Abstract
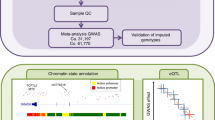
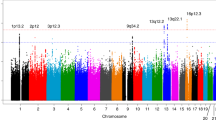
We conducted a genome-wide association study of pancreatic cancer in 3,851 affected individuals (cases) and 3,934 unaffected controls drawn from 12 prospective cohort studies and 8 case-control studies. Based on a logistic regression model for genotype trend effect that was adjusted for study, age, sex, self-described ancestry and five principal components, we identified eight SNPs that map to three loci on chromosomes 13q22.1, 1q32.1 and 5p15.33. Two correlated SNPs, rs9543325 (P = 3.27 × 10−11, per-allele odds ratio (OR) 1.26, 95% CI 1.18–1.35) and rs9564966 (P = 5.86 × 10−8, per-allele OR 1.21, 95% CI 1.13–1.30), map to a nongenic region on chromosome 13q22.1. Five SNPs on 1q32.1 map to NR5A2, and the strongest signal was at rs3790844 (P = 2.45 × 10−10, per-allele OR 0.77, 95% CI 0.71–0.84). A single SNP, rs401681 (P = 3.66 × 10−7, per-allele OR 1.19, 95% CI 1.11–1.27), maps to the CLPTM1L-TERT locus on 5p15.33, which is associated with multiple cancers. Our study has identified common susceptibility loci for pancreatic cancer that warrant follow-up studies.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Ferlay, J., Bray, F., Pisani, P. & Parkin, D.M. GLOBOCAN 2002: Cancer Incidence, Mortality and Prevalence Worldwide. IARC CancerBase vol. 5 (IARCPress, Lyon, 2004).
Anderson, K.E., Mack, T.M. & Silverman, D. Cancer of the pancreas. in Cancer Epidemiology and Prevention (ed. Schottenfeld, D. & Fraumeni, J.J.) 721–762 (Oxford University Press, New York, (2006)).
Shi, C., Hruban, R.H. & Klein, A.P. Familial pancreatic cancer. Arch. Pathol. Lab. Med. 133, 365–374 (2009).
Jones, S. et al. Exomic sequencing identifies PALB2 as a pancreatic cancer susceptibility gene. Science 324, 217 (2009).
Tischkowitz, M.D. et al. Analysis of the gene coding for the BRCA2-interacting protein PALB2 in familial and sporadic pancreatic cancer. Gastroenterology 137, 1183–1186 (2009).
Amundadottir, L. et al. Genome-wide association study identifies variants in the ABO locus associated with susceptibility to pancreatic cancer. Nat. Genet. 41, 986–990 (2009).
Wellcome Trust Case Control Consortium. Genome-wide association study of 14,000 cases of seven common diseases and 3,000 shared controls. Nature 447, 661–678 (2007).
Li, Y., Sung, W.K. & Liu, J.J. Association mapping via regularized regression analysis of single-nucleotide-polymorphism haplotypes in variable-sized sliding windows. Am. J. Hum. Genet. 80, 705–715 (2007).
Yu, Z. & Schaid, D.J. Sequential haplotype scan methods for association analysis. Genet. Epidemiol. 31, 553–564 (2007).
Dong, J.T. & Chen, C. Essential role of KLF5 transcription factor in cell proliferation and differentiation and its implications for human diseases. Cell. Mol. Life Sci. 66, 2691–2706 (2009).
Nakamura, Y. et al. Kruppel-like factor 12 plays a significant role in poorly differentiated gastric cancer progression. Int. J. Cancer 125, 1859–1867 (2009).
Chen, C. et al. Defining a common region of deletion at 13q21 in human cancers. Genes Chromosom. Cancer 31, 333–344 (2001).
Baudis, M. & Cleary, M.L. Progenetix.net: an online repository for molecular cytogenetic aberration data. Bioinformatics 17, 1228–1229 (2001).
Kainu, T. et al. Somatic deletions in hereditary breast cancers implicate 13q21 as a putative novel breast cancer susceptibility locus. Proc. Natl. Acad. Sci. USA 97, 9603–9608 (2000).
Paré, J.F. et al. The fetoprotein transcription factor (FTF) gene is essential to embryogenesis and cholesterol homeostasis and is regulated by a DR4 element. J. Biol. Chem. 279, 21206–21216 (2004).
Lee, Y.K. & Moore, D.D. Liver receptor homolog-1, an emerging metabolic modulator. Front. Biosci. 13, 5950–5958 (2008).
Botrugno, O.A. et al. Synergy between LRH-1 and beta-catenin induces G1 cyclin-mediated cell proliferation. Mol. Cell 15, 499–509 (2004).
Yamamoto, K., Okamoto, A., Isonishi, S., Ochiai, K. & Ohtake, Y. A novel gene, CRR9, which was up-regulated in CDDP-resistant ovarian tumor cell line, was associated with apoptosis. Biochem. Biophys. Res. Commun. 280, 1148–1154 (2001).
Bodnar, A.G. et al. Extension of life-span by introduction of telomerase into normal human cells. Science 279, 349–352 (1998).
