Abstract
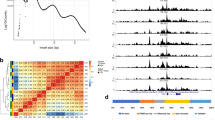
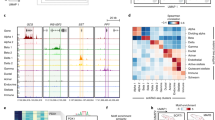
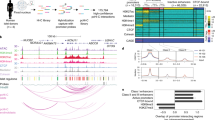
Tissue-specific transcriptional regulation is central to human disease1. To identify regulatory DNA active in human pancreatic islets, we profiled chromatin by formaldehyde-assisted isolation of regulatory elements2,3,4 coupled with high-throughput sequencing (FAIRE-seq). We identified ∼80,000 open chromatin sites. Comparison of FAIRE-seq data from islets to that from five non-islet cell lines revealed ∼3,300 physically linked clusters of islet-selective open chromatin sites, which typically encompassed single genes that have islet-specific expression. We mapped sequence variants to open chromatin sites and found that rs7903146, a TCF7L2 intronic variant strongly associated with type 2 diabetes5, is located in islet-selective open chromatin. We found that human islet samples heterozygous for rs7903146 showed allelic imbalance in islet FAIRE signals and that the variant alters enhancer activity, indicating that genetic variation at this locus acts in cis with local chromatin and regulatory changes. These findings illuminate the tissue-specific organization of cis-regulatory elements and show that FAIRE-seq can guide the identification of regulatory variants underlying disease susceptibility.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Birney, E. et al. Identification and analysis of functional elements in 1% of the human genome by the ENCODE pilot project. Nature 447, 799–816 (2007).
Giresi, P.G., Kim, J., McDaniell, R.M., Iyer, V.R. & Lieb, J.D. FAIRE (Formaldehyde-Assisted Isolation of Regulatory Elements) isolates active regulatory elements from human chromatin. Genome Res. 17, 877–885 (2007).
Hogan, G.J., Lee, C.K. & Lieb, J.D. Cell cycle-specified fluctuation of nucleosome occupancy at gene promoters. PLoS Genet. 2, e158 (2006).
Nagy, P.L., Cleary, M.L., Brown, P.O. & Lieb, J.D. Genomewide demarcation of RNA polymerase II transcription units revealed by physical fractionation of chromatin. Proc. Natl. Acad. Sci. USA 100, 6364–6369 (2003).
Grant, S.F. et al. Variant of transcription factor 7-like 2 (TCF7L2) gene confers risk of type 2 diabetes. Nat. Genet. 38, 320–323 (2006).
Bell, G.I. & Polonsky, K.S. Diabetes mellitus and genetically programmed defects in beta-cell function. Nature 414, 788–791 (2001).
Oliver-Krasinski, J.M. & Stoffers, D.A. On the origin of the beta cell. Genes Dev. 22, 1998–2021 (2008).
Henikoff, S. Nucleosome destabilization in the epigenetic regulation of gene expression. Nat. Rev. Genet. 9, 15–26 (2008).
Wallrath, L.L., Lu, Q., Granok, H. & Elgin, S.C. Architectural variations of inducible eukaryotic promoters: preset and remodeling chromatin structures. Bioessays 16, 165–170 (1994).
Gunton, J.E. et al. Loss of ARNT/HIF1beta mediates altered gene expression and pancreatic-islet dysfunction in human type 2 diabetes. Cell 122, 337–349 (2005).
Odom, D.T. et al. Control of pancreas and liver gene expression by HNF transcription factors. Science 303, 1378–1381 (2004).
Di Lorenzo, T.P., Peakman, M. & Roep, B.O. Translational mini-review series on type 1 diabetes: systematic analysis of T cell epitopes in autoimmune diabetes. Clin. Exp. Immunol. 148, 1–16 (2007).
McCarthy, M.I. & Zeggini, E. Genome-wide association scans for Type 2 diabetes: new insights into biology and therapy. Trends Pharmacol. Sci. 28, 598–601 (2007).
Sladek, R. et al. A genome-wide association study identifies novel risk loci for type 2 diabetes. Nature 445, 881–885 (2007).
Kim, T.H. et al. Analysis of the vertebrate insulator protein CTCF-binding sites in the human genome. Cell 128, 1231–1245 (2007).
Xi, H. et al. Identification and characterization of cell type-specific and ubiquitous chromatin regulatory structures in the human genome. PLoS Genet. 3, e136 (2007).
Bao, L., Zhou, M. & Cui, Y. CTCFBSDB: a CTCF-binding site database for characterization of vertebrate genomic insulators. Nucleic Acids Res. 36, D83–D87 (2008).
Heintzman, N.D. et al. Histone modifications at human enhancers reflect global cell-type-specific gene expression. Nature 459, 108–112 (2009).
Cuddapah, S. et al. Global analysis of the insulator binding protein CTCF in chromatin barrier regions reveals demarcation of active and repressive domains. Genome Res. 19, 24–32 (2009).
