Abstract
The emergence of apomixis—the transition from sexual to asexual reproduction—is a prominent feature of modern citrus. Here we de novo sequenced and comprehensively studied the genomes of four representative citrus species. Additionally, we sequenced 100 accessions of primitive, wild and cultivated citrus. Comparative population analysis suggested that genomic regions harboring energy- and reproduction-associated genes are probably under selection in cultivated citrus. We also narrowed the genetic locus responsible for citrus polyembryony, a form of apomixis, to an 80-kb region containing 11 candidate genes. One of these, CitRWP, is expressed at higher levels in ovules of polyembryonic cultivars. We found a miniature inverted-repeat transposable element insertion in the promoter region of CitRWP that cosegregated with polyembryony. This study provides new insights into citrus apomixis and constitutes a promising resource for the mining of agriculturally important genes.
Similar content being viewed by others
Main
Asexual reproduction is a remarkable feature of perennial fruit crops that facilitates the faithful propagation of commercially valuable individuals by avoiding the uncertainty associated with the sexual reproduction of hybrid plants. Many fruit crops reproduce or are propagated by asexual mechanisms. These mechanisms include natural means of propagation, such as layering, cutting and grafting1, and modern technologies, such as tissue culture. Apomixis is a naturally occurring mode of asexual reproduction that yields offspring that are genetically identical to the mother plant2. It is uncommon in agriculturally important crops apart from citrus and apple3. The first example of apomixis in citrus was reported in 1719, when Leeuwenhoek observed the development of two plantlets from the same seed4. A better understanding of apomixis and its introgression into agronomically important crops could potentially help revolutionize modern agriculture5. As demonstrated by fruit crops, apomixis enables breeders to fix valuable traits and heterozygosity. Most previous studies have focused on gametophytic apomixis2,6. Sporophytic apomixis, including nucellar embryony, is not as well studied.
Apomixis in citrus is sporophytic—the embryos develop from somatic nucellar cells—and very stable among commercial varieties, including sweet oranges, mandarins, grapefruits and lemons7. In general, 2–10 embryos develop within a seed. However, in particular genotypes, 30 or more embryos can develop in one seed3,8. This phenomenon is typically called 'polyembryony' and is considered an important trait for breeding purposes. On the one hand, polyembryony is widely employed in citrus nurseries and propagation programs to generate large numbers of uniform rootstocks from seeds. On the other hand, polyembryony has sometimes caused problems for breeding that required sexual crosses. Data from crosses between monoembryonic and polyembryonic cultivars indicate that one or two dominant loci control polyembryony9,10. Molecular markers linked to a polyembryony locus have been reported7,11. This locus was later mapped to a genomic region that spans approximately 380 kb12. Genes differentially expressed during polyembryogenesis have also been identified13,14,15.
Citrus fruit trees, designated as Citrinae, are a large group belonging to the subfamily Aurantioideae and the family Rutaceae16. Citrinae are classified into primitive citrus, near citrus and true citrus on the basis of botanical characteristics16. Atalantia buxifolia, also known as Chinese box orange and formerly named Severinia buxifolia, is an example of a primitive citrus species. The true citrus group comprises almost all of the commercially cultivated citrus, with major varieties belonging to the Citrus genus (Supplementary Note 1). The best known citrus varieties are the sweet orange (Citrus sinensis), mandarin (Citrus reticulata), lemon (Citrus limon), grapefruit (Citrus paradisi) and pummelo (Citrus grandis or Citrus maxima). There are a range of reproductive systems among the different varieties of primitive and modern citrus species that include sexual reproduction, clonal propagation and mixtures of these systems.
We aimed to understand the genomic characteristics of primitive and cultivated citrus and the genetic basis of asexual reproduction in citrus. We de novo sequenced pummelo and three representative species of Citrinae. We also sequenced a population of 100 citrus accessions that ranged from primitive to cultivated citrus. Our genetic analyses of segregating and natural populations, and our transcriptome profiling, provide new insights into the genetic basis of apomixis in citrus.
Results
Citrinae phylogeny and reproductive systems
Conserved single-copy genes from eight species of Citrinae were used for the construction of a phylogenetic tree (Fig. 1a). As expected, the primitive citrus Atalantia was located at a basal position in the tree. The spiny and wild species Citrus ichangensis (Ichang papeda, hereafter referred to as papeda) is phylogenetically located between primitive and cultivated citrus. The three basic species, C. reticulata (mandarin), C. grandis (pummelo) and C. medica (citron) showed close relationships to each other. Primitive, wild and cultivated citrus each have nine haploid chromosomes (Supplementary Fig. 1). Citrus species exhibit broad sexual compatibility. Interspecific and intergeneric hybridizations are feasible even between primitive and cultivated citrus17. Our experiments also support the grafting compatibility and partial sexual compatibility of Atalantia and Citrus (Supplementary Fig. 2). Atalantia and papeda are mainly seed propagated, with large or many seeds that mostly fill the locules of the fruit (Fig. 1b), and are not suitable for propagation by grafting owing to strong and numerous prickles and stiff twigs (Supplementary Fig. 3). In contrast, the vegetative propagule that facilitates propagation by grafting has an ideal morphology in mandarin, pummelo, sweet orange and other cultivated citrus (Supplementary Fig. 3).
(a) Citrinae phylogeny and genomes. A phylogenetic tree was constructed using the maximum likelihood method. Asterisks indicate the de novo genomes sequenced in this study. Circles indicate previously published genomes. Purple indicates primitive citrus; green indicates wild citrus; brown indicates the three basic species of citrus. Bootstrap values >60 are indicated. (b) Differences among the fruits and seeds from primitive, wild and cultivated citrus. Cross-sections of young (left) and mature (right) fruit from different species are shown. The length of each line is based on the percentage of seed weight in the fruit. Scale bars, 1 cm.
