Abstract
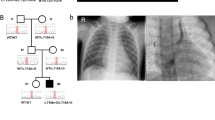
Tetralogy of Fallot (TOF), the most common severe congenital heart malformation, occurs sporadically, without other anomaly, and from unknown cause in 70% of cases. Through a genome-wide survey of 114 subjects with TOF and their unaffected parents, we identified 11 de novo copy number variants (CNVs) that were absent or extremely rare (<0.1%) in 2,265 controls. We then examined a second, independent TOF cohort (n = 398) for additional CNVs at these loci. We identified CNVs at chromosome 1q21.1 in 1% (5/512, P = 0.0002, OR = 22.3) of nonsyndromic sporadic TOF cases. We also identified recurrent CNVs at 3p25.1, 7p21.3 and 22q11.2. CNVs in a single subject with TOF occurred at six loci, two that encode known (NOTCH1, JAG1) disease-associated genes. Our findings predict that at least 10% (4.5–15.5%, 95% confidence interval) of sporadic nonsyndromic TOF cases result from de novo CNVs and suggest that mutations within these loci might be etiologic in other cases of TOF.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Ferencz, C. et al. Congenital heart disease: prevalence at livebirth. The Baltimore-Washington Infant Study. Am. J. Epidemiol. 121, 31–36 (1985).
Schott, J.J. et al. Congenital heart disease caused by mutations in the transcription factor NKX2–5. Science 281, 108–111 (1998).
Basson, C.T. et al. Mutations in human TBX5 cause limb and cardiac malformation in Holt-Oram syndrome. Nat. Genet. 15, 30–35 (1997).
Eldadah, Z.A. et al. Familial tetralogy of Fallot caused by mutation in the jagged1 gene. Hum. Mol. Genet. 10, 163–169 (2001).
Tomita-Mitchell, A., Maslen, C.L., Morris, C.D., Garg, V. & Goldmuntz, E. GATA4 sequence variants in patients with congenital heart disease. J. Med. Genet. 44, 779–783 (2007).
McDaniell, R. et al. NOTCH2 mutations cause Alagille syndrome, a heterogeneous disorder of the notch signaling pathway. Am. J. Hum. Genet. 79, 169–173 (2006).
Garg, V. et al. Mutations in NOTCH1 cause aortic valve disease. Nature 437, 270–274 (2005).
Goldmuntz, E. et al. Frequency of 22q11 deletions in patients with conotruncal defects. J. Am. Coll. Cardiol. 32, 492–498 (1998).
Yagi, H. et al. Role of TBX1 in human del22q11.2 syndrome. Lancet 362, 1366–1373 (2003).
Thienpont, B. et al. Submicroscopic chromosomal imbalances detected by array-CGH are a frequent cause of congenital heart defects in selected patients. Eur. Heart J. 28, 2778–2784 (2007).
Richards, A.A. et al. Cryptic chromosomal abnormalities identified in children with congenital heart disease. Pediatr. Res. 64, 358–363 (2008).
Loffredo, C.A. Epidemiology of cardiovascular malformations: prevalence and risk factors. Am. J. Med. Genet. 97, 319–325 (2000).
Korn, J.M. et al. Integrated genotype calling and association analysis of SNPs, common copy number polymorphisms and rare CNVs. Nat. Genet. 40, 1253–1260 (2008).
McCarroll, S.A. et al. Integrated detection and population-genetic analysis of SNPs and copy number variation. Nat. Genet. 40, 1166–1174 (2008).
Sato, M. et al. The validity of a rheumatoid arthritis medical records-based index of severity compared with the DAS28. Arthritis Res. Ther. 8, R57 (2006).
Redon, R. et al. Global variation in copy number in the human genome. Nature 444, 444–454 (2006).
Xu, B. et al. Strong association of de novo copy number mutations with sporadic schizophrenia. Nat. Genet. 40, 880–885 (2008).
Stefansson, H. et al. Large recurrent microdeletions associated with schizophrenia. Nature 455, 232–236 (2008).
Sebat, J. et al. Strong association of de novo copy number mutations with autism. Science 316, 445–449 (2007).
Christiansen, J. et al. Chromosome 1q21.1 contiguous gene deletion is associated with congenital heart disease. Circ. Res. 94, 1429–1435 (2004).
de Vries, B.B. et al. Diagnostic genome profiling in mental retardation. Am. J. Hum. Genet. 77, 606–616 (2005).
Brunetti-Pierri, N. et al. Recurrent reciprocal 1q21.1 deletions and duplications associated with microcephaly or macrocephaly and developmental and behavioral abnormalities. Nat. Genet. 40, 1466–1471 (2008).
Mefford, H.C. et al. Recurrent rearrangements of chromosome 1q21.1 and variable pediatric phenotypes. N. Engl. J. Med. 359, 1685–1699 (2008).
