb), performed as in ref. 11, indicates that the genetic distance, based on DNA sequence divergence per base pair between members of Rosaceae, increases from apple to pear to peach (Supplementary Table 20). When predicted gene spaces of apple and pear were compared, a value of 96.35% nucleotide identity was calculated between these two species of the tribe Pyreae. The estimate for nucleotide identity between the tribes Pyreae and Amygdaleae (apple and peach) was 90.64%. When grape was compared with apple and pear, nucleotide identity was estimated at 85.31%. When the frequency of transitions and transversions was considered (Fig. 4), the ratio R (transitions/transversions) was similar for apple-specific and pear-specific mutations. For peach-specific mutations, the R value is more difficult to interpret, as it is probably biased by the existence of recent GWD in apple and pear. The comparison of apple and pear with grape showed that although transitions were only 20% more frequent than transversions, T-to-G transversions represented 12% of the total number of mutations observed (Fig. 4d), implying that Vitaceae is strongly divergent taxonomically from core members of the Rosaceae.
(a–c) Mutations identified in a three-way comparison of apple, pear and peach. Numbers of transitions and transversions where apple (Ap) differs from pear (Pr) and peach (Pc) (a; 12,273 total), where pear differs from apple and peach (b; 13,124 total), and where peach differs from apple and pear (c; 381,619 total). (d) Number of transitions and transversions in a two-way alignment of grape DNA (Gr) to a consensus sequence (con.) of apple and pear (26,693 total). Note the high rate of T-to-G transversions. R is the transitions/transversions ratio. Methods and computer calculations were similar to those in ref. 11 (Supplementary Note).
The granule-bound starch synthase (gbss) genes, also known as waxy (Wx) genes (divided in two groups, Wx1 and Wx2), were also used4 as a tool to study molecular taxonomy of Rosaceae (Supplementary Table 21). We identified six Wx genes in the apple genome, located on chromosomes 7, 9 and 16 (Wx1 type) and 8, 6 and 14 (Wx2 type) (Supplementary Fig. 10). After counting Wx genes of apple, including putative gene losses in syntenic chromosomal segments, we were able to identify eight two-by-two syntenic regions containing or expected to contain Wx loci. If Wx1-1 on chromosome 7 is not considered (because neither a syntenic Wx-1-1 region nor a paralogous Wx-1-1 copy was found ), four Wx loci should have been present in the nine-chromosome Pyreae ancestor, a result that is consistent with an ancestral paleopolyploid state. When the genomic Wx gene sequences were integrated in the phylogenetic analysis based on sequences present in the Rosaceae database12, the three Wx-1 and the three Wx-2 genes were mapped to two separate clades, both of which also included Wx genes of Gillenia (Supplementary Fig. 11). However, Prunus and Spiraea sequences clustered in separate clades, supporting the conclusions that the tight relationships between apple and Gillenia Wx1 genes, as well as between apple and Gillenia Wx2 genes, were probably generated by the recent GWD (the Pyreae event)—the founding step of the Pyreae genome—and that Prunus- and Spiraea-related species are less likely to have contributed to the Pyreae genome. Hence, we tested the Rosaceae molecular taxonomy12 by Bayesian analysis of the sequences of seven nuclear and chloroplast genes. A major clade with the maximum statistical support included all Pyreae (x = 17) as well as Gillenia (x = 9) (Supplementary Fig. 12). Notably, the genera Spiraea (x = 9) and Prunus (x = 8) were not included in this clade.
Apple domestication
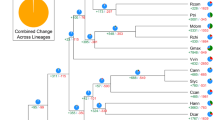
Although M. sieversii has been considered to be the ancestor of the domesticated apple13, this has been challenged by the identification of molecular similarities between domestic apple and M. sylvestris14. To test these two hypotheses, we surveyed molecular differences at 23 genes across the genus Malus (Supplementary Table 22). The 74 accessions we considered included 12 M. × domestica cultivars, 10 M. sieversii, 21 M. sylvestris, all major wild apple species and two Pyrus species (Supplementary Table 23). For M. × domestica, we included the cultivars 'Cox's Orange Pippin', 'Golden Delicious', 'McIntosh', 'Red Delicious' and 'Jonathan', the most important 'founders' of modern apple breeding15 (Supplementary Note). For each gene and accession, a PCR amplicon was resequenced and the data were analyzed as a concatenated data set with a total length of ∼11,300 bp, with 1,507 polymorphic informative sites. A neighbor-net planar graph16 was constructed from the molecular differences among accessions (Fig. 5 and Supplementary Fig. 13). Although the clade containing M. sylvestris was well separated from the clade with M. × domestica, M. sieversii and M. × domestica genotypes shared a large common clade that also included accessions of M. orientalis and M. × asiatica. The average polymorphism rate within the domestic cultivars was 4.8 SNPs per kb, with 5.7 SNPs per kb between 'Golden Delicious' and M. sieversii, and 9.6 SNPs per kb between and M. sylvestris (Supplementary Table 24). The genetic differentiation was categorized as 'moderate' between M. × domestica and M. sieversii (Fst = 0.14), and 'great' between M. × domestica and M. sylvestris and between M. sieversii and M. sylvestris (Fst = 0.17 and Fst = 0.21, respectively)17. The mean numbers of haplotypes per gene were 6.4, 5.8 and 10.0 for M. × domestica, M. sieversii and M. sylvestris, respectively (Supplementary Table 25).