Atouf, F., Czernichow, P. & Scharfmann, R. Expression of neuronal traits in pancreatic beta cells. Implication of neuron-restrictive silencing factor/repressor element silencing transcription factor, a neuron-restrictive silencer. J. Biol. Chem. 272, 1929–1934 (1997).
Fujitani, Y. et al. Targeted deletion of a cis-regulatory region reveals differential gene dosage requirements for Pdx1 in foregut organ differentiation and pancreas formation. Genes Dev. 20, 253–266 (2006).
Gerrish, K., Van Velkinburgh, J.C. & Stein, R. Conserved transcriptional regulatory domains of the pdx-1 gene. Mol. Endocrinol. 18, 533–548 (2004).
Sander, M. et al. Homeobox gene Nkx6.1 lies downstream of Nkx2.2 in the major pathway of beta-cell formation in the pancreas. Development 127, 5533–5540 (2000).
Edwards, C.A. et al. The evolution of the DLK1–DIO3 imprinted domain in mammals. PLoS Biol. 6, e135 (2008).
Zeggini, E. et al. Meta-analysis of genome-wide association data and large-scale replication identifies additional susceptibility loci for type 2 diabetes. Nat. Genet. 40, 638–645 (2008).
Mohlke, K.L., Boehnke, M. & Abecasis, G.R. Metabolic and cardiovascular traits: an abundance of recently identified common genetic variants. Hum. Mol. Genet. 17, R102–R108 (2008).
Lyssenko, V. et al. Clinical risk factors, DNA variants, and the development of type 2 diabetes. N. Engl. J. Med. 359, 2220–2232 (2008).
Bouatia-Naji, N. et al. A polymorphism within the G6PC2 gene is associated with fasting plasma glucose levels. Science 320, 1085–1088 (2008).
Helgason, A. et al. Refining the impact of TCF7L2 gene variants on type 2 diabetes and adaptive evolution. Nat. Genet. 39, 218–225 (2007).
Guelen, L. et al. Domain organization of human chromosomes revealed by mapping of nuclear lamina interactions. Nature 453, 948–951 (2008).
Dillon, N. Gene regulation and large-scale chromatin organization in the nucleus. Chromosome Res. 14, 117–126 (2006).
Gilbert, N. et al. Chromatin architecture of the human genome: gene-rich domains are enriched in open chromatin fibers. Cell 118, 555–566 (2004).
Bucher, P. et al. Assessment of a novel two-component enzyme preparation for human islet isolation and transplantation. Transplantation 79, 91–97 (2005).
Latif, Z.A., Noel, J. & Alejandro, R. A simple method of staining fresh and cultured islets. Transplantation 45, 827–830 (1988).
Boj, S.F., Parrizas, M., Maestro, M.A. & Ferrer, J. A transcription factor regulatory circuit in differentiated pancreatic cells. Proc. Natl. Acad. Sci. USA 98, 14481–14486 (2001).
Li, H., Ruan, J. & Durbin, R. Mapping short DNA sequencing reads and calling variants using mapping quality scores. Genome Res. 18, 1851–1858 (2008).
Boyle, A.P., Guinney, J., Crawford, G.E. & Furey, T.S. F-Seq: a feature density estimator for high-throughput sequence tags. Bioinformatics 24, 2537–2538 (2008).
Rozowsky, J. et al. PeakSeq enables systematic scoring of ChIP-seq experiments relative to controls. Nat. Biotechnol. 27, 66–75 (2009).
Buck, M.J., Nobel, A.B. & Lieb, J.D. ChIPOTle: a user-friendly tool for the analysis of ChIP-chip data. Genome Biol. 6, R97 (2005).
Miller, W. et al. 28-way vertebrate alignment and conservation track in the UCSC Genome Browser. Genome Res. 17, 1797–1808 (2007).
Blanchette, M. et al. Genome-wide computational prediction of transcriptional regulatory modules reveals new insights into human gene expression. Genome Res. 16, 656–668 (2006).
Wingender, E. et al. TRANSFAC: an integrated system for gene expression regulation. Nucleic Acids Res. 28, 316–319 (2000).
Xie, X., Rigor, P. & Baldi, P. MotifMap: a human genome-wide map of candidate regulatory motif sites. Bioinformatics 25, 167–174 (2009).
Frith, M.C. et al. Detection of functional DNA motifs via statistical over-representation. Nucleic Acids Res. 32, 1372–1381 (2004).
Sandelin, A., Alkema, W., Engstrom, P., Wasserman, W.W. & Lenhard, B. JASPAR: an open-access database for eukaryotic transcription factor binding profiles. Nucleic Acids Res. 32, D91–D94 (2004).
Giardine, B. et al. Galaxy: a platform for interactive large-scale genome analysis. Genome Res. 15, 1451–1455 (2005).
Dennis, G. Jr. et al. DAVID: Database for Annotation, Visualization, and Integrated Discovery. Genome Biol. 4, 3 (2003).
Thomas, P.D. et al. PANTHER: a library of protein families and subfamilies indexed by function. Genome Res. 13, 2129–2141 (2003).
