Abstract
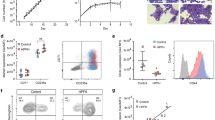
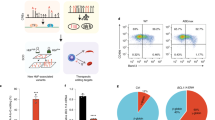
Hereditary persistence of fetal hemoglobin (HPFH) is characterized by persistent high levels of fetal hemoglobin (HbF) in adults. Several contributory factors, both genetic and environmental, have been identified1 but others remain elusive. HPFH was found in 10 of 27 members from a Maltese family. We used a genome-wide SNP scan followed by linkage analysis to identify a candidate region on chromosome 19p13.12–13. Sequencing revealed a nonsense mutation in the KLF1 gene, p.K288X, which ablated the DNA-binding domain of this key erythroid transcriptional regulator2. Only family members with HPFH were heterozygous carriers of this mutation. Expression profiling on primary erythroid progenitors showed that KLF1 target genes were downregulated in samples from individuals with HPFH. Functional assays suggested that, in addition to its established role in regulating adult globin expression, KLF1 is a key activator of the BCL11A gene, which encodes a suppressor of HbF expression3. These observations provide a rationale for the effects of KLF1 haploinsufficiency on HbF levels.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Serjeant, G. Geographic heterogeneity of sickle cell disease. in Disorders of Hemoglobin (eds. Steinberg, M.H., Forget, B.G., Higgs, D.R. & Nagel, R.L.) 895–906 (Cambridge Univ. Press, 2001).
Bieker, J.J. Probing the onset and regulation of erythroid cell-specific gene expression. Mt. Sinai J. Med. 72, 333–338 (2005).
Sankaran, V.G. et al. Human fetal hemoglobin expression is regulated by the developmental stage-specific repressor BCL11A. Science 322, 1839–1842 (2008).
Stamatoyannopoulos, G. & Grosveld, F. Hemoglobin switching. in The Molecular Basis of Blood Diseases (eds. Stamatoyannopoulos, G., Majerus, P.W., Perlmutter, R.M. & Varmus, H.) 135–182 (WB Saunders Company, Philadelphia, 2001).
Thein, S.L., Menzel, S., Lathrop, M. & Garner, C. Control of fetal hemoglobin: new insights emerging from genomics and clinical implications. Hum. Mol. Genet. 18, R216–R223 (2009).
Craig, J.E. et al. Dissecting the loci controlling fetal haemoglobin production on chromosomes 11p and 6q by the regressive approach. Nat. Genet. 12, 58–64 (1996).
Gilman, J.G. & Huisman, T.H. DNA sequence variation associated with elevated fetal G gamma globin production. Blood 66, 783–787 (1985).
Close, J. et al. Genome annotation of a 1.5 Mb region of human chromosome 6q23 encompassing a quantitative trait locus for fetal hemoglobin expression in adults. BMC Genomics 5, 33 (2004).
Garner, C. et al. Haplotype mapping of a major quantitative-trait locus for fetal hemoglobin production on chromosome 6q23. Am. J. Hum. Genet. 62, 1468–1474 (1998).
Lettre, G. et al. DNA polymorphisms at the BCL11A, HBS1L-MYB, and β-globin loci associate with fetal hemoglobin levels and pain crises in sickle cell disease. Proc. Natl. Acad. Sci. USA 105, 11869–11874 (2008).
Menzel, S. et al. A QTL influencing F cell production maps to a gene encoding a zinc-finger protein on chromosome 2p15. Nat. Genet. 39, 1197–1199 (2007).
Abecasis, G.R., Cherny, S.S., Cookson, W.O. & Cardon, L.R. Merlin—rapid analysis of dense genetic maps using sparse gene flow trees. Nat. Genet. 30, 97–101 (2002).
Hoffmann, K. & Lindner, T.H. easyLINKAGE-Plus—automated linkage analyses using large-scale SNP data. Bioinformatics 21, 3565–3567 (2005).
Leykin, I. et al. Comparative linkage analysis and visualization of high-density oligonucleotide SNP array data. BMC Genet. 6, 7 (2005).