Hahn, W.C. et al. Creation of human tumor cells with defined genetic elements. Nature 400, 464–468 (1999).
Kim, N.W. et al. Specific association of human telomerase activity with immortal cells and cancer. Science 266, 2011–2015 (1994).
Wang, Y. et al. Common 5p15.33 and 6p21.33 variants influence lung cancer risk. Nat. Genet. 40, 1407–1409 (2008).
McKay, J.D. et al. Lung cancer susceptibility locus at 5p15.33. Nat. Genet. 40, 1404–1406 (2008).
Rafnar, T. et al. Sequence variants at the TERT-CLPTM1L locus associate with many cancer types. Nat. Genet. 41, 221–227 (2009).
Stacey, S.N. et al. New common variants affecting susceptibility to basal cell carcinoma. Nat. Genet. 41, 909–914 (2009).
Shete, S. et al. Genome-wide association study identifies five susceptibility loci for glioma. Nat. Genet. 41, 899–904 (2009).
Landi, M.T. et al. A genome-wide association study of lung cancer identifies a region of chromosome 5p15 associated with risk for adenocarcinoma. Am. J. Hum. Genet. 85, 679–691 (2009).
Zienolddiny, S. et al. The TERT-CLPTM1L lung cancer susceptibility variant associates with higher DNA adduct formation in the lung. Carcinogenesis 30, 1368–1371 (2009).
Calado, R.T. et al. Constitutional hypomorphic telomerase mutations in patients with acute myeloid leukemia. Proc. Natl. Acad. Sci. USA 106, 1187–1192 (2009).
Savage, S.A. & Alter, B.P. Dyskeratosis congenita. Hematol. Oncol. Clin. North Am. 23, 215–231 (2009).
Yamaguchi, H. et al. Mutations in TERT, the gene for telomerase reverse transcriptase, in aplastic anemia. N. Engl. J. Med. 352, 1413–1424 (2005).
Calle, E.E. et al. The American Cancer Society Cancer Prevention Study II Nutrition Cohort: rationale, study design, and baseline characteristics. Cancer 94, 2490–2501 (2002).
The ATBC Cancer Prevention Study Group. The alpha-tocopherol, beta-carotene lung cancer prevention study: design, methods, participant characteristics, and compliance. Ann. Epidemiol. 4, 1–10 (1994).
Riboli, E. et al. European Prospective Investigation into Cancer and Nutrition (EPIC): study populations and data collection. Public Health Nutr. 5, 1113–1124 (2002).
Gallicchio, L. et al. Single nucleotide polymorphisms in inflammation-related genes and mortality in a community-based cohort in Washington County, Maryland. Am. J. Epidemiol. 167, 807–813 (2008).
Wolpin, B.M. et al. Circulating insulin-like growth factor binding protein-1 and the risk of pancreatic cancer. Cancer Res. 67, 7923–7928 (2007).
Zeleniuch-Jacquotte, A. et al. Postmenopausal levels of sex hormones and risk of breast carcinoma in situ: results of a prospective study. Int. J. Cancer 114, 323–327 (2005).
Hayes, R.B. et al. Methods for etiologic and early marker investigations in the PLCO trial. Mutat. Res. 592, 147–154 (2005).
Anderson, G.L. et al. Implementation of the Women's Health Initiative study design. Ann. Epidemiol. 13, S5–S17 (2003).
Rexrode, K.M., Lee, I.M., Cook, N.R., Hennekens, C.H. & Buring, J.E. Baseline characteristics of participants in the Women's Health Study. J. Womens Health Gend. Based Med. 9, 19–27 (2000).
Eppel, A., Cotterchio, M. & Gallinger, S. Allergies are associated with reduced pancreas cancer risk: a population-based case-control study in Ontario, Canada. Int. J. Cancer 121, 2241–2245 (2007).
Duell, E.J. et al. Detecting pathway-based gene-gene and gene-environment interactions in pancreatic cancer. Cancer Epidemiol. Biomarkers Prev. 17, 1470–1479 (2008).
Hassan, M.M. et al. Risk factors for pancreatic cancer: case-control study. Am. J. Gastroenterol. 102, 2696–2707 (2007).
Olson, S.H. et al. Allergies, variants in IL-4 and IL-4R alpha genes, and risk of pancreatic cancer. Cancer Detect. Prev. 31, 345–351 (2007).
Risch, H.A. Etiology of pancreatic cancer, with a hypothesis concerning the role of N-nitroso compounds and excess gastric acidity. J. Natl. Cancer Inst. 95, 948–960 (2003).
McWilliams, R.R. et al. Polymorphisms in DNA repair genes, smoking, and pancreatic adenocarcinoma risk. Cancer Res. 68, 4928–4935 (2008).
Wigginton, J.E., Cutler, D.J. & Abecasis, G.R. A note on exact tests of Hardy-Weinberg equilibrium. Am. J. Hum. Genet. 76, 887–893 (2005).
de Bakker, P.I. et al. Practical aspects of imputation-driven meta-analysis of genome-wide association studies. Hum. Mol. Genet. 17, R122–R128 (2008).
