Abstract
Several mechanisms of action have been proposed for DNA methyltransferase and histone deacetylase inhibitors (DNMTi and HDACi), primarily based on candidate-gene approaches. However, less is known about their genome-wide transcriptional and epigenomic consequences. By mapping global transcription start site (TSS) and chromatin dynamics, we observed the cryptic transcription of thousands of treatment-induced non-annotated TSSs (TINATs) following DNMTi and HDACi treatment. The resulting transcripts frequently splice into protein-coding exons and encode truncated or chimeric ORFs translated into products with predicted abnormal or immunogenic functions. TINAT transcription after DNMTi treatment coincided with DNA hypomethylation and gain of classical promoter histone marks, while HDACi specifically induced a subset of TINATs in association with H2AK9ac, H3K14ac, and H3K23ac. Despite this mechanistic difference, both inhibitors convergently induced transcription from identical sites, as we found TINATs to be encoded in solitary long terminal repeats of the ERV9/LTR12 family, which are epigenetically repressed in virtually all normal cells.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Change history
18 August 2017
In the version of this article initially published, the e-mail address of corresponding author Christoph Plass was incorrect. The correct e-mail address is c.plass@dkfz.de. The error has been corrected in the HTML and PDF versions of the article.
References
Navada, S.C., Steinmann, J., Lübbert, M. & Silverman, L.R. Clinical development of demethylating agents in hematology. J. Clin. Invest. 124, 40–46 (2014).
West, A.C. & Johnstone, R.W. New and emerging HDAC inhibitors for cancer treatment. J. Clin. Invest. 124, 30–39 (2014).
Jones, P.A. At the tipping point for epigenetic therapies in cancer. J. Clin. Invest. 124, 14–16 (2014).
Chiappinelli, K.B. et al. Inhibiting DNA methylation causes an interferon response in cancer via dsRNA including endogenous retroviruses. Cell 162, 974–986 (2015).
Roulois, D. et al. DNA-demethylating agents target colorectal cancer cells by inducing viral mimicry by endogenous transcripts. Cell 162, 961–973 (2015).
Li, H. et al. Immune regulation by low doses of the DNA methyltransferase inhibitor 5-azacitidine in common human epithelial cancers. Oncotarget 5, 587–598 (2014).
Cameron, E.E., Bachman, K.E., Myöhänen, S., Herman, J.G. & Baylin, S.B. Synergy of demethylation and histone deacetylase inhibition in the re-expression of genes silenced in cancer. Nat. Genet. 21, 103–107 (1999).
Andersson, R. et al. An atlas of active enhancers across human cell types and tissues. Nature 507, 455–461 (2014).
Pertea, M. et al. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol. 33, 290–295 (2015).
Iyer, M.K. et al. The landscape of long noncoding RNAs in the human transcriptome. Nat. Genet. 47, 199–208 (2015).
Prensner, J.R. et al. The long noncoding RNA SChLAP1 promotes aggressive prostate cancer and antagonizes the SWI/SNF complex. Nat. Genet. 45, 1392–1398 (2013).
Almeida, L.G. et al. CTdatabase: a knowledge-base of high-throughput and curated data on cancer-testis antigens. Nucleic Acids Res. 37, D816–D819 (2009).
Ingolia, N.T., Lareau, L.F. & Weissman, J.S. Ribosome profiling of mouse embryonic stem cells reveals the complexity and dynamics of mammalian proteomes. Cell 147, 789–802 (2011).
van Heesch, S. et al. Extensive localization of long noncoding RNAs to the cytosol and mono- and polyribosomal complexes. Genome Biol. 15, R6 (2014).
Xie, M. et al. DNA hypomethylation within specific transposable element families associates with tissue-specific enhancer landscape. Nat. Genet. 45, 836–841 (2013).
Göke, J. et al. Dynamic transcription of distinct classes of endogenous retroviral elements marks specific populations of early human embryonic cells. Cell Stem Cell 16, 135–141 (2015).
Rafehi, H. et al. Vascular histone deacetylation by pharmacological HDAC inhibition. Genome Res. 24, 1271–1284 (2014).
Cohen, C.J., Lock, W.M. & Mager, D.L. Endogenous retroviral LTRs as promoters for human genes: a critical assessment. Gene 448, 105–114 (2009).