Black, orange and green, accessions of M. × domestica cultivars, M. sieversii and M. sylvestris, respectively; A, O and P, accessions of M. × asiatica, M. orientalis and Pyrus, respectively; squares, accessions of all other wild species. Full information on accessions is provided in Supplementary Table 23 and Supplementary Figure 13. The split separating the M. × domestica–M. sieversii–M. orientalis–M. × asiatica complex from other species is highlighted in red, and the split separating M. sylvestris is highlighted in green. Genetic distances were obtained as Hamming distances, with pairwise alignment of nucleotide positions. The planar graph was constructed with Splits-Tree 4.10.
Discussion
The putative gene content in apple (57,386 putative genes plus 31,678 transposable element–related ORFs) is high compared to Arabidopsis thaliana (27,228), poplar (45,654), papaya (28,027), Brachypodium distachyon (25,532), grape (33,514), rice (40,577), sorghum (34,496), cucumber (26,682), soybean (46,430) and maize (32,540). Putative apple-specific genes, identified as described in Supplementary Note, totaled 11,444. The gene density in apple (Table 2) is within the range of those in poplar and grape, but lower than those in Arabidopsis, Brachypodium and rice. The existence of hemizygous DNA in the heterozygous variety 'Golden Delicious' may have contributed to this gene number, as has also been noted for grape2.
The apple genome has a relatively high number of repeated sequences, which are difficult to assemble or anchor. As seen in grape and cereals, retrotransposons represent the most abundant transposable-element fraction, comprising 38% of the total genome and 89% of all transposable elements (Table 2 and Supplementary Table 7). In contrast, apple has the lowest content of DNA transposons (including the CACTA superfamily) among the reported plant genomes.
The number of transcription factors identified (4,021; Supplementary Table 7) was among the highest of the sequenced plant genomes (Table 2), although the allocation of transcription factor genes to gene families was similar to other sequenced plant species (Supplementary Fig. 3). Partial exceptions were the families C2H2, CCAAT and NAC, which were notably more represented in apple.
The fraction of nucleotide-binding site–leucine-rich repeat (NBS-LRR) resistance genes is considerably higher in eurosids II (apple, poplar and grape) than in eurosids I (Arabidopsis). In monocotyledons (rice), this class of genes predominates. The content of Toll/interleukin region (TIR)-NBS-LRR genes is highest in Arabidopsis (52%), lower in other eurosids (11–32%) and absent in monocots (Table 2). In addition to NBS genes, the apple genome contains 575 LRR-kinase genes.
As seen in other genomes, different classes of apple genes differ greatly in their degree of duplication (Supplementary Table 11 and Supplementary Fig. 4). Across the ten genomes considered, there are gene families with either low or high numbers of paralogous copies. This is particularly evident for genes likely to be involved in metabolism of anthocyanins and flavonoids, isoflavones and isoflavonones, and terpenes (Supplementary Table 7). Relevant cases in each pathway are flavonone 3-hydroxylase (2–13 copies in nine plant genomes) and isoflavone reductase (3–19 copies) compared to isoflavone synthase (54–151 copies); squalene synthase (13 copies) compared to squalene monooxygenase (1–27 copies). It seems that, for some gene classes, the number of paralogous copies may already have been established in the genome of common progenitor(s) of higher plants.
An intriguing aspect of the apple's biology concerns its characteristic fruit, the pome, which is found only in the Pyreae tribe12. This indicates that the pome probably evolved after a relatively recent Pyreae-specific GWD, a polyploidization step that we hypothesize has contributed to the apple's developmental and metabolic specificity (Supplementary Table 7). Pome fruit is derived by enlargement of the receptacle, which is the region below the whorl of sepals in the apple flower. MADS-box genes may regulate pome development, as they determine the eventual fate of floral tissues in all plant species analyzed so far18. For example, it has recently been shown that an apple MADS-box gene that is a member of the AP1 clade, common to all flowering plants19 and closely related to Arabidopsis FRUITFULL (FUL), is differentially expressed during pome development20. In addition, a substantial number of apple type II MADS-box genes belong, phylogenetically, to the StMADS11 subclade, a group named for its first reported member, which was isolated from potato (Supplementary Fig. 14a)21. This subclade includes only two Arabidopsis genes, SVP and AGL24. Ectopic overexpression of SVP and related genes in Arabidopsis leads to foliose sepal syndrome—that is, the formation of large sepals22. In apple, this specific subclade not only includes two genes expressed in the pome but is also expanded to include 15 other genes.