A high-quality and three draft genomes of Citrinae
A high-quality genome of citrus was generated from haploid pummelo (Supplementary Fig. 4) by single-molecule sequencing. A total of 56.8× coverage of single-molecule sequences on the PacBio RS II platform via a shotgun approach was used for genome assembly (Supplementary Table 1). Additionally, 370.3× Illumina sequencing data were used to correct sequencing errors and to fill in the gaps (Supplementary Fig. 5 and Supplementary Table 2). The contig N50 and N90 of the final assembly were 2.2 Mb and 70 kb, respectively. Moreover, the scaffold N50 and N90 were 4.2 Mb and 565 kb, respectively (Table 1). The sequence contiguity represents 18-fold and a 44-fold improvements of the C. clementina and C. sinensis genomes, respectively, as evaluated by contig N50 (refs. 18,19). Ninety percent of the assembly is in 101 scaffolds larger than 565 kb. The largest scaffold is 14.3 Mb. A total of 117 scaffolds, accounting for 87% of the assembled genome, were anchored by 1,563 molecular markers (Supplementary Figs. 6 and 7 and Supplementary Table 3).
Three draft genomes of heterozygous Citrinae species—atalantia (A. buxifolia, primitive citrus), papeda (C. ichangensis, wild citrus) and citron (C. medica) (Fig. 1a)—were sequenced and assembled with Illumina shotgun sequencing reads. High-coverage sequencing data that ranged from 164.4× to 401.9× were produced from multiple libraries, with small insert sizes that ranged from 140 bp to 500 bp and long insert sizes of 2 kb, 5 kb and 20 kb (Supplementary Tables 4–6). The sequenced reads were assembled using the de Bruijn graph algorithm from the SOAPdenovo package20, resulting in scaffold N50 values of 1,074 kb, 504 kb and 367 kb (with contig N50 values of 23.9 kb, 76.6 kb and 46.5 kb) for atalantia, papeda and citron, respectively (Supplementary Table 7).
Ab initio gene prediction programs, homology searches and RNA-seq analysis were integrated to annotate the pummelo genome. We identified 30,123 protein-coding genes with 42,886 transcripts. The gene transcripts had an average length of 1,572 bp, a mean coding sequence size of 1,141 bp and an average exon length of 277 bp. RNA-seq data revealed that 12,763 gene loci (42.4%) encoded two or more transcriptional isoforms. Regarding the core and dispensable portions of these genomes, all of the genes in the six genomes (29,655 in sweet orange; 24,533 in clementine mandarin; 30,123 in pummelo; 32,579 in citron; 32,067 in papeda; 28,420 in atalantia) (Supplementary Table 7) were classified into 23,829 gene families on the basis of the homology of their encoded proteins. A total of 12,457 (52%) gene families were shared by all six of these genomes (Supplementary Fig. 8 and Supplementary Table 8). Further analysis indicated a similar number of genes in almost all of the transcription factor gene families in the six citrus genomes (Supplementary Tables 9 and 10).
Genomic characteristics of citrus
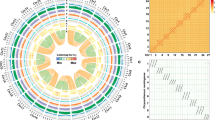
The four sequenced species from this study (atalantia, papeda, citron and pummelo), and the two previously published sequenced species (sweet orange and clementine mandarin)18,19 range from primitive to cultivated citrus (Fig. 1). An overview of the genome synteny and sequence variation is presented in Figure 2 and Supplementary Figures 9–13. The genome sizes are similar among the Citrus species. We observed the most variation in genome size between Atalantia and the Citrus species (Supplementary Table 11 and Supplementary Note 2). On the basis of our presence–absence variation (PAV) analysis, 33.28% of the unique regions in pummelo (relative to atalantia) contained transposable elements (TEs), and 2.68% of the unique regions contained protein-coding genes (Supplementary Table 12). Approximately one-fifth of the genes in the unique regions (153 of 780 annotated genes) of pummelo were predicted to have known functions. These genes were significantly enriched (P value < 0.05 and FDR < 0.05) in three GO terms: defense response and proteolysis (biological process category) and pectate lyase activity (molecular function category) relative to the entire genome of pummelo (Supplementary Fig. 14). Among the 153 genes with functional annotations, 93 are homologous to one or more genes (60.78%) (Supplementary Table 13). Conversely, the unique regions in atalantia relative to pummelo had a higher TE content (54.20%) than the entire atalantia genome (43.55%).
SNVs were confidently identified by sequence alignment (Supplementary Fig. 15) and filtered using resequencing data. A pairwise comparison of each species versus pummelo showed that the transition/transversion ratios varied from 1.38 to 1.66, with an average of 1.57 (Supplementary Table 14). The distributions of SNVs in gene structures displayed the highest density in the 5′ and 3′ UTRs (an average of 18.8 and 17.2 SNVs per kb for 5′ and 3′ UTRs, respectively). The second highest density of SNVs was in introns (an average of 14.2 SNVs per kb). The lowest density of SNVs was in coding sequence (CDS) regions (an average of 9.7 SNVs per kb) (Supplementary Table 15).
The ratio of nonsynonymous to synonymous substitutions (dN/dS) was calculated for each of the 8,551 single-copy orthologs shared by these six citrus genomes. We found that 207, 100, 104, 66, 106 and 77 genes showed higher dN/dS values in atalantia, papeda, citron, pummelo, clementine mandarin and sweet orange (Supplementary Table 16), respectively, relative to other species. For the dN/dS of single-copy orthologous genes, 12.79% of the genes showed a value greater than one in atalantia (Supplementary Table 17). This proportion was higher than the average value of 6.82% from other citrus (Supplementary Tables 18–22), indicating greater sequence divergence in atalantia relative to other citrus. These genes may have functional associations with the biological features of atalantia. For instance, Cg4g018790 encodes a cytokinin oxidase/dehydrogenase homologous to the rice OsCKX2 gene, which has been reported to control seed number21. In this gene, higher sequence polymorphisms were observed in atalantia relative to other citrus species (Supplementary Fig. 16).