Sokol, M., Jessen, K.M. & Pedersen, F.S. Human endogenous retroviruses sustain complex and cooperative regulation of gene-containing loci and unannotated megabase-sized regions. Retrovirology 12, 32 (2015).
Hashimoto, K. et al. CAGE profiling of ncRNAs in hepatocellular carcinoma reveals widespread activation of retroviral LTR promoters in virus-induced tumors. Genome Res. 25, 1812–1824 (2015).
Yu, X. et al. The long terminal repeat (LTR) of ERV-9 human endogenous retrovirus binds to NF-Y in the assembly of an active LTR enhancer complex NF-Y/MZF1/GATA-2. J. Biol. Chem. 280, 35184–35194 (2005).
New, M., Olzscha, H. & La Thangue, N.B. HDAC inhibitor-based therapies: can we interpret the code? Mol. Oncol. 6, 637–656 (2012).
Klco, J.M. et al. Genomic impact of transient low-dose decitabine treatment on primary AML cells. Blood 121, 1633–1643 (2013).
de Parseval, N., Lazar, V., Casella, J.F., Benit, L. & Heidmann, T. Survey of human genes of retroviral origin: identification and transcriptome of the genes with coding capacity for complete envelope proteins. J. Virol. 77, 10414–10422 (2003).
Katoh, I. & Kurata, S. Association of endogenous retroviruses and long terminal repeats with human disorders. Front. Oncol. 3, 234 (2013).
Lander, E.S. et al. Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001).
La Mantia, G. et al. Identification of regulatory elements within the minimal promoter region of the human endogenous ERV9 proviruses: accurate transcription initiation is controlled by an Inr-like element. Nucleic Acids Res. 20, 4129–4136 (1992).
Lania, L. et al. Structural and functional organization of the human endogenous retroviral ERV9 sequences. Virology 191, 464–468 (1992).
Ling, J. et al. The solitary long terminal repeats of ERV-9 endogenous retrovirus are conserved during primate evolution and possess enhancer activities in embryonic and hematopoietic cells. J. Virol. 76, 2410–2423 (2002).
Pi, W. et al. Long-range function of an intergenic retrotransposon. Proc. Natl. Acad. Sci. USA 107, 12992–12997 (2010).
Krönung, S.K. et al. LTR12 promoter activation in a broad range of human tumor cells by HDAC inhibition. Oncotarget 7, 33484–33497. (2016).
Beyer, U., Krönung, S.K., Leha, A., Walter, L. & Dobbelstein, M. Comprehensive identification of genes driven by ERV9-LTRs reveals TNFRSF10B as a re-activatable mediator of testicular cancer cell death. Cell Death Differ. 23, 64–75 (2016).
Sherrill-Mix, S., Ocwieja, K.E. & Bushman, F.D. Gene activity in primary T cells infected with HIV89.6: intron retention and induction of genomic repeats. Retrovirology 12, 79 (2015).
Lock, F.E. et al. Distinct isoform of FABP7 revealed by screening for retroelement-activated genes in diffuse large B-cell lymphoma. Proc. Natl. Acad. Sci. USA 111, E3534–E3543 (2014).
Mak, K.S. et al. Repression of chimeric transcripts emanating from endogenous retrotransposons by a sequence-specific transcription factor. Genome Biol. 15, R58 (2014).
Wiesner, T. et al. Alternative transcription initiation leads to expression of a novel ALK isoform in cancer. Nature 526, 453–457 (2015).
Vizoso, M. et al. Epigenetic activation of a cryptic TBC1D16 transcript enhances melanoma progression by targeting EGFR. Nat. Med. 21, 741–750 (2015).
Chiappinelli, K.B., Zahnow, C.A., Ahuja, N. & Baylin, S.B. Combining epigenetic and immunotherapy to combat cancer. Cancer Res. 76, 1683–1689 (2016).
Juergens, R.A. et al. Combination epigenetic therapy has efficacy in patients with refractory advanced non-small cell lung cancer. Cancer Discov. 1, 598–607 (2011).
Kim, K. et al. Eradication of metastatic mouse cancers resistant to immune checkpoint blockade by suppression of myeloid-derived cells. Proc. Natl. Acad. Sci. USA 111, 11774–11779 (2014).
Schroeder, T. et al. Azacitidine and donor lymphocyte infusions as first salvage therapy for relapse of AML or MDS after allogeneic stem cell transplantation. Leukemia 27, 1229–1235 (2013).