Carbohydrate metabolism is another important aspect of fruit composition. In Rosaceae, photosynthesis-derived carbohydrates are transported mainly as sorbitol23,24. Compared with other plant genomes, apple has considerably more copies of key genes related to sorbitol metabolism. These include aldose 6-P reductase (A6PR), which is rate-limiting for sorbitol biosynthesis, sorbitol-dehydrogenase (SDH), which converts sorbitol to fructose in the fruit25, and sorbitol transporter PcSOT2, which is specific to Rosaceae fruit26,27. In total, there are 71 sorbitol metabolism genes in apple; in other species, the number ranges between 9 and 43 (Supplementary Tables 7 and 26, and Supplementary Fig. 14b–d). In the Rosaceae, an evolutionary trend toward fruit organ specialization may have been partially based on gene duplication, which has created large families of specific paralogous genes (particularly evident for SDH; Supplementary Fig. 14c). Gene families expanded in apple, such as StMADS11-like and SDH-like, have yet to be tested functionally for their involvement in fruit characteristics.
A number of models have been proposed to explain the uniquely high number of chromosomes in Pyreae, the most popular being the 'wide-hybridization' hypothesis based on an allopolyploidization event between spireoid (x = 9) and amygdaloid (x = 8) ancestors28,29. More recent molecular phylogeny studies point to the possibility that Pyreae originated by autopolyploidization or by hybridization between two sister taxa with x = 9 (similar to extant Gillenia), followed by diploidization and aneuploidization4 to x = 17. This hypothesis takes into account that Gillenia and related taxa are New World species and that the earliest fossil evidence of specimens belonging to extant genera of Pyreae are from North America.
Our results support the autopolyploidization hypothesis4, as the derivation from a Gillenia-like taxon best fits the available data. First, the apple genome derives from a relatively recent duplication. Relationships between its homologous chromosomes based on genome sequence extend observations based on synteny and collinearity of molecular markers30,31. The timing of such a GWD, as estimated from our genomic data (Fig. 1c and Supplementary Figs. 15 and 16), agrees with archeobotanical dates of 48–50 Mya32.
Second, molecular phylogeny of Wx genes in the apple genome confirms the close relationship of Gillenia (x = 9) with the Pyreae (x = 17) lineage, as the Wx gene sequences of Prunus, Spiraea and other Rosaceae genera belong to a different phylogenetic cluster (Supplementary Fig. 11). The monophyletic origin of Pyreae and Gillenia was confirmed by a molecular phylogeny of a broader set of genes (Supplementary Fig. 12).
In addition, a simple and parsimonious pattern of chromosome breakage and fusion explains the derivation of the current x = 17 Pyreae karyotype from a polyploidization event of two x = 9 genomes (Fig. 3). The rate of chromosome rearrangements after polyploidization (12 chromosome events in 60 My) is similar to that for poplar (∼16 events in 60 My)6 and lower than in maize (at least 17 chromosome fusion events in 5 My)33 or in artificial neopolyploids34. In this sense, molecular clocks of perennial woody species seem slower than those of annual species, in terms of both nucleotide substitutions and chromosome rearrangements9. For the genus Helianthus, a similar observation that only some of the ancestor chromosomes are rearranged in the extant chromosomes has been discussed in detail. In this genus, such rearrangement was associated with chromosomal differences between two sister species contributing to a GWD allopolyploid event35.
Similarly, the collinearity between Pyrus and Malus genetic maps31,36 suggests that the Pyreae genome reorganization occurred before the divergence of the two genera. A rapid genome rearrangement after polyploidization is expected in species lacking the Ph1-like function that prevents the pairing of homologous chromosomes in wheat37.
It has been proposed that central Asia is the center of origin of domesticated apple38. Between 25 and 47 different Malus species, including M. × domestica, are currently recognized39. As asiatic M. × asiatica, M. baccata, M. micromalus, M. orientalis, M. prunifolia and M. sieversii, and European M. sylvestris, are the species taxonomically closest to M. × domestica39, they are considered to have contributed, to differing extents, to the domestic gene pool. M. sieversii, common in the Tian Shan region of central Asia, is the only wild species sharing all the qualities of the domesticated apple in terms of fruit and tree morphology40.
Apples are known to have been gathered in the Neolithic and Bronze Age in the Near East and Europe, and all archaeological findings indicate a fruit size compatible with those of the wild M. sylvestris41, a species bearing small astringent and acidulate fruits. Sweet apples corresponding to extant domestic apples appeared in the Near East around 4,000 years ago41, at the time when the grafting technology used to propagate the highly heterozygous and self-incompatible apple was becoming available. From the Middle East, the domesticated apple passed to the Greeks and Romans, who spread fruit cultivation across Europe13,41.
On the basis of our molecular results, M. × domestica cultivars appear more closely related to accessions of the wild species M. sieversii and less closely related to accessions of M. sylvestris, M. baccata, M. micromalus and M. prunifolia. The already known42,43 genetic similarity of M. sieversii to M. orientalis and to M. × asiatica (a Chinese cultivated apple form) is also confirmed by our data.
The data support the formation of the M. × domestica gene pool from M. sieversii. Once grafting was introduced, the crop passed through a process described as 'instant domestication'44. This could explain apple's lack of domestication syndrome, which is the loss of sexual reproduction, seed dispersion and seed dormancy. Despite evidence of intrageneric hybridizations14,45, the possibility of substantial genetic contributions to the domestic gene pool of other wild Malus species, such as M. sylvestris14, was rejected in our analysis.
Our study also fully supports the proposal that M. × domestica and M. sieversii are the same species, for which the more appropriate nomenclature of M. pumila Mill. could be adopted13,46.