Genetic diversity and comparative population analysis
Citrinae species exhibit high levels of morphological diversity (Fig. 3a), which is a reflection of their diverse genetic backgrounds. We selected and sequenced 100 citrus accessions with an average depth of 30-fold genome coverage (Supplementary Table 23). Sequencing data from three mandarins, one orange and four pummelos from previous studies18,19 were also analyzed. This set of 108 citrus accessions (Supplementary Fig. 17) was used for population diversity analysis, genetic differentiation analysis and association analysis of the apomixis trait. For comparative population analysis, 15 atalantia accessions, 11 papeda accessions and 19 pummelo accessions, representing primitive, wild and cultivated citrus, were used (Fig. 3b and Supplementary Fig. 18). The fixation index (FST) values among these citrus populations fluctuated around 0.28 (Supplementary Fig. 19), which indicates moderate genetic differentiations among citrus populations. For a comparison, the FST was 0.34 between two wild populations of common bean and 0.44 between wild and cultivated cucumber22,23.
(a) The diversity of citrus germplasms in this study. Inner circle, primitive citrus; middle circle, wild and cultivated citrus; outer circle, cultivated citrus. (b) Maximum likelihood phylogenetic tree of 45 citrus accessions (15 atalantia, 11 papeda and 19 pummelo) used in the comparative population analysis. Bootstrap values are indicated in the tree. (c) The distribution of π along the chromosomes among the populations of atalantia, papeda and pummelo, respectively. These values were calculated in 2-Mb sliding windows with 1-Mb steps. (d) Distribution of the reduction of diversity (log10 π ratios) and differentiation (FST values) from the pairwise comparison of the atalantia and pummelo populations. π ratios were calculated as πatalantia/πpummelo in 50-kb sliding windows with 10-kb steps. The region above the dashed line in the distribution of log10 π ratios corresponds to the 5% right tail of the empirical distribution (log10 π ratio = 0.36). The regions above the dashed line in the FST values distribution are in the 5% right tail of the empirical distribution (FST is 0.40).
Nucleotide diversity (π) analysis indicated that atalantia has the highest genetic diversity and that papeda has a higher level of genetic diversity than pummelo (Fig. 3c). We observed a dramatic reduction in genetic diversity between atalantia and papeda. Additional sequence analysis indicated that atalantia has more highly diverged regions (threefold more highly divergent regions, on average) than the other two populations (Supplementary Table 24). For example, a long region at the end of chromosome 5 (39.64–49.38 Mb, about one-fifth of the chromosome) displayed the highest diversity in atalantia and a dramatic reduction in diversity in papeda and pummelo (Fig. 3c).
Genomic regions that showed both high differentiation and reduced diversity were identified among the primitive, wild and cultivated citrus (Supplementary Fig. 20 and Supplementary Tables 25–27). The results of pairwise comparisons indicated that the distribution of such regions was uneven along the chromosomes (Fig. 3d and Supplementary Fig. 21). On the basis of the pairwise comparison between atalantia and pummelo, we concluded that a total of 18.1 Mb of genomic regions, containing 2,430 genes, were probably under selection in pummelo (Fig. 3d and Supplementary Table 28). These regions include genes associated with vegetative growth, such as Cg7g021010, Cg4g017400 and Cg9g010410, which are homologous to TCP15 (ref. 24), LAS25 and YUCCA3 (ref. 26), respectively (Supplementary Fig. 22). Notably, we observed a strong signal on chromosome 9, spanning 220 kb, from 13.8 Mb to 14.0 Mb (Fig. 3d and Supplementary Fig. 23) in the pummelo genome. Thirteen of the fifteen annotated genes in this region have functions related to cytochrome c biogenesis, mitochondrial proteins, ATP synthase, NADH dehydrogenase and mitochondrial ribosomal proteins (Supplementary Table 25). Comparative analysis also indicated that flowering-related genes, such as the gene encoding the WUSCHEL-related homeobox (WOX)27 transcription factor (Cg4g011660; Supplementary Fig. 24) and the gene encoding the flowering time control protein FPA (Cg2g027110; Supplementary Fig. 25), showed strongly reduced diversity and a high degree of differentiation. Therefore, we suggest that these genes were probably under selection. These regions were shown to harbor signals of selection as defined by recent studies22,28. The effect caused by other possibilities, such as genetic drift and bottlenecks29, could not be excluded but might be limited, as indicated by the similar pairwise sequentially Markovian coalescent (PSMC) curve of these three populations (Supplementary Fig. 26). FPA has a critical role in the regulation of flowering time in Arabidopsis30. This finding is consistent with the observation that atalantia has a prolonged flowering season (from May to December) and that in contrast, the flowering season is synchronized among all of the other cultivated citrus.
The genomic basis of citrus apomixis (nucellar polyembryony)
The emergence of a particular type of apomixis, nucellar polyembryony, in mandarin and most cultivated varieties of citrus is one of the prominent features of the recent evolution of citrus (Fig. 4a). We scored the polyembryony phenotypes of 124 fruit-yielding progeny from a segregating population derived from a cross between HB pummelo (monoembryony) and Fairchild mandarin (polyembryony) that exhibited contrasting phenotypes (Supplementary Table 29). For the bulk segregant analysis (BSA), the transformed Δ(SNP index) was calculated and plotted against the genomic sequence in 250-kb sliding windows with step sizes of 10 kb (Fig. 4b). The peak value for the transformed Δ(SNP index) was localized to a region spanning from 23.695 to 25.655 Mb on chromosome 4. These data indicated that this 1.96-Mb region harbors a major locus contributing to nucellar polyembryony.
(a) The phenotype of citrus polyembryony. (b) Mapping the polyembryony by bulk segregant analysis (BSA) of a segregating population derived from HB pummelo × Fairchild mandarin cross. Red arrow indicates the position of the 1.96-Mb peak. The transformed Δ(SNP index) is the product of the Δ(SNP index) and normalized SNP density in each 250-kb sliding window (10-kb steps). (c) Gene-based local association analysis in the relevant region. Top, heat map for association levels in the relevant region. The −log10-transformed P (two-tailed Fisher's exact test) was calculated in each 50-kb sliding window (10-kb steps). Middle, the 80-kb candidate region with the strongest association. Regression line (red) and confidence intervals (gray) were calculated using a linear model at a confidence level of 0.95. Bottom, genes in the candidate region in diploid pummelo. Boxes represent genes; horizontal black line represents intergenic regions; vertical lines show positions of 5′ end of the genes. Genes annotated from the haploid pummelo genome are presented in Supplementary Figure 28.