Steinmann, J. et al. 5-Azacytidine and DLI can induce long-term remissions in AML patients relapsed after allograft. Bone Marrow Transplant. 50, 690–695 (2015).
Rizvi, N.A. et al. Cancer immunology. Mutational landscape determines sensitivity to PD-1 blockade in non-small cell lung cancer. Science 348, 124–128 (2015).
Bhuvanagiri, M. et al. 5-azacytidine inhibits nonsense-mediated decay in a MYC-dependent fashion. EMBO Mol. Med. 6, 1593–1609 (2014).
Setiadi, A.F. et al. Epigenetic enhancement of antigen processing and presentation promotes immune recognition of tumors. Cancer Res. 68, 9601–9607 (2008).
Peng, D. et al. Epigenetic silencing of TH1-type chemokines shapes tumour immunity and immunotherapy. Nature 527, 249–253 (2015).
Almstedt, M. et al. The DNA demethylating agent 5-aza-2′-deoxycytidine induces expression of NY-ESO-1 and other cancer/testis antigens in myeloid leukemia cells. Leuk. Res. 34, 899–905 (2010).
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106 (2010).
Zhao, M., Kim, P., Mitra, R., Zhao, J. & Zhao, Z. TSGene 2.0: an updated literature-based knowledgebase for tumor suppressor genes. Nucleic Acids Res. 44, D1023–D1031 (2015).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Kong, L. et al. CPC: assess the protein-coding potential of transcripts using sequence features and support vector machine. Nucleic Acids Res. 35, W345–W349 (2007).
Nielsen, M. et al. NetMHCpan, a method for quantitative predictions of peptide binding to any HLA-A and -B locus protein of known sequence. PLoS One 2, e796 (2007).
Zhao, X., Valen, E., Parker, B.J. & Sandelin, A. Systematic clustering of transcription start site landscapes. PLoS One 6, e23409 (2011).
Kim, D., Langmead, B. & Salzberg, S.L. HISAT: a fast spliced aligner with low memory requirements. Nat. Methods 12, 357–360 (2015).
Haberle, V., Forrest, A.R., Hayashizaki, Y., Carninci, P. & Lenhard, B. CAGEr: precise TSS data retrieval and high-resolution promoterome mining for integrative analyses. Nucleic Acids Res. 43, e51 (2015).
Lawrence, M. et al. Software for computing and annotating genomic ranges. PLoS Comput. Biol. 9, e1003118 (2013).
Robinson, M.D., McCarthy, D.J. & Smyth, G.K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Lister, R. et al. Human DNA methylomes at base resolution show widespread epigenomic differences. Nature 462, 315–322 (2009).
Wang, Q. et al. Tagmentation-based whole-genome bisulfite sequencing. Nat. Protoc. 8, 2022–2032 (2013).
Hansen, K.D., Langmead, B. & Irizarry, R.A. BSmooth: from whole genome bisulfite sequencing reads to differentially methylated regions. Genome Biol. 13, R83 (2012).
Rice, P., Longden, I. & Bleasby, A. EMBOSS: the European molecular biology open software suite. Trends Genet. 16, 276–277 (2000).
Claus, R. et al. Quantitative analyses of DAPK1 methylation in AML and MDS. Int. J. Cancer. 131, E138–E142 (2012).
Calvo, S.E., Pagliarini, D.J. & Mootha, V.K. Upstream open reading frames cause widespread reduction of protein expression and are polymorphic among humans. Proc. Natl. Acad. Sci. USA 106, 7507–7512 (2009).
Kozak, M. An analysis of 5′-noncoding sequences from 699 vertebrate messenger RNAs. Nucleic Acids Res. 15, 8125–8148 (1987).
Jackson, R.J., Hellen, C.U. & Pestova, T.V. The mechanism of eukaryotic translation initiation and principles of its regulation. Nat. Rev. Mol. Cell Biol. 11, 113–127 (2010).
Grillo, G. et al. UTRdb and UTRsite (RELEASE 2010): a collection of sequences and regulatory motifs of the untranslated regions of eukaryotic mRNAs. Nucleic Acids Res. 38, D75–D80 (2010).
Smith, T.F. & Waterman, M.S. Identification of common molecular subsequences. J. Mol. Biol. 147, 195–197 (1981).
