Abstract
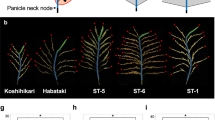
Identification of alleles that improve crop production and lead to higher-yielding varieties are needed for food security. Here we show that the quantitative trait locus WFP (WEALTHY FARMER'S PANICLE) encodes OsSPL14 (SQUAMOSA PROMOTER BINDING PROTEIN-LIKE 14, also known as IPA1). Higher expression of OsSPL14 in the reproductive stage promotes panicle branching and higher grain yield in rice. OsSPL14 controls shoot branching in the vegetative stage and is affected by microRNA excision. We also demonstrate the feasibility of using the OsSLP14WFP allele to increase rice crop yield. Introduction of the high-yielding OsSPL14WFP allele into the standard rice variety Nipponbare resulted in increased rice production.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Sasaki, T. From the editor's desk. Rice 1, 1–2 (2008).
Ashikari, M. et al. Cytokinin oxidase regulates rice grain production. Science 309, 741–745 (2005).
Ikeda-Kawakatsu, K. et al. Expression level of ABERRANT PANICLE ORGANIZATION1 determines rice inflorescence form through control of cell proliferation in the meristem. Plant Physiol. 150, 736–747 (2009).
Fan, C. et al. GS3, a major QTL for grain length and weight and minor QTL for grain width and thickness in rice, encodes a putative transmembrane protein. Theor. Appl. Genet. 112, 1164–1171 (2006).
Song, X.J., Huang, W., Shi, M., Zhu, M.Z. & Lin, H.X.A. QTL for rice grain width and weight encodes a previously unknown RING-type E3 ubiquitin ligase. Nat. Genet. 39, 623–630 (2007).
Shomura, A. et al. Deletion in a gene associated with grain size increased yields during rice domestication. Nat. Genet. 40, 1023–1028 (2008).
Huang, X. et al. Natural variation at the DEP1 locus enhances grain yield in rice. Nat. Genet. 41, 494–497 (2009).
Komatsu, K. et al. LAX and SPA: major regulators of shoot branching in rice. Proc. Natl. Acad. Sci. USA 100, 11765–11770 (2003).
Oikawa, T. & Kyozuka, J. Two-step regulation of LAX PANICLE1 protein accumulation in axillary meristem formation in rice. Plant Cell 21, 1095–1108 (2009).
Xie, K., Wu, C. & Xiong, L. Genomic organization, differential expression, and interaction of SQUAMOSA promoter-binding-like transcription factors and microRNA156 in rice. Plant Physiol. 142, 280–293 (2006).
Kakutani, T. Epi-alleles in plants: inheritance of epigenetic information over generations. Plant Cell Physiol. 43, 1106–1111 (2002).
Lindroth, A.M. et al. Requirement of CHROMOMETHYLASE3 for maintenance of CpXpG methylation. Science 292, 2077–2080 (2001).
Jackson, J., Lindroth, A., Cao, X. & Jacobsen, S. Control of CpNpG DNA methylation by the KRYPTONITE histone H3 methyltransferase. Nature 416, 556–560 (2002).
Zilberman, D., Cao, X. & Jacobsen, S. ARGONAUTE4 control of locus-specific siRNA accumulation and DNA and histone methylation. Science 299, 716–719 (2003).
Chan, S., Henderson, I. & Jacobsen, S. Gardening the genome: DNA methylation in Arabidopsis thaliana. Nat. Rev. Genet. 6, 351–360 (2005).
Kinoshita, Y. et al. Control of FWA gene silencing in Arabidopsis thaliana by SINE-related direct repeats. Plant J. 49, 38–45 (2007).
Saze, H., Shiraishi, A., Miura, A. & Kakutani, T. Control of genic DNA methylation by a jmjC domain-containing protein in Arabidopsis thaliana. Science 319, 462–465 (2008).
Miura, K. et al. A metastable DWARF1 epigenetic mutant affecting plant stature in rice. Proc. Natl. Acad. Sci. USA 106, 11218–11223 (2009).
Ashikari, M. & Matsuoka, M. Identification, isolation and pyramiding of quantitative trait loci for rice breeding. Trends Plant Sci. 11, 344–350 (2006).