Singleton, B.K., Burton, N.M., Green, C., Brady, R.L. & Anstee, D.J. Mutations in EKLF/KLF1 form the molecular basis of the rare blood group In(Lu) phenotype. Blood 112, 2081–2088 (2008).
Miller, I.J. & Bieker, J.J. A novel, erythroid cell-specific murine transcription factor that binds to the CACCC element and is related to the Kruppel family of nuclear proteins. Mol. Cell. Biol. 13, 2776–2786 (1993).
Feng, W.C., Southwood, C.M. & Bieker, J.J. Analyses of β-thalassemia mutant DNA interactions with erythroid Kruppel-like factor (EKLF), an erythroid cell-specific transcription factor. J. Biol. Chem. 269, 1493–1500 (1994).
Leberbauer, C. et al. Different steroids co-regulate long-term expansion versus terminal differentiation in primary human erythroid progenitors. Blood 105, 85–94 (2005).
Pilon, A.M. et al. Failure of terminal erythroid differentiation in EKLF-deficient mice is associated with cell cycle perturbation and reduced expression of E2F2. Mol. Cell. Biol. 28, 7394–7401 (2008).
Moffat, J. et al. A lentiviral RNAi library for human and mouse genes applied to an arrayed viral high-content screen. Cell 124, 1283–1298 (2006).
Antoniou, M. Induction of erythroid-specific expression in murine erythroleukemia (MEL) cell lines. Methods Mol. Biol. 7, 421–434 (1991).
Mühlemann, O., Eberle, A.B., Stalder, L. & Zamudio Orozco, R. Recognition and elimination of nonsense mRNA. Biochim. Biophys. Acta 1779, 538–549 (2008).
Donze, D., Townes, T.M. & Bieker, J.J. Role of erythroid Kruppel-like factor in human γ- to β-globin gene switching. J. Biol. Chem. 270, 1955–1959 (1995).
Drissen, R. et al. The active spatial organization of the β-globin locus requires the transcription factor EKLF. Genes Dev. 18, 2485–2490 (2004).
Zhou, D., Pawlik, K.M., Ren, J., Sun, C.W. & Townes, T.M. Differential binding of erythroid Krupple-like factor to embryonic/fetal globin gene promoters during development. J. Biol. Chem. 281, 16052–16057 (2006).
Bottardi, S., Ross, J., Pierre-Charles, N., Blank, V. & Milot, E. Lineage-specific activators affect beta-globin locus chromatin in multipotent hematopoietic progenitors. EMBO J. 25, 3586–3595 (2006).
Wijgerde, M. et al. The role of EKLF in human β-globin gene competition. Genes Dev. 10, 2894–2902 (1996).
Drissen, R. et al. The erythroid phenotype of EKLF-null mice: defects in hemoglobin metabolism and membrane stability. Mol. Cell. Biol. 25, 5205–5214 (2005).
Miller, S.A., Dykes, D.D. & Polesky, H.F. A simple salting out procedure for extracting DNA from human nucleated cells. Nucleic Acids Res. 16, 1215 (1988).
Trifillis, P., Ioannou, P., Schwartz, E. & Surrey, S. Identification of four novel δ-globin gene mutations in Greek Cypriots using polymerase chain reaction and automated fluorescence-based DNA sequence analysis. Blood 78, 3298–3305 (1991).
Craig, J.E., Barnetson, R.A., Prior, J., Raven, J.L. & Thein, S.L. Rapid detection of deletions causing δβ-thalassemia and hereditary persistence of fetal hemoglobin by enzymatic amplification. Blood 83, 1673–1682 (1994).
Higgs, D.R. & Wood, W.G. Genetic complexity in sickle cell disease. Proc. Natl. Acad. Sci. USA 105, 11595–11596 (2008).
O'Connell, J.R. & Weeks, D.E. PedCheck: a program for identification of genotype incompatibilities in linkage analysis. Am. J. Hum. Genet. 63, 259–266 (1998).
Schmittgen, T.D. & Livak, K.J. Analyzing real-time PCR data by the comparative C(T) method. Nat. Protoc. 3, 1101–1108 (2008).