Su, A.I. et al. A gene atlas of the mouse and human protein-encoding transcriptomes. Proc. Natl. Acad. Sci. USA 101, 6062–6067 (2004).
Gudmundsson, J. et al. Two variants on chromosome 17 confer prostate cancer risk, and the one in TCF2 protects against type 2 diabetes. Nat. Genet. 39, 977–983 (2007).
Prokopenko, I. et al. Variants in MTNR1B influence fasting glucose levels. Nat. Genet. 41, 77–81 (2009).
Sandhu, M.S. et al. Common variants in WFS1 confer risk of type 2 diabetes. Nat. Genet. 39, 951–953 (2007).
Unoki, H. et al. SNPs in KCNQ1 are associated with susceptibility to type 2 diabetes in East Asian and European populations. Nat. Genet. 40, 1098–1102 (2008).
Yasuda, K. et al. Variants in KCNQ1 are associated with susceptibility to type 2 diabetes mellitus. Nat. Genet. 40, 1092–1097 (2008).
Luco, R.F. et al. A conditional model reveals that induction of hepatocyte nuclear factor-1alpha in Hnf1alpha-null mutant beta-cells can activate silenced genes postnatally, whereas overexpression is deleterious. Diabetes 55, 2202–2211 (2006).
Luco, R.F., Maestro, M.A., Sadoni, N., Zink, D. & Ferrer, J. Targeted deficiency of the transcriptional activator Hnf1alpha alters subnuclear positioning of its genomic targets. PLoS Genet. 4, e1000079 (2008).
Ishihara, H. et al. Pancreatic beta cell line MIN6 exhibits characteristics of glucose metabolism and glucose-stimulated insulin secretion similar to those of normal islets. Diabetologia 36, 1139–1145 (1993).
Acknowledgements
We thank X. Garcia for experimental support in rs7903146 functional studies, I. Moran for insights and help in the analysis of COREs, P. Maechler (Univ. Geneva Medical Center) for generously providing insulin release data, L. Piemonti and R. Nano (San Raffaele Scientific Institute), and M. Nacher (IDIBELL) for human islets. This work was supported by the European Union VI Framework Programme project Eurodia to J.F., Ministerio de Ciencia e Innovación (SAF2008-03116) to J.F., Juvenile Diabetes Research Foundation (26-2008-633 to J.F., 31-2008-416 to T.B., 6-2005-1178 and 31-2008-416 to A.S.), the US National Human Genome Research Institute Encyclopedia Of DNA Elements (NHGRI ENCODE) project (U54 HG004563 subcontract to J.D.L.) and R01 DK072193 (US National Institutes of Health) to K.L.M. K.L.M. is a Pew Scholar in the Biomedical Sciences.
Author information
Authors and Affiliations
Contributions
J.F. and J.D.L. conceived the study. K.J.G., T.N., L.P., J.M.S., K.L.M., J.D.L. and J.F. designed the experiments, interpreted results and wrote the manuscript. T.N. conducted FAIRE experiments, and developed and performed allelic imbalance assays. P.G.G. optimized the FAIRE protocol and performed microarray studies. K.J.G., J.M.S., and P.G.G. performed sequence analysis and K.J.G., L.P., T.N. and J.M.S. performed data analysis. L.P. conducted the analysis of COREs. M.P.F. and T.M.P. conducted reporter assays. P.M. conducted high-throughput sequencing. A.S., D.B., T.B. and E.M. provided purified human islet samples.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–4 and Supplementary Tables 1–9. (PDF 3941 kb)
Supplementary Table 2
RefSeq transcripts with preferential islet FAIRE enrichment (XLS 201 kb)
Supplementary Table 4
Over- and under-represented transcription factor binding motifs in intergenic islet FAIRE sites (XLS 57 kb)
Supplementary Table 5
Islet-selective Clusters of Open Regulatory Elements (COREs) (XLS 552 kb)
Supplementary Table 7
Islet-selective CORES that extend > 2 kb from the transcription start or termination site of overlapping genes (XLS 92 kb)
Rights and permissions
About this article
Cite this article
Gaulton, K., Nammo, T., Pasquali, L. et al. A map of open chromatin in human pancreatic islets. Nat Genet 42, 255–259 (2010). https://doi.org/10.1038/ng.530
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.530
This article is cited by
-
Chromatin accessibility differences between alpha, beta, and delta cells identifies common and cell type-specific enhancers
BMC Genomics (2023)
-
A genome-wide CRISPR screen identifies CALCOCO2 as a regulator of beta cell function influencing type 2 diabetes risk
Nature Genetics (2023)
-
Comprehensive assessment of differential ChIP-seq tools guides optimal algorithm selection
Genome Biology (2022)
-
Multidimensional chromatin profiling of zebrafish pancreas to uncover and investigate disease-relevant enhancers
Nature Communications (2022)
-
Genetic regulation of RNA splicing in human pancreatic islets
Genome Biology (2022)