International Schizophrenia Consortium. Rare chromosomal deletions and duplications increase risk of schizophrenia. Nature 455, 237–241 (2008).
Walsh, T. et al. Rare structural variants disrupt multiple genes in neurodevelopmental pathways in schizophrenia. Science 320, 539–543 (2008).
Sharp, A.J. et al. Discovery of previously unidentified genomic disorders from the duplication architecture of the human genome. Nat. Genet. 38, 1038–1042 (2006).
Razzaque, M.A. et al. Germline gain-of-function mutations in RAF1 cause Noonan syndrome. Nat. Genet. 39, 1013–1017 (2007).
Pandit, B. et al. Gain-of-function RAF1 mutations cause Noonan and LEOPARD syndromes with hypertrophic cardiomyopathy. Nat. Genet. 39, 1007–1012 (2007).
Bassett, A.S. et al. Clinical features of 78 adults with 22q11 deletion syndrome. Am. J. Med. Genet. A. 138, 307–313 (2005).
McDonald-McGinn, D.M. et al. Phenotype of the 22q11.2 deletion in individuals identified through an affected relative: cast a wide FISHing net!. Genet. Med. 3, 23–29 (2001).
Niessen, K. & Karsan, A. Notch signaling in cardiac development. Circ. Res. 102, 1169–1181 (2008).
Krantz, I.D. et al. Spectrum and frequency of jagged1 (JAG1) mutations in Alagille syndrome patients and their families. Am. J. Hum. Genet. 62, 1361–1369 (1998).
Schubbert, S. et al. Germline KRAS mutations cause Noonan syndrome. Nat. Genet. 38, 331–336 (2006).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Stern, R.F. et al. Multiplex ligation-dependent probe amplification using a completely synthetic probe set. Biotechniques 37, 399–405 (2004).
Schouten, J.P. et al. Relative quantification of 40 nucleic acid sequences by multiplex ligation-dependent probe amplification. Nucleic Acids Res. 30, e57 (2002).
Yau, S.C., Bobrow, M., Mathew, C.G. & Abbs, S.J. Accurate diagnosis of carriers of deletions and duplications in Duchenne/Becker muscular dystrophy by fluorescent dosage analysis. J. Med. Genet. 33, 550–558 (1996).
Kim, J.B. et al. Polony multiplex analysis of gene expression (PMAGE) in mouse hypertrophic cardiomyopathy. Science 316, 1481–1484 (2007).
Acknowledgements
We gratefully acknowledge the participation of families. We also thank C. Sougnez and M. Parkin for technical assistance and R. Geggel for supplying TOF images. This work was supported by grants from the Howard Hughes Medical Institute (C.E.S.), US National Institutes of Health (to C.E.S., J.G.S. and R.E.B. and to the Broad Institute (National Center for Research Resources)), Pediatric Scientist Development Program (S.C.G.) and Sarnoff Cardiovascular Research Foundation (J.C.L.). Multiple sclerosis controls were genotyped in collaboration with Affymetrix, Inc.
Author information
Authors and Affiliations
Contributions
S.C.G., A.C.P., R.E.B., J.G.S. and C.E.S. designed the experiments. S.C.G., J.C.L., S.J.I. and J.M.G. performed the experiments. S.C.G., S.R.D., J.M.K., S.A.M., S.G., D.M.A., J.G.S. and C.E.S. were involved in genotyping and data analysis. E.E., J.H.C., A.C.P., S.M.M., M.d.L.Q.-D., M.A.A., R.D.E., R.M.P., N.A.S., M.E.W., P.L.D.J., D.A.H. and R.E.B. recruited subjects and collected DNA. S.C.G., J.G.S. and C.E.S. wrote the paper with input from all authors.
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–3 and Supplementary Tables 1–3 (PDF 2123 kb)
Rights and permissions
About this article
Cite this article
Greenway, S., Pereira, A., Lin, J. et al. De novo copy number variants identify new genes and loci in isolated sporadic tetralogy of Fallot. Nat Genet 41, 931–935 (2009). https://doi.org/10.1038/ng.415
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.415
This article is cited by
-
Genomic frontiers in congenital heart disease
Nature Reviews Cardiology (2022)
-
Best practices for variant calling in clinical sequencing
Genome Medicine (2020)
-
Dysregulation of Notch signaling in cardiac mesenchymal cells of patients with tetralogy of Fallot
Pediatric Research (2020)
-
Copy number variation analysis in bicuspid aortic valve-related aortopathy identifies TBX20 as a contributing gene
European Journal of Human Genetics (2019)
-
Notch and interacting signalling pathways in cardiac development, disease, and regeneration
Nature Reviews Cardiology (2018)