To fine map the polyembryony locus, a local association analysis with the variant information from the protein-encoding genes in the 1.96 Mb region was performed using the genome sequencing data from the 108 aforementioned accessions, including 45 polyembryonic and 63 monoembryonic accessions (Supplementary Table 23). We employed P values from the two-tailed Fisher's exact test to measure the correlations between nucleotide variation and phenotype (Supplementary Fig. 27), which were adjusted with a false discovery rate (FDR) correction for multiple testing. This analysis localized the polyembryony locus to an ∼80-kb region on chromosome 4, between 24.54 and 24.62 Mb, that contains 11 genes (Fig. 4c and Supplementary Fig. 28). Consistent with this result, the population differentiation between the polyembryonic and monoembryonic accessions was substantially higher for this 80-kb region than the surrounding regions (Supplementary Fig. 29). The linkage disequilibrium (LD) among polyembryonic accessions was larger than the LD among monoembryonic accessions (Supplementary Fig. 29), which indicates a selection signature between monoembryonic and polyembryonic citrus.
Among the 11 candidate genes, Cg4g018910 and Cg4g018970 showed the highest association with the polyembryony phenotype (Fig. 5a). Expression analysis of all candidate genes indicated that Cg4g018970 was highly expressed in ovules and specifically expressed in polyembryonic citrus cultivars but not in monoembryonic citrus cultivars (Fig. 5b,c and Supplementary Fig. 30). Cg4g018970 encodes a RWP-RK domain–containing protein similar to the Arabidopsis RKD family of proteins, which serve as regulators of egg cell–related genes31. Thus, we hereafter refer to Cg4g018970 as CitRWP. A sequence alignment of CitRWP from polyembryonic and monoembryonic citrus identified four SNPs in the gene sequence and an insertion of a miniature inverted-repeat TE (MITE) in the promoter region (g.24610316_24610317ins(203) in chromosome 4 in the reference of pummelo genome (MKYQ00000000)) (Fig. 5d and Supplementary Fig. 31). To test whether this transposon is linked to the polyembryony locus, we screened the 124 fruit-yielding individuals from the segregating population of 217 progeny derived from an HB pummelo × Fairchild mandarin cross. We did not find this MITE insertion in any of the monoembryonic progeny (n = 66). In contrast, we found the MITE insertion in all of the polyembryonic progeny (n = 58) (Supplementary Fig. 32). Thus, we conclude that this MITE insertion is associated with the polyembryony locus. Next, we used a germplasm collection containing 786 accessions of citrus to independently test this association. Consistent with our results, we found this MITE insertion in all of the polyembryonic accessions (n = 213). In contrast, all the monoembryonic accessions (n = 573) lack this particular MITE insertion (Fig. 5e,f and Supplementary Table 30).
(a) Beeswarm and box plot for associated SNPs in the 11 candidate genes. The dots in different colors represent associated SNPs in the corresponding gene sequence. Boxes, median and interquartile range; whiskers maximum and minimum values. (b) Relative expression of CitRWP as determined by qRT-PCR in different tissues of sweet orange. Data are mean ± s.d.; n = 3 technical replicates of 2 pooled tissue samples. (c) qRT-PCR validation of CitRWP expression in the ovules of monoembryonic cultivars (PML, pummelo; CTR, citron; CLM, clementine mandarin) and polyembryonic cultivars (GRP, grapefruit; LM, lemon; PK, ponkan). Data are mean ± s.d.; n = 3 technical replicates of 2 pooled tissue samples. (d) Schematic diagram of CitRWP. Asterisks indicate positions of SNPs associated with polyembryony; triangle indicates MITE insertion; orange rectangles indicate exons. (e) Representative PCR-based detection of MITE insertions in citrus cultivars. (f) PCR-based analysis of the MITE insertion in 786 citrus accessions and the F1 segregating population derived from the HB pummelo (HB) × Fairchild mandarin (FC) cross.
Transcriptome of sexual and nucellar embryos from citrus
To investigate the molecular processes and genes involved in sexual and nucellar embryogenesis in citrus, we compared the RNA-seq data derived from leaves, ovules, fruits and seeds of pummelo (sexual type) and sweet orange (asexual type). We found that 1,637 and 1,840 genes were highly expressed in ovules (≥2-fold higher than in other tissues) of pummelo and sweet orange, respectively (Fig. 6a and Supplementary Tables 31 and 32). We found that 853 of these genes were highly expressed in both pummelo and sweet orange. Gene Ontology (GO) analysis indicated that the genes associated with microtubule-based movement, DNA packaging, cell cycle, DNA replication, mitosis and transcription regulator activity were over-represented among the genes that were highly expressed in ovules (Supplementary Table 33). The genes that were highly expressed in the ovules of sweet orange were significantly (P < 0.05; FDR < 0.05) enriched in almost the same gene ontologies as the group of 853 genes that were highly expressed in the ovules of both pummelo and sweet orange, such as regulation of transcription, mitosis, nuclear division and mitotic cell cycle, which contribute to cell division (Supplementary Table 34). These data highlight similarities in the transcriptomes of nucellar and sexual embryos.
(a) Heat map of genes that are specifically expressed in ovules of both pummelo (monoembryony) and sweet orange (polyembryony). Leaf, ovule, fruit and seed tissues from pummelo (odd numbers) and sweet orange (even numbers) were used for the comparative analysis. (b) Heat map of the genes with significantly different expression in the ovules of monoembryonic (PML, pummelo; CTR, citron; CLM, clementine mandarin) and polyembryonic cultivars (GRP, grapefruit; LM, lemon; PK, ponkan). Data are derived from 2 replicates of plant tissue. Normalized FPKM values after scaling and centering are shown. (c) Model for genes that are involved in monoembryony and polyembryony, including the candidate gene CitRWP (yellow star). Blue dots indicate genes highly expressed in ovules and showed no significant difference in expression between monoembryonic and polyembryonic citrus cultivars (Supplementary Tables 36 and 37). Red dots indicate genes highly expressed in the ovules of polyembryonic citrus cultivars. M, megaspore mother cell; ES, embryo sac; N, nucellar embryo initiation cell.