A practical goal of sequencing the complex heterozygous apple genome is to accelerate the breeding of this economically important perennial crop species. Many genes related to disease resistance, aroma and taste, plant development and reaction to the environment have been identified and mapped to the chromosomes. In addition, SNP molecular markers have been made available at a frequency of 4.4 SNPs per kb. These markers are currently being used in advanced breeding programs and comparative genetic studies31 that should speed cultivar development. The anchored sequence of the apple genome will be a tool to initiate a new era in the breeding of this crop. The availability of nearly all apple gene sequences should benefit apple researchers by enabling genome-wide functional studies and accelerating establishment of gene-trait relationships.
URLs.
Arabidposis thaliana (TAIR Release 8.0), ftp://ftp.arabidopsis.org; Carica papaya, ftp://asgpb.mhpcc.hawaii.edu/papaya/annotation/; Populus trichocarpa (assembly release v1.0, annotation v1.1.), http://genome.jgi-psf.org/Poptr1_1/Poptr1_1.home.html; Vitis vinifera (assembly release v1.0, annotation v2.0), http://genomics.research.iasma.it; Oryza sativa (MSU Rice Genome Annotation Project Release 6.0 assembly), http://rice.plantbiology.msu.edu/index.shtml; Sorghum bicolor (assembly release v1.0, annotation v1.4), http://www.phytozome.net/sorghum; Cucumis sativus (assembly release v1.0, annotation v1.0), http://cucumber.genomics.org.cn/page/cucumber/index.jsp; Glycine max (assembly release Glyma1, annotation Glyma1.0), http://genome.jgi-psf.org/soybean/soybean.home.html; Zea mays (assembly release B73 RefGen_v1), http://www.maizesequence.org/Zea_mays/Info/Index; Brachypodium distachyon (assembly release v1.0, annotation v1.0), http://www.brachybase.org; integrated genetic map, http://genomics.research.iasma.it; RepBase14.01, http://www.girinst.org.
Methods
Plant material.
The DNA of Malus × domestica, variety 'Golden Delicious', was extracted from young leaves of a two-year-old plant grown in the greenhouse at Fondazione Edmund Mac–Istituto Agrario di San Michele all'Adige. The dihaploid 'Golden Delicious' derivative genotype used at Washington State University and the University of Washington to produce 1.5× of 454 sequence was developed by the French National Institute for Agricultural Research47 after a spontaneous duplication of a haploid individual selected in the progeny of a selfed derivative from 'Golden Delicious'47.
'Golden Delicious' was chosen for genome sequencing because of its extensive use in apple breeding programs worldwide. Its heterozygous status did not hamper the genome assembly, thanks to expertise gained in heterozygous grape sequencing1. Indeed, it allowed the inference of both haplotypes, thus giving access to both allelic versions for further genomic projects, and the development of SNP markers. The dihaploid genotype was important for a more accurate haplotype phase determination.
Bacterial artificial chromosomes, shotgun libraries and Sanger sequencing.
The apple bacterial artificial chromosome (BAC) library was from high–molecular weight genomic DNA (Amplicon Express), prepared as described48. The fosmid and shotgun libraries were from genomic DNA provided by R. Meilan (Oregon State University). The shotgun libraries were from DNA sheared with a Gene Machines Hydroshear device. The DNA was size-selected for inserts from 2 to 12 kb to produce libraries of 2, 3, 6, 9 and 11 kb (average sizes). DNA was amplified with the Templiphi kit (GE Healthcare) and sequenced with the Sanger method.
Libraries and 454 pyrosequencing.
Two random shotgun genomic libraries were created by fragmentation of 10 μg of genomic DNA with the GS FLX Titanium library preparation kit (454 Life Sciences). Sequencing was performed with the GS FLX instrument (454 Life Sciences). Further details on library construction and pyrosequencing are in the Supplementary Note.
Genome assembly and anchoring.
From 27 libraries, 39.2 million reads (11.6 billion Q20 bases) were produced by Sanger sequencing and sequencing by synthesis (Supplementary Table 1). Chloroplast and mitochondrial sequences were identified with 847× and 168× coverage, respectively. Chloroplast (160,068 bp) and mitochondrial (396,947 bp) genomes were used to assess sequence quality and clone size in each library. Preliminary estimates of one to two SNPs per 1,000 bp were adopted in the assembly process. The actual SNP rate (4.4 SNPs per 1,000 bp) indicates that the preliminary value was conservative. Metacontigs were constructed on the basis of paired reads matching to nonrepetitive parts of contigs. Merging of contigs into metacontigs accepted a maximum total average coverage of 20×. Fifteen BAC clones were sequenced and individually assembled for quality assessment of sequencing accuracy and genome assembly.
Genetic maps used in metacontig anchoring were derived from six F1 populations totaling 720 individuals (Supplementary Note). Simple sequence repeat primer sequences49,50,51,52 enabled detection of 196 polymorphic markers. Thirty-four SNP-based markers were from apple EST sequences, and 1,489 from genomic electronic SNPs, deduced by genomic sequence comparison between the two haplotypes present in the heterozygous genotype of 'Golden Delicious'. The consensus genetic maps for the six populations were used to generate an integrated genetic map (Supplementary Fig. 1) with TMAP53 and a minimum logarithmic odds of 10.