Pritchard, J.K., Stephens, M. & Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 155, 945–959 (2000).
Frazer, K.A. et al. A second generation human haplotype map of over 3.1 million SNPs. Nature 449, 851–861 (2007).
Thomas, G. et al. Multiple loci identified in a genome-wide association study of prostate cancer. Nat. Genet. 40, 310–315 (2008).
Hunter, D.J. et al. A genome-wide association study identifies alleles in FGFR2 associated with risk of sporadic postmenopausal breast cancer. Nat. Genet. 39, 870–874 (2007).
Yu, K. et al. Population substructure and control selection in genome-wide association studies. PLoS One 3, e2551 (2008).
Price, A.L. et al. Principal components analysis corrects for stratification in genome-wide association studies. Nat. Genet. 38, 904–909 (2006).
Patterson, N., Price, A.L. & Reich, D. Population structure and eigenanalysis. PLoS Genet. 2, e190 (2006).
Sun, L., Wilder, K. & McPeek, M.S. Enhanced pedigree error detection. Hum. Hered. 54, 99–110 (2002).
Clayton, D. Testing for association on the X chromosome. Biostatistics 9, 593–600 (2008).
Lettre, G., Lange, C. & Hirschhorn, J.N. Genetic model testing and statistical power in population-based association studies of quantitative traits. Genet. Epidemiol. 31, 358–362 (2007).
Higgins, J.P. & Thompson, S.G. Quantifying heterogeneity in a meta-analysis. Stat. Med. 21, 1539–1558 (2002).
Fearnhead, P. SequenceLDhot: detecting recombination hotspots. Bioinformatics 22, 3061–3066 (2006).
Fearnhead, P., Harding, R.M, Schneider, J.A, Myers, S. & Donnelly, P. Application of coalescent methods to reveal fine-scale rate variation and recombination hotspots. Genetics 167, 2067–2081 (2004).
Crawford, D.C. et al. Evidence for substantial fine-scale variation in recombination rates across the human genome. Nat. Genet. 36, 700–706 (2004).
Li, N.P. & Stephens, M. Modeling linkage disequilibrium and identifying recombination hotspots using single-nucleotide polymorphism data. Genetics 165, 2213–2233 (2003).
Acknowledgements
The authors gratefully acknowledge the energy and contribution of our late colleague Sheila Bingham. Additional acknowledgments are in the Supplementary Note.
Author information
Authors and Affiliations
Contributions
G.M.P., L.A., C.S.F., P.K., R.Z.S.-S., K.B.J., S.M.L., J.B.M., G.S.T., R.N.H., P.H. and S.J.C. organized and designed the study. L.A., A.H., K.B.J., G.T. and S.J.C. supervised genotyping of samples. L.A., P.K., R.Z.S.-S., C.S.F., K.B.J., C.K., H.P., Z.W., K.Y., R.N.H., P.H. and S.J.C. contributed to the design and execution of statistical analysis. L.A., G.M.P., P.K., R.Z.S.-S., R.N.H., P.H. and S.J.C. wrote the first draft of the manuscript. G.M.P., C.S.F., R.Z.S.-S., A.A.A., H.B.B., S.G., M.G., K.H., E.A.H., E.J.J., A.P.K., A.L., D.L., M.T.M., S.H.O., H.A.R., W.Z., D.A., W.R.B., C.D.B., M.-C.B.-R., J.E.B., P.M.B., F.C., S.C., M.C., M.deA., E.J.D., J.M.G., E.L.G., M.G., G.H., S.E.H., M.H., B.H., D.J.H., M.J., R.K., V.K., R.C.K., R.R.M., D.S.M., A.V.P., P.H.M.P., A.R., E.R., L.R., X.-O.S., A.T., D.T., S.K.V.D.E., J.V., J.W.-W., B.M.W., H.Y., A.Z.-J. and J.F.F.Jr. conducted the epidemiologic studies and contributed samples to the PanScan GWAS and/or replication. All authors contributed to the writing of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6, Supplementary Tables 1–3 and Supplementary Note (PDF 730 kb)
Rights and permissions
About this article
Cite this article
Petersen, G., Amundadottir, L., Fuchs, C. et al. A genome-wide association study identifies pancreatic cancer susceptibility loci on chromosomes 13q22.1, 1q32.1 and 5p15.33. Nat Genet 42, 224–228 (2010). https://doi.org/10.1038/ng.522
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.522
This article is cited by
-
Inhibiting NR5A2 targets stemness in pancreatic cancer by disrupting SOX2/MYC signaling and restoring chemosensitivity
Journal of Experimental & Clinical Cancer Research (2023)
-
A deep learning algorithm to predict risk of pancreatic cancer from disease trajectories
Nature Medicine (2023)
-
Exploring the Neandertal legacy of pancreatic ductal adenocarcinoma risk in Eurasians
Biological Research (2023)
-
Association between circulating vitamin E and ten common cancers: evidence from large-scale Mendelian randomization analysis and a longitudinal cohort study
BMC Medicine (2022)
-
Common variability in oestrogen-related genes and pancreatic ductal adenocarcinoma risk in women
Scientific Reports (2022)