Acknowledgements
We thank C. Weigel, R. Toth, and K. Chiappinelli for helpful feedback and discussions and A. Tanay for critical proofreading of the manuscript. We also thank K. Bauer, K. Klimo, M. Bähr and the Genomics and Proteomics Core Facility at the German Cancer Research Center for their excellent technical support and expertise. D.B. is supported by the German-Israeli Helmholtz Research School in Cancer Biology. D.L., J.L., H.S.J., B.Z., and T.W. are supported by the American Cancer Society grant RSG-14-049-01-DMC and NIH grants R01HG007354, R01HG007175, R01ES024992, U01CA200060, U01HG009391, and U24ES 026699. This work was supported in part by the Helmholtz Association, the DFG funded Priority Program SPP1463, the German Center for Lung Research (DZL), and BMBF funded CancerEpiSys program and the ICGC Data-Mining project.
Author information
Authors and Affiliations
Contributions
C.R.S., M.D., D.B., D.B.L., S.L., M.S.I., H.B., S.H., M.H., A.L., A.R., G.S., J.S., R.W., J.P.M., K.R., D.W., C.C.O., and C.P. designed the experiments and performed experimental work. D.B., C.S., M.D., D.L., J.L., H.S.J., N.M.S., Y.H., B.Z., Y.A., C.D.I., B.B., and T.W. performed data analysis. I.O., O.W., and M.L. provided clinical expertise and data. D.B., M.D., T.W., and C.P. prepared the manuscript and figures. T.W., C.G., B.B., M.E., C.C.O., and C.P. provided project leadership. All authors contributed to the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Schematic representation of the experimental approach used in this study.
NCI-H1299 cells containing a dual fluorescence/resistance reporter at the epigenetically silenced DAPK1 locus were treated with DAC, SAHA, SB939, or DAC+SB as indicated. After treatment, cells were harvested and subjected to downstream transcriptomic or epigenomic profiling. CAGE = Cap-analysis of gene expression; ChIP = Chromatin immunoprecipitation; WGBS = Whole-genome bisulfite sequencing.
Supplementary Figure 2 Engineering of reporter cell line, epigenetic compound screen, and cryptic transcripts.
a) Schematic cloning overview of the dual-fluorescence/resistance reporter construct. Key components are depicted by the numbers 1-10. Arrows indicate activity of the CMV and SFFV promoter driving mCherry-Zeo and HSV-TK expression, respectively. HA-L/R = homology arm left/right; SA-2A = Splice-acceptor with T2A self-cleavage peptide; pA = polyA termination sequence; CMV = Cytomegalovirus promoter; SFFV = Spleen focus-forming virus promoter; HSV-TK = Herpex Simplex Virus Thymidine Kinase.
b) Predicted model of the DAPK1 locus after successful reporter integration. The reporter integration (Rep) results in a novel fusion transcript containing DAPK1 exons 1-3, spliced to the EGFP-NEO cassette. The T2A cleavage peptide creates separate proteins for EGFP-NEO and a truncated DAPK1 protein upon translation. mCherry-ZEO is constitutively expressed from the CMV promoter.
c) MassARRAY EpiTYPER quantitative DNA methylation analysis of the DAPK1 promoter CpG island in NCI-H1299 reporter cells after increasing DAC concentrations. DAPK1 methylation is calculated as average methylation value of all covered CpG-units.
d) DAPK1 mRNA levels and EGFP fluorescence after increasing doses of DAC. Expression values were normalized to housekeeping genes and depicted relative to the individual mock treated sample (left axis). Percentage of EGFP positive cells relative to untreated cells (right axis).
e) DNMT1 protein levels assessed via western blot after knockdown with a DNMT1-specific shRNA. ACTB = β-Actin.
f) DAPK1 expression (black) and promoter methylation (grey) after knockdown of DNMT1. Expression is normalized to housekeeping genes and depicted relative to the individual NT-control treated sample. Promoter methylation was calculated as average methylation value of all CpG-units covered. NT = non-targeting.
g) Relative viability of NCI-H1299 reporter cells after compound treatment and G418 selection. Cells were seeded 24 h prior to compound treatment in 96-well plates in triplicates. Compounds were refreshed every 24 h for 72 h and then G418-selection performed for 10 d. After selection, cell viability was quantified by Calcein-AM assay and normalized to DMSO controls. All bars represent mean values ± SD (n=3). Results are displayed sorted by drug target and quantified viability.
