Abstract
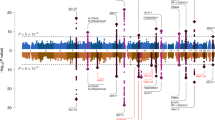
Saccular intracranial aneurysms are balloon-like dilations of the intracranial arterial wall; their hemorrhage commonly results in severe neurologic impairment and death. We report a second genome-wide association study with discovery and replication cohorts from Europe and Japan comprising 5,891 cases and 14,181 controls with ∼832,000 genotyped and imputed SNPs across discovery cohorts. We identified three new loci showing strong evidence for association with intracranial aneurysms in the combined dataset, including intervals near RBBP8 on 18q11.2 (odds ratio (OR) = 1.22, P = 1.1 × 10−12), STARD13-KL on 13q13.1 (OR = 1.20, P = 2.5 × 10−9) and a gene-rich region on 10q24.32 (OR = 1.29, P = 1.2 × 10−9). We also confirmed prior associations near SOX17 (8q11.23–q12.1; OR = 1.28, P = 1.3 × 10−12) and CDKN2A-CDKN2B (9p21.3; OR = 1.31, P = 1.5 × 10−22). It is noteworthy that several putative risk genes play a role in cell-cycle progression, potentially affecting the proliferation and senescence of progenitor-cell populations that are responsible for vascular formation and repair.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Rinkel, G.J., Djibuti, M., Algra, A. & van Gijn, J. Prevalence and risk of rupture of intracranial aneurysms: a systematic review. Stroke 29, 251–256 (1998).
Bilguvar, K. et al. Susceptibility loci for intracranial aneurysm in European and Japanese populations. Nat. Genet. 40, 1472–1477 (2008).
Iwamoto, H. et al. Prevalence of intracranial saccular aneurysms in a Japanese community based on a consecutive autopsy series during a 30-year observation period. The Hisayama study. Stroke 30, 1390–1395 (1999).
Salmela, E. et al. Genome-wide analysis of single nucleotide polymorphisms uncovers population structure in Northern Europe. PLoS One 3, e3519 (2008).
Jakkula, E. et al. The genome-wide patterns of variation expose significant substructure in a founder population. Am. J. Hum. Genet. 83, 787–794 (2008).
Marchini, J., Howie, B., Myers, S., McVean, G. & Donnelly, P. A new multipoint method for genome-wide association studies by imputation of genotypes. Nat. Genet. 39, 906–913 (2007).
Devlin, B. & Roeder, K. Genomic control for association studies. Biometrics 55, 997–1004 (1999).
Wakefield, J. A Bayesian measure of the probability of false discovery in genetic epidemiology studies. Am. J. Hum. Genet. 81, 208–227 (2007).
Wellcome Trust Case Control Consortium. Genome-wide association study of 14,000 cases of seven common diseases and 3,000 shared controls. Nature 447, 661–678 (2007).
Stephens, M. & Balding, D.J. Bayesian statistical methods for genetic association studies. Nat. Rev. Genet. 10, 681–690 (2009).
Wacholder, S., Chanock, S., Garcia-Closas, M., El Ghormli, L. & Rothman, N. Assessing the probability that a positive report is false: an approach for molecular epidemiology studies. J. Natl. Cancer Inst. 96, 434–442 (2004).
Helgadottir, A. et al. The same sequence variant on 9p21 associates with myocardial infarction, abdominal aortic aneurysm and intracranial aneurysm. Nat. Genet. 40, 217–224 (2008).
Kuro-o, M. et al. Mutation of the mouse klotho gene leads to a syndrome resembling ageing. Nature 390, 45–51 (1997).
Visel, A. et al. Targeted deletion of the 9p21 non-coding coronary artery disease risk interval in mice. Nature 464, 409–412 (2010).
Yun, M.H. & Hiom, K. CtIP-BRCA1 modulates the choice of DNA double-strand-break repair pathway throughout the cell cycle. Nature 459, 460–463 (2009).
Leung, T.H. et al. Deleted in liver cancer 2 (DLC2) suppresses cell transformation by means of inhibition of RhoA activity. Proc. Natl. Acad. Sci. USA 102, 15207–15212 (2005).
Urakawa, I. et al. Klotho converts canonical FGF receptor into a specific receptor for FGF23. Nature 444, 770–774 (2006).
Schievink, W.I. Genetics of intracranial aneurysms. Neurosurgery 40, 651–662 discussion 662–663 (1997).
Cannon Albright, L.A. et al. A genealogical assessment of heritable predisposition to aneurysms. J. Neurosurg. 99, 637–643 (2003).
Kamatani, Y. et al. A genome-wide association study identifies variants in the HLA-DP locus associated with chronic hepatitis B in Asians. Nat. Genet. 41, 591–595 (2009).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Clayton, D. & Leung, H.T. An R package for analysis of whole-genome association studies. Hum. Hered. 64, 45–51 (2007).
de Bakker, P. et al. Practical aspects of imputation-driven meta-analysis of genome-wide association studies. Hum. Mol. Genet. 17, R122–R128 (2008).
Clayton, D.G. et al. Population structure, differential bias and genomic control in a large-scale, case-control association study. Nat. Genet. 37, 1243–1246 (2005).
Patterson, N., Price, A.L. & Reich, D. Population structure and eigenanalysis. PLoS Genet. 2, e190 (2006).
Price, A.L. et al. Principal components analysis corrects for stratification in genome-wide association studies. Nat. Genet. 38, 904–909 (2006).
Belkin, M. & Niyogi, P. Laplacian eigenmaps for dimensionality reduction and data representation. Neural Comput. 15, 1373–1396 (2003).
Lee, A., Luca, D., Klei, L., Devlin, B. & Roeder, K. Discovering genetic ancestry using spectral graph theory. Genet. Epidemiol. 34, 51–59 (2009).
von Luxburg, U. A tutorial on spectral clustering. Stat. Comput. 17, 395–416 (2007).