Follenzi, A., Sabatino, G., Lombardo, A., Boccaccio, C. & Naldini, L. Efficient gene delivery and targeted expression to hepatocytes in vivo by improved lentiviral vectors. Hum. Gene Ther. 13, 243–260 (2002).
Zufferey, R., Nagy, D., Mandel, R.J., Naldini, L. & Trono, D. Multiply attenuated lentiviral vector achieves efficient gene delivery in vivo. Nat. Biotechnol. 15, 871–875 (1997).
Andrews, N.C. & Faller, D.V. A rapid micropreparation technique for extraction of DNA-binding proteins from limiting numbers of mammalian cells. Nucleic Acids Res. 19, 2499 (1991).
Follows, G.A. et al. Epigenetic consequences of AML1-ETO action at the human c-FMS locus. EMBO J. 22, 2798–2809 (2003).
Acknowledgements
We thank the family members for their cooperation; T. van Gent for help with statistical analysis; T. van Dijk for expertise in lentiviral technology; N. Papazian for preparation of human fetal liver samples; and M. Pizzuto for administrative support. The population samples from Malta were obtained from the Malta BioBank (EuroBioBank). Recombinant human erythropoietin (Epo) was a kind gift from Ortho-Biotech, and recombinant human Stem Cell Factor (SCF) was a kind gift from Amgen. Supported by the University of Malta and Mater Dei Hospital (A.E.F.); the Malta Government (J.B.); the European Molecular Biology Organization (J.B., P.P.); the Netherlands Scientific Organization (VENI 863.09.012 to L.G. and DN 82-294 and 912-07-019 to S.P.); the Netherlands Genomics Initiative, Erasmus MC (MRace; 296088 to S.P.); the Landsteiner Foundation for Blood Transfusion Research (0615 to S.P.); the Centre for Biomedical Genetics (F.G.G.); the European Commission FP6 EuTRACC consortium (037445 to F.G.G.); the US National Institutes of Health (NIH) (R01-HL073455 to F.G.G.); the Research Promotion Foundation of Cyprus (ΠΔE046_02 to G.P.P.); and the European Commission FP7 GEN2PHEN (200754 to G.P.P).
Author information
Authors and Affiliations
Contributions
F.G.G., A.E.F., G.P.P. and S.P. designed experiments. J.B., P.P., M.G., L.G., G.G., P.F., M.P., C.A.S., W.C., R.G., Z.Ö., N.G. and M.v.L. performed experiments. J.B., P.P., M.G., L.G. and G.G. analyzed results. P.J.v.d.S., F.G.G., A.E.F., G.P.P. and S.P. supervised data analysis. P.J.v.d.S., W.v.IJ. and M.B. provided expertise, analysis tools and infrastructure. A.J.M.H.V., J.H. and M.B. analyzed data. J.B., P.P., M.G., F.G.G., M.v.L., A.E.F., G.P.P. and S.P. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5, Supplementary Tables 1–3 and Supplementary Note (PDF 810 kb)
Rights and permissions
About this article
Cite this article
Borg, J., Papadopoulos, P., Georgitsi, M. et al. Haploinsufficiency for the erythroid transcription factor KLF1 causes hereditary persistence of fetal hemoglobin. Nat Genet 42, 801–805 (2010). https://doi.org/10.1038/ng.630
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.630
This article is cited by
-
Identification of a genomic DNA sequence that quantitatively modulates KLF1 transcription factor expression in differentiating human hematopoietic cells
Scientific Reports (2023)
-
Epigenomic analysis of KLF1 haploinsufficiency in primary human erythroblasts
Scientific Reports (2022)
-
Hemolysis contributes to anemia during long-duration space flight
Nature Medicine (2022)
-
Targeting Genetic Modifiers of HBG Gene Expression in Sickle Cell Disease: The miRNA Option
Molecular Diagnosis & Therapy (2022)
-
CD14+ monocytes repress gamma globin expression at early stages of erythropoiesis
Scientific Reports (2021)