h) Genome browser view of the newly identified intragenic transcripts at the DAPK1 locus (ALPHA, BETA, and GAMMA). All three transcripts initiated in the intron between DAPK1 exons 2 and 3. Key functional annotations were derived from the ENCODE cell line data and visualized relative to the transcript start sites to characterize the genomic architecture in the vicinity of the cryptic TSSs: Enhancer mark H3K27Ac, transcription factor binding, transposable elements, and vertebrate conservation. LTR = long terminal repeat, SINE = short interspersed nuclear element, LINE = long interspersed nuclear element, DNA = DNA transposon.
Supplementary Figure 3 CAGE QC and characterization of TINATs
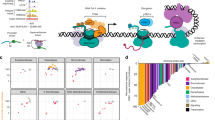
a) Principle component analysis based on the variance stabilized expression1 of the 1000 most variable CAGE-clusters. For each treatment condition (DMSO = purple; SB939 = orange; SAHA = green; DAC = blue; DAC+SB = red) both replicates are shown.
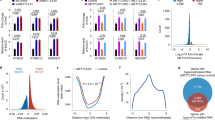
b) Boxplot showing normalized CAGE-tags for TSSs associated with 116 AIMs previously described to be ubiquitously up-regulated after DNMTi across three cancer entities2. AIM expression levels in NCI-H1299 cells are shown after DMSO, DAC, SAHA, SB939, and DAC+SB treatment. Statistical significance was calculated using the Wilcoxon and Mann-Whitney two-sided test.
c) Boxplot showing normalized CAGE-tags for TSSs associated with 276 CTAs3. Statistical significance was calculated using the Wilcoxon and Mann-Whitney two-sided test.
d) CAGE-tags mapping to Syncytin-1 (ERVW-1) before and after treatment in NCI-H1299 cells. Reads were aligned against the ERVW-1 sequence (hg38) using HISAT4 and coverage was calculated using the R/Bioconductor package GenomicAlignments5.
e) Comparison of expression changes relative to DMSO for DAC against SB939 (left) and SB939 against SAHA (right).
f) Gene ontology enrichment of SB939 (orange) and DAC+SB (red) TINAT proximal genes calculated by GREAT6. 224,491 TSS coordinates consisting of our and previously described TSSs (Gencode.v19.TSS.notlow.gff) were used as background regions. The top 5 most significantly enriched biological processes are shown. DAC TINATs showed no enrichment.
g) TINAT expression correlates with expression of nearby genes. Spearman correlation for the expression of each TINAT – nearest gene pair was calculated across all conditions (green) and compared to the null distribution obtained by permuting sample labels (red).
h) Density plot showing the difference in directionality based on CAGE tags located within 1400 bps of TINATs or 200 ubiquitous cell line enhancers (green)7. Directionality scores of ±1 or 0 correspond to perfectly strand-specific or balanced transcription, respectively.
i) TINAT expression across various human tissue and cell types from the FANTOM57 project. TINATs less than 150 bps away from a FANTOM5 CAGE cluster supported by at least one tag were considered expressed in a given sample.
Supplementary Figure 4 Genomic distribution of TINATs
Chromosome-wise genomic distribution of TINATs identified after DAC (blue), SB939 (orange), and DAC+SB (red) treatment.
Supplementary Figure 5 Translation of TINATs into proteins
a) NetMHCpan8 was used to predict the binding affinity of 12 major HLA alleles (columns) for 178 DAC+SB out-of-frame peptide sequences (rows). Shown are the log-transformed binding affinity IC50 values (values > 0.426 correspond to an IC50 value stronger than 500 nM). Binding affinity ranges from low (white) to high (brown). The presence of a TINAT within the adult thymus is displayed.
b) Coupled in vitro transcription and translation of TINAT-exon fusion candidates. cDNAs of DAC+SB treated NCI-H1299 reporter cells was PCR-amplified with gene-specific primers introducing a T7 promoter at the 5’ end (top panel). PCR fragments were transcribed and translated in the presence of radioactively labeled methionine. Translation products were size-separated and visualized by autoradiography (bottom panel). The full-length canonical CRYGC mRNA served as positive control.