To identify genes that are differentially expressed between polyembryony and monoembryony, a comparative transcriptome analysis was performed on the ovules from three pairs of monoembryonic and polyembryonic citrus cultivars (pummelo–grapefruit, citron–lemon, clementine–ponkan) at the key stage of nucellar embryo initiation (i.e., 3 d before anthesis)14,15. Considering the genetic differences between each pair of citrus cultivars, we performed a pairwise comparison of the three pairs of RNA-seq data. Among the 2,624 genes that were preferentially expressed in the ovules of pummelo and sweet orange, 29 showed markedly different (≥2-fold) expression levels between each pair of monoembryonic and polyembryonic cultivars (Fig. 6b and Supplementary Table 35). The annotation of these genes and analysis of homologous Arabidopsis sequences indicated that 11 of the 29 genes encode proteins with unknown function. Almost half of the remaining genes were associated with the biological processes of stress response (RNS1, ATSOD1, Cg4g011120 and Cg9g004560) and oxidation reduction (ATBBE18 and Cg3g004030). Three additional genes (EDA24, AGL2 and ACR4) contribute to flower and embryo development in Arabidopsis and other plants32,33,34. These genes are expressed in the ovules of both monoembryonic and polyembryonic cultivars and are expressed at dramatically higher levels in polyembryonic cultivars. CitRWP shows a similar expression trend in polyembryonic cultivars, which is consistent with our conclusion, from genetic analysis, that CitRWP is the key candidate gene controlling polyembryony (Fig. 6c and Supplementary Tables 36 and 37).
Discussion
We sequenced and assembled one high-quality genome and three draft genomes of Citrinae species. The haploid pummelo genome represents the most contiguous citrus genome assembly to date. The sequence contiguity (contig N50) of the long reads assembly using PacBio data is at least 18-fold higher than that of recently published citrus genomes18,19. The shotgun approach using PacBio single-molecule sequencing in combination with Illumina data is effective and efficient for a de novo genome assembly. This high-quality pummelo genome together with the three draft genomes of citron, papeda and atalantia constitute a valuable platform for citrus genetic and genomic research.
Change in the reproductive system is one of the striking features in the evolution and domestication of fruit crops35. Consistent with this observation, the sequence analysis of the six genomes that included primitive, wild and cultivated citrus indicated frequent variations in flowering- and seed-related genes, such as CKX21. The comparative population analysis of these species indicated that mitochondria- and reproduction-associated genes, such as WOX27 and FPA30, are probably under selection in cultivated citrus.
Apomixis has received considerable attention because of its capability to permanently fix valuable traits and hybrid genotypes (hybrid vigor). It is possible that apomixis was used inadvertently during the history of citrus breeding and selection. When desirable variations and traits were observed, the apomictic lines tended to have a greater chance of being selected, maintained and dispersed. This may also explain why most of the commercial and elite citrus are apomictic. In this study, we provided a comprehensive analysis of citrus apomixis (polyembryony) using genetic, genomic and transcriptomic approaches. The segregation ratios of both the phenotype and genotype in a segregating F1 population are consistent with the single dominant gene model for nucellar embryony9. The cosegregation of the MITE insertion in the promoter region of CitRWP with the polyembryonic phenotype lends more support to the single dominant mutation model for the emergence of apomixis in mandarin. Our genomic analysis localized the candidate region to an 80-kb interval containing 11 genes, which was embedded in a 380-kb region reported by a previous study12. In this interval, we identified a promising candidate gene for the single dominant allele responsible for polyembryony in citrus. Notably, this candidate gene, CitRWP, is specifically expressed in ovules and at higher levels in polyembryonic cultivars. A gene harboring the same domain, AtRKD4 (GROUNDED), was reported to promote embryogenesis in somatic cells when transiently overexpressed36,37. Fully elucidating the contribution of CitRWP to apomixis will require knocking out CitRWP in citrus.
Our transcriptome analyses provide global insights into the molecular processes involved in the development of sexual and asexual (nucellar) embryos in citrus. The results indicate a close developmental relationship between sexual reproduction and apomixis in citrus, which is consistent with a report on apomictic Boechera gunnisoniana38. The unusually high expression of three sexual embryogenesis–associated genes during apomixis (AGL2, EDA24 and ACR4) provides more support for the notion that nucellar embryogenesis and sexual reproduction may utilize similar genes or pathways. Previous studies reported that extreme stress induces the initiation of autonomous embryo development in somatic cells in response to exogenous or endogenous signals39. Here we identified six nucellar embryo-associated genes that are regulated by stress, which is in agreement with a previous report on apomixis in Hieracium praealtum40.
This genomic information on primitive, wild and cultivated citrus may also be valuable for the future analysis of metabolism and to further investigate the basis of historical uses of citrus species. For example, C. medica was historically used for medicinal purposes, and atalantia was also used as a traditional medicinal plant for folk medicines aiming to treat malaria, chronic rheumatism, paralysis and snakebites41. Additionally, wild and primitive citrus, which have been largely neglected during the history of citrus breeding, exhibit higher genetic diversity that is useful for managing disease in cultivated citrus with narrow genetic backgrounds resulting from apomixis, clonal propagation and human selection. Future characterization and utilization of genes controlling apomixis, metabolism, disease resistance and other agriculturally important traits is a promising path for the improvement of crop breeding.
Methods
Plant materials.
A total of 100 citrus accessions (Supplementary Table 23) were sequenced in this study. The germplasm containing 786 citrus accessions (Supplementary Table 30) was collected at the National Citrus Breeding Center, Huazhong Agricultural University, Wuhan, China, and the Citrus Experiment Stations from the MATS program of the Ministry of Agriculture of China. The haploid pummelo was obtained from the Citrus Research Institute, Southwest University, Chongqing, China (Supplementary Fig. 4).
The segregating population derived from the HB pummelo × Fairchild mandarin cross comprises 217 individuals including 124 fruit-bearing progeny (Supplementary Table 29). The population was created in 2004 and began flowering in 2012 at the National Citrus Breeding Center, Huazhong Agricultural University, Wuhan, China.