Repetitive elements.
The highest-coverage sequences were characterized as repetitive elements. Identified elements were iteratively masked, and the remaining sequences were searched for the next highest–coverage sequence. For each type, members were searched (BLASTN and BLASTX) against RepBase14.01, the NCBI databases and the Uniprot database54,55.
Gene prediction and annotation.
FgenesH56, Twinscan57, GlimmerHMM58 and GeneWise59 were used. The predicted protein sequences were searched with BLAST against Uniprot, protein domain data banks and plant protein databases annotated with GO terms. The GO terms were extracted by Argot60 and InterproScan61. Unique genes were searched against proteins from rice, poplar, papaya, barrel medic, sorghum, Arabidopsis and grape by BLAST with an e-value cutoff of e−10.
All-versus-all BLAST.
Protein sequences from apple, poplar6 and grape2 were extracted from a BLAST database, and pairwise similarities between all genes was obtained by BLASTP e-value (cutoff e−5; 500 hits)62.
Gene families.
Tribe-MCL63 was adopted, with parameter I set to 2 and parameter 'scheme' to 4; other parameters were at default values.
Detection of collinearity.
Metacontig anchoring generated lists of apple genes, from which transposable element–related sequences were removed. Poplar and grape gene lists were as described2. Colinearity in the gene order was detected with i-ADHoRe 2.4 (ref. 64), with the following parameters: family blast type; alignment method, gg; gap size, 30; cluster gap, 35; q value, 0.9; prob cutoff, 0.0001; anchor points, 4; level 2 only, false.
Ks dating.
Homologous genes were aligned with CLUSTALW65. Ks dating was based on codeml66 with the following parameters: verbose, 0; noisy, 0; runmode, −2; seqtype, 1; model, 0; NSsites, 0; icode, 0; fix_alpha, 0; fix_kappa, 0; RateAncestor, 0.
Molecular distances, taxonomy and phylogeny.
Molecular distances were analyzed with EST data sets. A two-way alignment between apple and pear contigs (cDNA sequences, data not shown ) was first generated. Sequences from apple and pear were combined with the peach sequence (EST databases) in three-way alignments. Phylogenetic analysis of the Wx genes included gbss1 (Wx1) and gbss2 (Wx2) sequences from the apple genome and from ref. 12. Sequences were aligned by T-coffee67, and phylogenesis was by a Bayesian inference approach (MrBayes program). Phylogeny of Rosaceae was based on four chloroplast DNA sequences and on the nuclear internal transcribed spacer region. The data set included 6,308 positions in 85 operational taxonomic units, each representing one genus, aligned by a Bayesian method68.
Apple domestication.
A set of 74 Malus accessions, including 12 accessions of M. × domestica cultivars15, 10 of M. sieversii and 21 of M. sylvestris, was assembled. This included 31 of 34 recognized Malus species69. Twenty-three genes were resequenced and, after alignment67, a concatenated 11,300-bp multilocus sequence was generated for each accession. Genetic relationships analysis used Splits-Tree v4.10 (ref. 16) and Hamming distance per pair of accessions. Haplotypes were computed with Phase v2.1 (ref. 70). Nucleotide diversity (π), He, Ho and Fst values17 were computed with Arlequin 3.1 (ref. 71).
References
Zharkikh, A. et al. Sequencing and assembly of highly heterozygous genome of Vitis vinifera L. cv Pinot Noir: problems and solutions. J. Biotechnol. 136, 38–43 (2008).
Velasco, R. et al. A high quality draft consensus sequence of the genome of a heterozygous grapevine variety. PLoS ONE 2, e1326 (2007).
Hummer, K.E. & Janick, J. Rosaceae: taxonomy, economic importance, genomics. in Genetics and Genomics of Rosaceae (eds. Folta, K.M. & Gardiner, S.E.) 1–17 (Springer, New York, 2009).
Evans, R.C. & Campbell, C.S. The origin of the apple subfamily (Maloideae; Rosaceae) is clarified by DNA sequence data from duplicated GBSSI genes. Am. J. Bot. 89, 1478–1484 (2002).
Fawcett, J.A., Maere, S. & Van de Peer, Y. Plants with double genomes might have had a better chance to survive the Cretaceous-Tertiary extinction event. Proc. Natl. Acad. Sci. USA 106, 5737–5742 (2009).
Tuskan, G.A. et al. The genome of black cottonwood, Populus trichocarpa (Torr. & Gray). Science 313, 1596–1604 (2006).
Jaillon, O. et al. The grapevine genome sequence suggests ancestral hexaploidization in major angiosperm phyla. Nature 449, 463–467 (2007).
Tang, H. et al. Synteny and collinearity in plant genomes. Science 320, 486 (2008).
Van de Peer, Y., Fawcett, J.A., Proost, S., Sterck, L. & Vandepoele, K. The flowering world: a tale of duplications. Trends Plant Sci. 14, 680–688 (2009).
Luo, M.C. et al. Genome comparisons reveal a dominant mechanism of chromosome number reduction in grasses and accelerated genome evolution in Triticeae. Proc. Natl. Acad. Sci. USA 106, 15780–15785 (2009).