c) Polysome profiles of NCI-H1299 cells treated with (red) or without (black) harringtonine (top). 40S, 60S, and 80S subunits are highlighted. tRNA, 18S, and 28S rRNA levels in the presence (red) or absence (black) of harringtonine were measured across 14 fractions by ethidium bromide (EtBr) staining (middle). Lower panel shows Ncl mRNA levels as well as the spike-in control rabbit beta-globin as quantified by northern blot (NB).
d) Distribution of GAPDH, HPRT1 mRNA as well as nine TINAT-exon fusion transcripts along 14 polysome fractions isolated from DAC+SB treated NCI-H1299 cells in the presence (red) or absence (blue) of harringtonine. Colored squares below horizontal axis line indicate the fraction where half of the mRNAs have accumulated. The mean of three biological replicates is shown and error bars denote standard deviation.
e) Distribution of GAPDH, HPRT1 mRNA as well as 28 TINAT-exon fusion transcripts along 5 polysome fractions isolated from DAC+SB treated NCI-H1299 cells in the presence (red) or absence (blue) of harringtonine. Colored squares below horizontal axis line indicate the fraction where half of the mRNAs have accumulated. The mean of two biological replicates is shown and error bars denote standard deviation.
f) Expression levels of transcripts overlapping TINAT coordinates across different polysome fractions in untreated LS-174 T-pTER-β-catenin cells9. Intronic or intergenic TINAT coordinates were extended downstream by 500 bps or to the next exon and the overlap with bowtie hg19 aligned reads was counted and normalized by the total number of uniquely mapped reads. Expression levels of 21 TINAT candidate transcripts with a normalized read count >1 in at least one fraction are shown as Z-scores from -1.5 (white) to 1.5 (red).
Supplementary Figure 6 Original gel images
Western blot analysis of histone post-translational modifications at different time points following the treatment with SB939 or DMSO. Samples are arranged from left to right: DMSO 24h, SB939 24h, DMSO 9h, SB939 9h, DMSO 6h, SB939 6h, DMSO 3h, SB939 3h, DMSO 0h.
Supplementary Figure 7 Additional histone modifications around TINATs
a) ChIP-seq occupancy plots showing the average level of the indicated histone modifications 5 kb up- and downstream of all identified TINATs. Shaded areas indicate the 95% confidence interval and numbers indicate the normalized read counts.
b) Average level of the indicated histone modifications 5 kb up- and downstream of all identified TINATs after SAHA treatment. Shaded areas indicate the 95% confidence interval and numbers indicate the normalized read counts.
c) Within group sum of squares as a function of the number of k-means clusters.
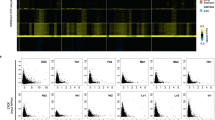
d) H3K27me3 (darkred), H3K4me1 (green), H3K36ac (orange), H2AK9ac (violet), H3K4ac (red), H3K18ac (blue), H3K23ac (brown), H4K8ac (lightgreen), and H4K12ac (lightred) levels around TINATs after DMSO (green bar), DAC (blue bar), SB939 (orange bar), or DAC+SB (red bar) treatment. Color intensity of the histone modifications represents Z-scores. TINATs are arranged as in Fig 4d.
Supplementary Figure 8 Characterization of transposable elements associated with TINATs
a) Normalized CAGE-tag expression levels of TINATs overlapping with LTRs (red), TEs other than LTRs (blue), or no TEs (green) after DAC (left), SB939 (middle), or DAC+SB (right) treatment. Y-axis is logarithmic.
b) Genomic distribution of LTR12 copies, all other LTRs, as well as a random set. HOMER10 software was used to annotate genomic features.
c) Enrichment of certain TE families across the three chromatin clusters identified in Figure 4d. Point size represents the significance of the overrepresentation which was calculated cluster-wise for each TE family using Fisher’s exact test. Color represents fold-enrichment (observed vs expected).
d) Fraction of TINATs overlapping with LTRs of the LTR12 subfamily, other LTRs, or non-LTR genomic regions.
e) Heatmap of normalized read counts for the 20 most significant differentially expressed TE subfamilies between DMSO and SAHA treated primary vascular endothelial cells11. The FDR corrected p-value for differential expression is shown in the bar plot to the right. CPM = counts per million
f) LTR12C expression in NCI-H1299 reporter cells normalized to housekeepers after HDACi (SB939) or treatment with different chemotherapeutic agents relative to DMSO. The results from a single experiment are shown.