De novo assembly and annotation of four citrus genomes.
For the haploid C. grandis, 30 cells of PacBio data (56.8 × genome coverage) and 141 Gb of Illumina data (370.3 × genome coverage; Supplementary Tables 1, 2 and 38) containing both short and long inserts were used. For the assembly of haploid pummelo genome, the PacBio long reads were corrected and assembled using the Hierarchical Genome Assembly Process (HGAP)42 in SMRT Analysis (v2.3.0). Another round of polishing was performed with Quiver42 to further improve the quality of the assembled contigs. Next, mate-paired reads corrected by Quake43 were used in scaffolding with the SSPACE-STANDARD package44, and GapCloser20 was used to fill the gaps with all of the Illumina reads. Then, the SSPACE-STANDARD package was used to further extend the gap-filled scaffolds. To improve the completeness of the assembly, long contigs assembled only with Illumina reads were aligned to the PacBio-based assembly, and the unaligned sequences were added to the final assembly (Supplementary Note 3).
For the heterozygous diploid A. buxifolia, C. ichangensis and C. medica, short-insert (i.e., insert sizes ranging from 140 bp to 500 bp) and long-insert (i.e., insert sizes of 2, 5, and 20 kb) DNA libraries were constructed (Illumina) (Supplementary Tables 4–6 and 38). In total, 131.9 Gb (∼401.9× genome coverage), 64.3 Gb (∼164.4× genome coverage), and 69.5 Gb (∼170.9× genome coverage) of raw data were generated for A. buxifolia, C. ichangensis and C. medica, respectively. The draft genomes were assembled using SOAPdenovo20 with Illumina reads corrected using QUAKE. Then, all the reads were aligned onto the contigs for scaffolding, and the GapCloser package was used to fill the gaps. For the atalantia genome, the 3.73 Gb PacBio reads were corrected using the Illumina reads and were then used to improve the assembly. For the annotation of each of the four genomes, the TEs were masked and the annotation was performed ab initio by integrating the evidence from sequence homology and transcriptomic data (Supplementary Notes 4–6 and Supplementary Table 39).
Phylogenetic tree of Citrinae.
The coding region sequences from 103 single-copy genes18 were concatenated (Supplementary Data Set 1), constrained to their amino acid sequences and used for tree construction. The maximum likelihood (ML) tree was produced with Clausena as the outgroup using the substitution model GTRGAMMA of the RAxML software45. A total of 1,000 rapid bootstrap inferences were performed.
Detection of single nucleotide variations (SNVs) and presence–absence variations (PAVs).
The six citrus genomes (A. buxifolia, C. ichangensis, C. medica, C. grandis, C. clementina and C. sinensis) were used for comparison and synteny analyses. The differences between pairs of genomes were found using the Nucmer program in the MUMmer software46 with the default parameters. Alignment results of the five query genome sequences (A. buxifolia, C. ichangensis, C. medica, C. clementina and C. sinensis) against the reference genome of C. grandis were used to extract SNVs, PAVs and indels (insertions and deletions), which were further filtered and validated with re-sequencing data. In addition, the enrichment analysis for genes in the unique regions was performed using agriGO47 against the background of the corresponding whole genome (Supplementary Note 7).
Genomic characterization of six citrus genomes.
Protein-encoding genes from A. buxifolia, C. ichangensis, C. medica, C. grandis, C. clementina and C. sinensis were compared using BLASTP48 with an E-value cutoff of 1 × 10−10 and clustered using OrthoMCL49. The results of all-by-all BLASTP was used to estimate the synteny for pairs of citrus genomes with the i-ADHoRe software50. The gene family members in each genome were identified using the HMMER software51, on the basis of the domain profiles of 62 transcription factor families, 22 groups of transcriptional regulators collected in the PlnTFDB database (see URLs) and the NB-ARC domain (PF00931) from Pfam (see URLs). The domains of the proteins encoded by the members of the MADS-box (Supplementary Data Set 2) and NBS-related genes were manually checked using PROSITE (see URLs). The MADS-box and NBS-related genes were classified (Supplementary Figs. 33 and 34).
The dN/dS values were calculated for each of the 8,551 single-copy orthologous genes shared by the six citrus genomes using the codeml program in PAML52 with the one-ratio branch model and the free-ratios branch model. Using a likelihood ratio test, the values of dN/dS for each single-copy orthologous gene were determined and the fast-evolving single-copy orthologous genes were identified for each citrus genome (P < 0.05 and FDR < 0.05).
Genetic diversity and FST analysis.
A total of 100 citrus accessions were sequenced on the Illumina platform with an average depth of 30-fold genome coverage (Supplementary Table 23). In addition, we used sequence data from three mandarins, one orange and four pummelos from previous studies18,19 for our analysis. The paired-end data were aligned to the pummelo reference genome using BWA53. Population-based SNP calling was performed using SAMtools54. To evaluate the relationships among the 108 accessions, a neighbor-joining tree was constructed using the maximum composite likelihood method in MEGA5 software55 with 1,000 bootstrap test (data in Supplementary Data Set 3). The shotgun genome sequencing data of 15 atalantia, 11 papeda and 19 pummelo were used to represent primitive, wild and cultivated citrus. The SNPs located in the regions of single-copy orthologous genes shared by the six citrus genomes were used to construct a phylogenetic tree using the substitution model GTRGAMMA of the maximum likelihood method with the RAxML software (Supplementary Data Set 4).
For genetic diversity and selection signature analysis, SNPs of the whole genome were used, and a sliding-window approach (50-kb windows sliding in 10-kb steps) was employed to quantify genetic differentiation (FST) and nucleotide diversity (π) for each pair of citrus populations using the VCFtools software56. Values of π were also calculated in 2-Mb sliding windows with steps sizes of 1 Mb. To detect the selection signatures, the log10 π ratio was calculated. The regions with significantly high FST values (in the 5% right tail of the empirical distribution of FST values) and significant reduction in diversity (in the 5% right tail of the empirical distribution of log10 π ratio) were considered to be under selection (Supplementary Note 8).