Wu, G. et al. The sequential action of miR156 and miR172 regulates developmental timing in Arabidopsis. Cell 138, 750–759 (2009).
Wang, J.W., Czech, B. & Weigel, D. miR156-regulated SPL transcription factors define an endogenous flowering pathway in Arabidopsis thaliana. Cell 138, 738–749 (2009).
Wang, J.W., Schwab, R., Czech, B., Mica, E. & Weigel, D. Dual effects of miR156-targeted SPL genes and CYP78A5/KLUH on plastochron length and organ size in Arabidopsis thaliana. Plant Cell 20, 1231–1243 (2008).
Schwarz, S., Grande, A.V., Bujdoso, N., Saedler, H. & Huijser, P. The microRNA regulated SBP-box genes SPL9 and SPL15 control shoot maturation in Arabidopsis. Plant Mol. Biol. 67, 183–195 (2008).
Yamaguchi, A. et al. The microRNA-regulated SBP-Box transcription factor SPL3 is a direct upstream activator of LEAFY, FRUITFULL, and APETALA1. Dev. Cell 17, 268–278 (2009).
Lee, J., Park, J.J., Kim, S.L., Yim, J. & An, G. Mutations in the rice liguleless gene result in a complete loss of the auricle, ligule, and laminar joint. Plant Mol. Biol. 65, 487–499 (2007).
Lander, E.S. et al. MAPMAKER: an interactive computer package for constructing primary genetic linkage maps of experimental and natural populations. Genomics 1, 174–181 (1987).
Lander, E.S. & Botstein, D. Mapping Mendelian factors underlying quantitative traits using RFLP linkage maps. Genetics 121, 185–199 (1989).
Liu, Y.G. et al. Complementation of plant mutants with large genomic DNA fragments by a transformation-competent artificial chromosome vector accelerates positional cloning. Proc. Natl. Acad. Sci. USA 96, 6535–6540 (1999).
Hood, E.E., Gelvin, S.B., Melches, L.S. & Hoekema, A. New Agrobacterium helper plasmids for gene transfer to plants. Transgenic Res. 2, 208–218 (1993).
Kouchi, H. & Hata, S. Isolation and characterization of novel nodulin cDNAs representing genes expressed at early stages of soybean nodule development. Mol. Gen. Genet. 238, 106–119 (1993).
Chiu, W. et al. Engineered GFP as a vital reporter in plants. Curr. Biol. 6, 325–330 (1996).
Hattori, Y. et al. The ethylene response factors SNORKEL1 and SNORKEL2 allow rice to adapt to deep water. Nature 460, 1026–1030 (2009).
Acknowledgements
We thank K. Imamura for providing the detailed protocol for in situ hybridization, S. Mizuno for maintenance of the paddy field and E. Kouketsu and K. Sakata for helping to produce transgenic plants. This work was supported by a grant from the Ministry of Agriculture, Forestry and Fisheries of Japan (Integrated Research Project for Plants, Insects and Animals using Genome Technology, QTL-1001).
Author information
Authors and Affiliations
Contributions
H.K., M.M. and M.A. designed the research. K.M., M.I., A.M., X.-J.S., M.I. and K.A. conducted the research. K.M., X.-J.S., H.K. and M.A. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Note, Supplementary Figures 1–11 and Supplementary Tables 1 and 2. (PDF 1117 kb)
Rights and permissions
About this article
Cite this article
Miura, K., Ikeda, M., Matsubara, A. et al. OsSPL14 promotes panicle branching and higher grain productivity in rice. Nat Genet 42, 545–549 (2010). https://doi.org/10.1038/ng.592
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.592
This article is cited by
-
Comparative analysis of SPL transcription factors from streptophyte algae and embryophytes reveals evolutionary trajectories of SPL family in streptophytes
Scientific Reports (2024)
-
Breeding rice for yield improvement through CRISPR/Cas9 genome editing method: current technologies and examples
Physiology and Molecular Biology of Plants (2024)
-
Genome-wide identification and analysis of SPL gene family in chickpea (Cicer arietinum L.)
Protoplasma (2024)
-
Genetic and functional mechanisms of yield-related genes in rice
Acta Physiologiae Plantarum (2024)
-
Fine mapping of the panicle length QTL qPL5 in rice
Molecular Breeding (2024)