g) Frequency of TINAT start positions along the 1577bp LTR12C consensus sequence. Height of the bar depicts TINAT frequency after DAC (green), SB939 (purple), and DACSB (red) treatment.
h) ChIP-seq occupancy plots showing the average intensity [reads per kilobase per million mapped reads (RPKM)] of H3K4me3 and H3K9ac for LTR12Cs without (left panel) and with TINATs (right panel) relative to the TINAT coordinates. For both groups, histone signal intensities are shown before (black/grey) and after (colored) treatment with DAC+SB.
i) qRT-PCR expression analysis of GATA2 transcript levels relative to housekeepers after different treatments in the presence (blue) or absence (red) of siRNAs targeting GATA2. Error bars represent standard deviation.
References
1 Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome biology 11, R106, doi:10.1186/gb-2010-11-10-r106 (2010).
2 Li, H. et al. Immune regulation by low doses of the DNA methyltransferase inhibitor 5-azacitidine in common human epithelial cancers. Oncotarget 5, 587-598, doi:10.18632/oncotarget.1782 (2014).
3 Almeida, L. G. et al. CTdatabase: a knowledge-base of high-throughput and curated data on cancer-testis antigens. Nucleic acids research 37, D816-819, doi:10.1093/nar/gkn673 (2009).
4 Kim, D., Langmead, B. & Salzberg, S. L. HISAT: a fast spliced aligner with low memory requirements. Nature methods 12, 357-360, doi:10.1038/nmeth.3317 (2015).
5 Lawrence, M. et al. Software for computing and annotating genomic ranges. PLoS computational biology 9, e1003118, doi:10.1371/journal.pcbi.1003118 (2013).
6 McLean, C. Y. et al. GREAT improves functional interpretation of cis-regulatory regions. Nature biotechnology 28, 495-501, doi:10.1038/nbt.1630 (2010).
7 Andersson, R. et al. An atlas of active enhancers across human cell types and tissues. Nature 507, 455-461, doi:10.1038/nature12787 (2014).
8 Nielsen, M. et al. NetMHCpan, a method for quantitative predictions of peptide binding to any HLA-A and -B locus protein of known sequence. PloS one 2, e796, doi:10.1371/journal.pone.0000796 (2007).
9 van Heesch, S. et al. Extensive localization of long noncoding RNAs to the cytosol and mono- and polyribosomal complexes. Genome biology 15, R6, doi:10.1186/gb-2014-15-1-r6 (2014).
10 Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Molecular cell 38, 576-589, doi:10.1016/j.molcel.2010.05.004 (2010).
11 Rafehi, H. et al. Vascular histone deacetylation by pharmacological HDAC inhibition. Genome research 24, 1271-1284, doi:10.1101/gr.168781.113 (2014).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8, Supplementary Table 2, 6, 7–11, and Supplementary Note 1. (PDF 3359 kb)
Supplementary Table 1
Sequencing summary. (XLSX 17 kb)
Supplementary Table 3
Details on ORF cloning. (XLSX 18 kb)
Supplementary Table 4
TINAT number per LTR family for each treatment. (XLSX 18 kb)
Supplementary Table 5
LTR12C literature comparison. (XLSX 26 kb)
Supplementary Dataset 1
Consensus clusters. (XLSX 10270 kb)
Supplementary Dataset 2
StringTie assembled transcripts. (ZIP 263 kb)
Rights and permissions
About this article
Cite this article
Brocks, D., Schmidt, C., Daskalakis, M. et al. DNMT and HDAC inhibitors induce cryptic transcription start sites encoded in long terminal repeats. Nat Genet 49, 1052–1060 (2017). https://doi.org/10.1038/ng.3889
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3889
This article is cited by
-
Towards targeting transposable elements for cancer therapy
Nature Reviews Cancer (2024)
-
Transposable Elements: Emerging Therapeutic Targets in Neurodegenerative Diseases
Neurotoxicity Research (2024)
-
T-bet+ B cells are activated by and control endogenous retroviruses through TLR-dependent mechanisms
Nature Communications (2024)
-
Proceedings of the inaugural Dark Genome Symposium: November 2022
Mobile DNA (2023)
-
Targeting the epigenome to reinvigorate T cells for cancer immunotherapy
Military Medical Research (2023)