Bulk segregant analysis (BSA) and pool sequencing.
Two DNA pools from 20 individuals exhibiting extreme monoembryonic and polyembryonic phenotypes were collected from a population derived from an HB pummelo × Fairchild mandarin cross (Supplementary Table 29). Sequencing was performed at tenfold and 60-fold depth for parents and DNA pools, respectively (Supplementary Table 40). Reads from the parents and from the monoembryonic and polyembryonic pools were separately mapped to the pummelo genome using BWA (mapping quality > 30). The SAMtools software was used for SNP calling (Q > 30).
According to the genotypes of HB pummelo (monoembryonic, pp) and Fairchild mandarin (polyembryonic, Pp), the allelic nucleotide polymorphisms fitting Pp in the polyembryonic pool and pp in the monoembryonic pool were identified. The sites were retained if they fit the following criteria: p% of the monoembryonic pool ≥ 0.9 and 0.4 ≤ p% ≤ 0.6 in the polyembryonic pool. The absolute value of their difference was calculated as a Δ(SNP index). Using the sliding-window approach, the average Δ(SNP index) and max–min normalized SNP density were calculated with 250-kb sliding windows with 10-kb step sizes. The final transformed Δ(SNP index) was the product of the average Δ(SNP index) and the normalized SNP density.
Local gene-based association analysis for the polyembryony locus.
The candidate polyembryony interval of 1.96 Mb was extracted from the genome sequences of atalantia, papeda, citron, sweet orange, and clementine mandarin by separately searching each genome with BLASTN. BWA was used to align the sequencing reads from the 108 accessions to the corresponding target sequences. The reads of each accession that mapped to the 1.96 Mb region were identified using the bam2fastq software (see URLs). These reads were remapped to gene sequences using the pummelo candidate regions as templates. After using iCORN2 (ref. 57) to substitute SNPs and small indels (1–3 bp), we obtained 292 gene sequences in the candidate region of each accession.
For each of the 292 genes in the candidate region, a multiple sequence alignment of homologous sequences from 108 accessions was performed using ClustalW58. For each polymorphic site, the P value from the two-tailed Fisher's exact test was calculated to quantify the associations between the nucleotide polymorphisms and the polyembryonic trait. The P values were adjusted using the FDR correction for multiple testing. The sites were retained if they fit the criteria of P < 0.01 and FDR < 0.01 (Supplementary Note 9). For the 11 candidate genes in the 80-kb region, all of the SNPs in the gene sequence from the 108 accessions were extracted and analyzed. For each SNP, the proportion of the accessions in which the genotype of the site was associated with the phenotype of the embryo was regarded as an association score. The SNPs with association scores greater than 80% were used for further SNP association analysis (Fig. 5a).
MITE marker development.
The 203-bp MITE insertion is located on chromosome 4 of the pummelo genome between nucleotides 24,610,316 and 24,610,317, which is in the promoter region of CitRWP. Two pairs of primers ('mite_p1' and 'mite_p2', Supplementary Table 41) were designed. Primer pair 'mite_p1' contains degenerate nucleotide to be used for the broad germplasm of 786 accessions. Primer pair 'mite_p2' was specifically designed to be used for the progeny from the HB pummelo × Fairchild mandarin cross. Both the primer pairs anneal approximately 100 bp upstream (forward primer) and 200 bp downstream (reversed primer) of the MITE insertion site (Supplementary Note 10). We performed a PCR-based screen to detect the MITE insertion. PCR was carried out in 20-μl reaction volumes containing 10 μl of 2× Taq Plus Mix (Vazyme Biotech), 100 ng genome template, 0.2 mM of each forward and reverse primer. The amplification was done at 95 °C for 5 min, followed by 30 cycles of 30 s at 95 °C, 30 s at 52 °C, and 60 s at 72 °C, and finally 5 min at 72 °C.
Transcriptome sequencing and analysis.
A total of 26 samples from leaves, ovules, fruits, and seeds were harvested for RNA-seq analysis. Two replicates of plant tissue were analyzed. RNA-seq libraries were constructed and sequenced on Illumina Genome Analyzer platform. An average of 4 Gb data were generated for each sample (Supplementary Table 42). The RNA-seq reads were aligned to the pummelo genome sequences using TopHat59. The expression level of each predicted transcript in each RNA-seq library was calculated as the FPKM with Cufflinks60. Gene Ontology (GO) enrichment analysis was performed using the web-based agriGO program with the default parameters (P < 0.05 and FDR < 0.05, Supplementary Tables 33, 34 and 43 and Supplementary Note 11).
Quantitative PCR analysis.
Total RNA from all the tissues was extracted using the TRIzol reagent (Takara). cDNA was synthesized using 1 μg total RNA and the HiScript II QRT SuperMix for qPCR (Vazyme, R223-01). qRT-PCR was performed on an ABI 7900 instrument (Applied Biosystems) using SYBR Green PCR Mastermix according to the manufacturer's instructions (Kapa, RR420). The cycling conditions included incubation for 3 min at 95 °C followed by 40 cycles of amplification (95 °C for 5 s and 60 °C for 35 s). Using the citrus β-actin gene as the internal reference gene, relative gene expression values were calculated using the 2−ΔΔCt method61. The primers used in qRT-PCR are listed in Supplementary Table 41.
Statistical analyses.
We used the Fisher's exact test to conduct the GO enrichment analysis of the target genes relative to the background of the corresponding entire genome using agriGO program. The likelihood ratio test was used to determine the branch model during the dN/dS analysis using PAML program. The two-tailed Fisher's exact tests were used to quantify the associations between the nucleotide polymorphisms and the polyembryonic trait. Moreover, all the P values from the tests mentioned above were adjusted using the FDR correction for multiple testing. The filtering criteria of P < 0.05 and FDR < 0.05 were given for the analysis of GO enrichment analysis and likelihood ratio test, and the filtering criteria were P < 0.01 and FDR < 0.01 for the associations analysis of citrus polyembryony.
Data availability.