Green, R.E. et al. Analysis of one million base pairs of Neanderthal DNA. Nature 444, 330–336 (2006).
Potter, D. et al. Phylogeny and classification of Rosaceae. Plant Syst. Evol. 266, 5–43 (2007).
Juniper, B.E. & Mabberley, D.J. The Story of the Apple (Timber Press, Portland, Oregon, USA, 2006).
Coart, E., Van Glabeke, S., De Loose, M., Larsen, A.S. & Roldan-Ruiz, I. Chloroplast diversity in the genus Malus: new insights into the relationship between the European wild apple (Malus sylvestris (L.) Mill.) and the domesticated apple (Malus domestica Borkh.). Mol. Ecol. 15, 2171–2182 (2006).
Noiton, D.A.M. & Alspach, P.A. Founding clones, inbreeding, coancestry, and status number of modern apple cultivars. J. Am. Soc. Hortic. Sci. 121, 773–782 (1996).
Huson, D.H. & Bryant, D. Application of phylogenetic networks in evolutionary studies. Mol. Biol. Evol. 23, 254–267 (2006).
Wright, S. Evolution and the Genetics of Populations Vol. 4 (University of Chicago Press, 1978).
Ng, M. & Yanofsky, M.F. Activation of the Arabidopsis B class homeotic genes by APETALA1. Plant Cell 13, 739–753 (2001).
Shan, H. et al. Evolution of plant MADS Box transcription factors: evidence for shifts in selection associated with early angiosperm diversification and concerted gene duplications. Mol. Biol. Evol. 26, 2229–2244 (2009).
Janssen, B.J. et al. Global gene expression analysis of apple fruit development from the floral bud to ripe fruit. BMC Plant Biol. 8, 16 (2008).
Becker, A. & Theissen, G. The major clades of MADS-box genes and their role in the development and evolution of flowering plants. Mol. Phylogenet. Evol. 29, 464–489 (2003).
Masiero, S. et al. INCOMPOSITA: a MADS-box gene controlling prophyll development and floral meristem identity in Antirrhinum. Development 131, 5981–5990 (2004).
Newcomb, R.D. et al. Analyses of expressed sequence tags from apple. Plant Physiol. 141, 147–166 (2006).
Bieleski, R.L. Sugar alcohols. in Encyclopedia of Plant Physiology New Series Vol. 13 (eds. Loewus, F.A. & Tanner, W.) 158–192 (Springer-Verlag, Berlin, 1982).
Loescher, W.H., Marlow, G.C. & Kennedy, R.A. Sorbitol metabolism and sink-source interconversions in developing apple leaves. Plant Physiol. 70, 335–339 (1982).
Watari, J. et al. Identification of sorbitol transporters expressed in the phloem of apple source leaves. Plant Cell Physiol. 45, 1032–1041 (2004).
Gao, Z. et al. Cloning, expression, and characterization of sorbitol transporters from developing sour cherry fruit and leaf sink tissues. Plant Physiol. 131, 1566–1575 (2003).
Chevreau, E., Lespinasse, Y. & Gallet, M. Inheritance of pollen enzymes and polyploid origin of apple. (Malus x domestica Borkh.). Theor. Appl. Genet. 71, 268–277 (1985).
Phipps, J.B., Robertson, K.R., Rohrer, J.R. & Smith, P.G. Origins and evolution of subfam. Maloideae (Rosaceae). Syst. Bot. 16, 303–332 (1991).
Maliepaard, C. et al. Aligning male and female linkage maps of apple (Malus pumila Mill.) using multi-allelic markers. Theor. Appl. Genet. 97, 60–73 (1998).
Celton, J.M., Tustin, D.S., Chagne, D. & Gardiner, S.E. Construction of a dense genetic linkage map for apple rootstocks using SSRs developed from Malus ESTs and Pyrus genomic sequences. Tree Genet. Genomes 5, 93–107 (2009).
Wolfe, J. & Wehr, W.J.A. Rosaceous Chamaebatiaria-like foliage from the Paleogene of western North America. Aliso 12, 177–200 (1988).
Salse, J. et al. Identification and characterization of shared duplications between rice and wheat provide new insight into grass genome evolution. Plant Cell 20, 11–24 (2008).
Doyle, J.J. et al. Evolutionary genetics of genome merger and doubling in plants. Annu. Rev. Genet. 42, 443–461 (2008).
Rieseberg, L.H., Sinervo, B., Linder, C.R., Ungerer, M.C. & Arias, D.M. Role of gene interactions in hybrid speciation: evidence from ancient and experimental hybrids. Science 272, 741–745 (1996).
Yamamoto, T. et al. Genetic linkage maps of Japanese and European pears aligned to the apple consensus map. Acta Hortic. 663, 51–56 (2004).
Griffiths, S. et al. Molecular characterization of Ph1 as a major chromosome pairing locus in polyploid wheat. Nature 439, 749–752 (2006).
Vavilov, N.I. Wild progenitors of the fruit trees of Turkestan and the Caucasus and the problem of the origin of fruit trees. in Proceedings of the 9th International Horticultural Congress 271–286 (The Royal Horticultural Society, London, 1930).