Rosenbaum, P. A characterization of optimal designs for observational studies. J.R. Statist. Soc. B 53, 597–610 (1991).
Hansen, B. & Klopfer, S. Optimal full matching and related designs via network flows. J. Comput. Graph. Statist. 15, 609–627 (2006).
Wigginton, J., Cutler, D. & Abecasis, G. A note on exact tests of Hardy-Weinberg equilibrium. Am. J. Hum. Genet. 76, 887–893 (2005).
Browning, B. & Browning, S. A unified approach to genotype imputation and haplotype-phase inference for large data sets of trios and unrelated individuals. Am. J. Hum. Genet. 84, 210–223 (2009).
Breslow, N. & Day, N. Statistical methods in cancer research. Volume I—the analysis of case-control studies. IARC Sci. Publ. 5–338 (1980).
Viechtbauer, W. Bias and efficiency of meta-analytic variance estimators in the random-effects model. J. Educ. Behav. Stat. 30, 261–293 (2005).
Higgins, J., Thompson, S., Deeks, J. & Altman, D. Measuring inconsistency in meta-analyses. Br. Med. J. 327, 557–560 (2003).
Goodman, S. Toward evidence-based medical statistics. 2: The Bayes factor. Ann. Intern. Med. 130, 1005–1013 (1999).
Wakefield, J. Reporting and interpretation in genome-wide association studies. Int. J. Epidemiol. 37, 641–653 (2008).
Clayton, D. Prediction and interaction in complex disease genetics: experience in type 1 diabetes. PLoS Genet. 5, e1000540 (2009).
Acknowledgements
We are grateful to the participants who made this study possible. We thank A. Chamberlain, B. Meseck-Selchow and members of the Keck Foundation Biotechnology Resource Laboratory for their technical help. This study was supported by the Yale Center for Human Genetics and Genomics and the Yale Program on Neurogenetics, the US National Institutes of Health (NIH) grants R01NS057756 (M.G.) and U24 NS051869 (S.M.) and the Howard Hughes Medical Institute (R.P.L.). This study is partially funded by the European Commission under the 6th Framework Programme through the @neurIST (www.aneurist.org) project under contract no. FP6-IST-2004-027703. The Frankfurt case cohort collection was supported by Bundesministerium für Bildung und Forschung (01GI9907), Utrecht Control cohort by the Prinses Beatrix Fonds and the Adessium foundation (L.H.v.d.B.). S.M. was supported in part by the Clinical and Translational Science Award UL1 RR024139, National Center for Research Resources, NIH. We would also like to acknowledge the use of Yale University Biomedical High Performance Computing Center (NIH grant: RR19895).
Author information
Authors and Affiliations
Contributions
Study Cohorts: ascertainment, characterization and DNA preparation: M.N., M.v.u.z.F., E.G., J.E.J., J.H. and A.P. (Finnish case-control); Y.M.R. and G.J.E.R. (NL cases); P.B., T.D., J. Blasco, G.Z., P.S., R.R., T.S., C.M.F., P.S., A.F.F., V.E., M.C.J.M.S., P.L., J. Byrne, J.M. and D.R. (@neurIST case series); B.K., G.A., M.S., D.K., F.W., A.O., B.S., C.S., J. Beck, F.R., C.R., D.B., C.G., E.I.S., B.M., A.R. and H.S. (DE case series); A.T., A.H., H.K. and I.I. (JP1); S.-K.L., H.Z. and Y.N. (JP2). Control Cohorts: A.A., L.P. and A.P. (Health2000); A.A., L.P. and A.P. (NFBC1966); C.M.v.D. and M.M.B.B. (Rotterdam Study); L.H.v.d.B. and C.W. (Utrecht); T.I. and H.E.W. (KORA-gen); S.S. (PopGen). Genotyping: K.B., Z.A., N.N., A.K.O., E.G., S.M., R.P.L. and M.G. (Yale); C. Perret, C. Proust and F.C. (Aneurist); S.-K.L., H.Z. and Y.N. (JP2). Data management and informatics: K.Y., K.B., Z.A., N.N. and M.G. (Yale); S.-K.L., H.Z. and Y.N. (JP2 cohort); Statistical analysis: K.Y. and M.G. Writing team: K.Y., K.B., M.W.S., R.P.L. and M.G. Study design and analysis plan: K.Y., R.P.L. and M.G.
Corresponding authors
Ethics declarations
Competing interests
The authors have a provisional patent application under consideration based on the findings of this work.
Supplementary information
Supplementary Text and Figures
Supplementary Note, Supplementary Tables 1–5 and Supplementary Figures 1 and 2 (PDF 489 kb)
Rights and permissions
About this article
Cite this article
Yasuno, K., Bilguvar, K., Bijlenga, P. et al. Genome-wide association study of intracranial aneurysm identifies three new risk loci. Nat Genet 42, 420–425 (2010). https://doi.org/10.1038/ng.563
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.563
This article is cited by
-
Endothelial nitric oxide synthase rs1799983 gene polymorphism is associated with the risk of developing intracranial aneurysm
Acta Neurochirurgica (2023)
-
The quest to unravel the complex genomics of intracranial aneurysms
Nature Cardiovascular Research (2022)
-
Epigenetic landscapes of intracranial aneurysm risk haplotypes implicate enhancer function of endothelial cells and fibroblasts in dysregulated gene expression
BMC Medical Genomics (2021)
-
Endogenous animal models of intracranial aneurysm development: a review
Neurosurgical Review (2021)
-
PPIL4 is essential for brain angiogenesis and implicated in intracranial aneurysms in humans
Nature Medicine (2021)