Robinson, J.P., Harris, S.A. & Juniper, B.E. Taxonomy of the genus Malus Mill. (Rosaceae) with emphasis on the cultivated apple, Malus domestica Borkh. Plant Syst. Evol. 226, 35–58 (2001).
Forsline, P.L., Aldwinckle, H.S., Dickson, E.E., Luby, J.J. & Hokanson, S.C. Collection, maintenance, characterization, and utilization of wild apples of Central Asia. Hortic. Rev. (Am. Soc. Hortic. Sci.) 29, 1–61 (2003).
Zohary, D. & Hopf, M. Domestication of Plants in the Old World: The Origin and Spread of Cultivated Plants in West Asia, Europe and the Nile Valley (Clarendon Press, Oxford, 1994).
Gharghani, A. et al. Genetic identity and relationships of Iranian apple (Malus x domestica Borkh.) cultivars and landraces, wild Malus species and representative old apple cultivars based on simple sequence repeat (SSR) marker analysis. Genet. Resour. Crop Evol. 56, 829–842 (2009).
Luby, J. Taxonomy, classification and brief history. in Apples: Botany, Production and Uses (eds. Ferree, D.C. & Warrington, I.J.) 1–14 (CABI, Cambridge, Massachusetts, USA, 2003).
Zohary, D. & Spiegelroy, P. Beginnings of fruit growing in the Old World. Science 187, 319–327 (1975).
Korban, S.S. Interspecific hybridization in Malus. HortScience 21, 41–48 (1986).
Korban, S.S. & Skirvin, R.M. Nomenclature of the cultivate apple. HortScience 19, 177–180 (1984).
Lespinasse, Y., Bouvier, L., Djulbic, M. & Chevreau, E. Haploidy in apple and pear. Acta Hortic. 538, 49–54 (1999).
Tao, Q., Wang, A. & Zhang, H.B. One large-insert plant-transformation-competent BIBAC library and three BAC libraries of Japonica rice for genome research in rice and other grasses. Theor. Appl. Genet. 105, 1058–1066 (2002).
Guilford, P. et al. Microsatellites in Malus X domestica (apple): abundance, polymorphism and cultivar identification. Theor. Appl. Genet. 94, 249–254 (1997).
Liebhard, R. et al. Development and characterisation of 140 new microsatellites in apple (Malus x domestica Borkh.). Mol. Breed. 10, 217–241 (2002).
Gianfranceschi, L., Seglias, N., Tarchini, R., Komjanc, M. & Gessler, C. Simple sequence repeats for the genetic analysis of apple. Theor. Appl. Genet. 96, 1069–1076 (1998).
Silfverberg-Dilworth, E. et al. Microsatellite markers spanning the apple (Malus x domestica Borkh.) genome. Tree Genet. Genomes 2, 202–224 (2006).
Cartwright, D.A., Troggio, M., Velasco, R. & Gutin, A. Genetic mapping in the presence of genotyping errors. Genetics 176, 2521–2527 (2007).
Uniprot Consortium. The universal protein resource (UniProt). Nucleic Acids Res. 36, D190–D195 (2008).
Wicker, T. et al. A unified classification system for eukaryotic transposable elements. Nat. Rev. Genet. 8, 973–982 (2007).
Solovyev, V., Kosarev, P., Seledsov, I. & Vorobyev, D. Automatic annotation of eukaryotic genes, pseudogenes and promoters. Genome Biol. 7, S10 (2006).
Korf, I., Flicek, P., Duan, D. & Brent, M.R. Integrating genomic homology into gene structure prediction. Bioinformatics 17, S140–S148 (2001).
Majoros, W.H., Pertea, M. & Salzberg, S.L. TigrScan and GlimmerHMM: two open source ab initio eukaryotic gene-finders. Bioinformatics 20, 2878–2879 (2004).
Birney, E., Clamp, M. & Durbin, R. GeneWise and genomewise. Genome Res. 14, 988–995 (2004).
Fontana, P., Cestaro, A., Velasco, R., Formentin, E. & Toppo, S. Rapid annotation of anonymous sequences from genome projects using semantic similarities and a weighting scheme in gene ontology. PLoS ONE 4, e4619 (2009).
Mulder, N. & Apweiler, R. InterPro and InterProScan—Tools for protein sequence classification and comparison. Methods Mol. Biol. 396, 59–70 (2007).
Altschul, S.F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402 (1997).
Enright, A.J., Van Dongen, S. & Ouzounis, C.A. An efficient algorithm for large-scale detection of protein families. Nucleic Acids Res. 30, 1575–1584 (2002).
Simillion, C., Janssens, K., Sterck, L. & Van de Peer, Y. i-ADHoRe 2.0: an improved tool to detect degenerated genomic homology using genomic profiles. Bioinformatics 24, 127–128 (2008).
Thompson, J.D., Higgins, D.G. & Gibson, T.J. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 22, 4673–4680 (1994).
Yang, Z. PAML: a program package for phylogenetic analysis by maximum likelihood. Comput. Appl. Biosci. 13, 555–556 (1997).
Notredame, C., Higgins, D.G. & Heringa, J. T-Coffee: a novel method for fast and accurate multiple sequence alignment. J. Mol. Biol. 302, 205–217 (2000).
Huelsenbeck, J.P. & Ronquist, F. MRBAYES: Bayesian inference of phylogenetic trees. Bioinformatics 17, 754–755 (2001).
Janick, J., Cummins, J.N., Brown, S.K. & Hemmat, M. Apples. in Fruit Breeding Vol. 1 (eds. Janick, J., Moore, J.J.) 1–77 (Wiley, New York, 1996).
Stephens, M., Smith, N.J. & Donnelly, P. A new statistical method for haplotype reconstruction from population data. Am. J. Hum. Genet. 68, 978–989 (2001).
Excoffier, L., Laval, G. & Schneider, S. Arlequin (version 3.0): an integrated software package for population genetics data analysis. Evol. Bioinform. Online 1, 47–50 (2005).
Acknowledgements
The Italian apple genome project was supported by the research office of the Provincia Autonoma di Trento. The US apple genome project was supported by Washington State University Agriculture Research Center, Washington Tree Fruit Research Commission and US Department of Agriculture National Research Initiative (USDA-NRI) grant 2008 −35300-04676 to A.D., A.K. and R.E.B. V.K. and C.W. received support from the USDA-NRI grant. S. Schaeffer and T.K. were supported by the US National Institutes of Health Protein Biotechnology Training Program and an Achievement Rewards for College Scientists fellowship. A.C.A., V.B., D.C., A.P.G., S.E.G., R.P.H. and R.N.C. were partially supported by the New Zealand Foundation for Research Science and Technology, contract no. C06X0812. We thank S. Attiya, E. Buglione and C. Celone from 454 Life Sciences-Roche Company as well as E. Stefani, A. Castelli and E. Potenza for technical support and V. Sgaramella for critical reading of the manuscript. Fosmid and shotgun libraries were prepared following the method developed by R. Meilan (Oregon State University).
Author information
Authors and Affiliations
Contributions
R. Velasco, A.D., A.K., R.E.B., M.S., M.E., F.S. and R. Viola managed the project. R. Velasco, A.Z., J.A., A.D., A.C., A.K., P.F., S.K.B., M.T., D.P., S. Salvi, J.L., A.G., R.E.B., S.E.G., M.S., M.E., Y.V.d.P. and F.S. designed the analyses. J.A., S.K.B., D.P., M. Pindo, G.C., D.M., G.E., L.M.F., N.G., T.M., J.T.M., J.R., B.W., C.K., Z.C., B.D., F.N., M. Palmer, T.K., D.J., S. Schaeffer, V.T.C., S.T.K., J.V., Q.T., A.M., A. Stormo, K.S. and R.B. performed DNA preparation and sequencing. R. Velasco, A.Z., A.D., A.C., A.K., P.F., M.T., D.P., P.B., V.C., A.D.R., M.K., P.M., D.M., P.L., Y.L., V.B., D.C., R.N.C., S.T., C.-E.D., A.G., R.E.B. and S.E.G. contributed to sequence assembly and anchoring to chromosomes. A.Z., A.D., A.C., A.K., P.F., M.T., S. Salvi, M. Pindo, S.C., M.C., F.C., V.G., S.L., G.M., M. Malnoy, D.M., M. Moretto, M. Perazzolli, A.S.-A., S.V., E.Z., V.K., C.W., D.E., A. Stella, A.V., M.M.K., S.M., A.C.A., R.N.C., A.P.G., E.L., J.A.F., S.P., P.R., L.S., S.T., B.L., R.P.H., Y.V.d.P. and F.S. contributed to automatic and manual genome annotation, genome structure and evolution analyses. R. Velasco, A.Z., J.A., A.D., A.K., P.F., M.T., S. Salvi, F.C., M. Malnoy, A.S.-A., S.V., R.E.B., A.V., S.M., J.A.F., L.S., S.T., B.L. and F.S. wrote the paper. M.M.K., A.C.A., R.N.C., A.P.G., R.P.H., C.-E.D., A.G., R.E.B., S.E.G., M.S., Y.V.d.P. and R. Viola revised the paper. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Note, Supplementary Tables 1–10 and 12–26 and Supplementary Figures 1–16 (PDF 10993 kb)
Supplementary Table 11
Resistance-related genes of Malus x domestica (XLS 197 kb)
Rights and permissions
About this article
Cite this article
Velasco, R., Zharkikh, A., Affourtit, J. et al. The genome of the domesticated apple (Malus × domestica Borkh.). Nat Genet 42, 833–839 (2010). https://doi.org/10.1038/ng.654
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.654
This article is cited by
-
Genome-wide investigation of UDP-Glycosyltransferase family in Tartary buckwheat (Fagopyrum tataricum)
BMC Plant Biology (2024)
-
Characterization of the actin (ACT) family in Rosaceae and role of PbrACT1 in pear pollen tube growth
Tree Genetics & Genomes (2024)
-
Water Loss: A Postharvest Quality Marker in Apple Storage
Food and Bioprocess Technology (2024)
-
Introduction of a diverse genetic background of Pyrus into Malus through intergeneric hybridization
Molecular Genetics and Genomics (2024)
-
Genomic-wide identification and expression analysis of R2R3-MYB transcription factors related to flavonol biosynthesis in Morinda officinalis
BMC Plant Biology (2023)