Genome assembly data for Citrus grandis, C. ichangensis, C. medica and A. buxifolia have been deposited at DDBJ/ENA/GenBank under accession numbers MKYQ00000000, MKYP00000000, MKYO00000000 and MKYR00000000, respectively. The versions described in this paper are MKYQ01000000, MKYP01000000, MKYO01000000 and MKYR01000000, respectively. Whole-genome sequencing data for C. grandis, C. ichangensis, C. medica, A. buxifolia, C. reticulata and C. sinensis have been deposited in BioProject under accession numbers PRJNA318855, PRJNA321657, PRJNA320023, PRJNA327148, PRJNA320985 and PRJNA321100, respectively. The RNA-seq data for C. grandis, C. ichangensis, C. medica, A. buxifolia, C. reticulata and C. sinensis have been deposited in BioProject under accession codes PRJNA339650, PRJNA340932, PRJNA341284, PRJNA341533, PRJNA341756 and PRJNA340305. The data for genome assembly and annotation of C. grandis, C. ichangensis, C. medica and A. buxifolia are available through our website at http://citrus.hzau.edu.cn/orange/download/index.php.
URLs.
PlnTFDB database, http://plntfdb.bio.uni-potsdam.de/v3.0/; Pfam, http://pfam.xfam.org/browse; PROSITE, http://prosite.expasy.org/prosite.html; bam2fastq, https://gsl.hudsonalpha.org/information/software/bam2fastq.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Accession codes
Primary accessions
BioProject
NCBI Reference Sequence
References
Meyer, R.S., DuVal, A.E. & Jensen, H.R. Patterns and processes in crop domestication: an historical review and quantitative analysis of 203 global food crops. New Phytol. 196, 29–48 (2012).
Conner, J.A., Mookkan, M., Huo, H., Chae, K. & Ozias-Akins, P. A parthenogenesis gene of apomict origin elicits embryo formation from unfertilized eggs in a sexual plant. Proc. Natl. Acad. Sci. USA 112, 11205–11210 (2015).
Koltunow, A.M. Apomixis: embryo sacs and embryos formed without meiosis or fertilization in ovules. Plant Cell 5, 1425–1437 (1993).
Batygina, T.B. & Vinogradova, G.Y. Phenomenon of polyembryony. Genetic heterogeneity of seeds. Russ. J. Dev. Biol. 38, 126–151 (2007).
Spillane, C., Curtis, M.D. & Grossniklaus, U. Apomixis technology development—virgin births in farmers' fields? Nat. Biotechnol. 22, 687–691 (2004).
Bicknell, R.A. & Koltunow, A.M. Understanding apomixis: recent advances and remaining conundrums. Plant Cell 16 (Suppl. 1), S228–S245 (2004).
Kepiro, J.L. & Roose, M.L. AFLP markers closely linked to a major gene essential for nucellar embryony (apomixis) in Citrus maxima × Poncirus trifoliata. Tree Genet. Genomes 6, 1–11 (2010).
Ueno, I., Iwamasa, M. & Nishiura, M. Embryo number of various varieties of Citrus and its relatives. Bull. Hort. Res. Sta. Japan 7, 11–22 (1967).
Iwamasa, M., Ueno, I. & Nishiura, M. Inheritance of nucellar embryony in citrus. Bull. Hort. Res. Sta. Japan 7, 1–8 (1967).
Cameron, J.W. & Soost, R.K. Sexual and nucellar embryony in F1 hybrids and advanced crosses of Citrus with Poncirus. J. Am. Soc. Hortic. Sci. 104, 408–410 (1979).
Raga, V., Bernet, G.P., Carbonell, E.A. & Asins, M.J. Segregation and linkage analyses in two complex populations derived from the citrus rootstock Cleopatra mandarin. Inheritance of seed reproductive traits. Tree Genet. Genomes 8, 1061–1071 (2012).
Nakano, M. et al. Characterization of genomic sequence showing strong association with polyembryony among diverse Citrus species and cultivars, and its synteny with Vitis and Populus. Plant Sci. 183, 131–142 (2012).
Nakano, M. et al. Characterization of genes associated with polyembryony and in vitro somatic embryogenesis in Citrus. Tree Genet. Genomes 9, 795–803 (2013).
Kumar, V., Malik, S.K., Pal, D., Srinivasan, R. & Bhat, S.R. Comparative transcriptome analysis of ovules reveals stress related genes associated with nucellar polyembryony in citrus. Tree Genet. Genomes 10, 449–464 (2014).
Long, J.M. et al. Genome-scale mRNA and small RNA transcriptomic insights into initiation of citrus apomixis. J. Exp. Bot. 67, 5743–5756 (2016).
Swingle, W.T. & Reece, P.C. The botany of Citrus and its wild relatives. in The Citrus Industry (University of California Press, 1967).
Medina-Filho, H.P., Bordignon, R. & Ballvé, R.M.L. Sunkifolias and Buxisunkis: sexually obtained reciprocal hybrids of Citrus sunki × Severinia buxifolia. Genet. Mol. Biol. 21, 129–133 (1998).
Xu, Q. et al. The draft genome of sweet orange (Citrus sinensis). Nat. Genet. 45, 59–66 (2013).
Wu, G.A. et al. Sequencing of diverse mandarin, pummelo and orange genomes reveals complex history of admixture during citrus domestication. Nat. Biotechnol. 32, 656–662 (2014).
Luo, R. et al. SOAPdenovo2: an empirically improved memory-efficient short-read de novo assembler. Gigascience 1, 18 (2012).
Ashikari, M. et al. Cytokinin oxidase regulates rice grain production. Science 309, 741–745 (2005).
Schmutz, J. et al. A reference genome for common bean and genome-wide analysis of dual domestications. Nat. Genet. 46, 707–713 (2014).
Qi, J. et al. A genomic variation map provides insights into the genetic basis of cucumber domestication and diversity. Nat. Genet. 45, 1510–1515 (2013).
Kieffer, M., Master, V., Waites, R. & Davies, B. TCP14 and TCP15 affect internode length and leaf shape in Arabidopsis. Plant J. 68, 147–158 (2011).
Greb, T. et al. Molecular analysis of the LATERAL SUPPRESSOR gene in Arabidopsis reveals a conserved control mechanism for axillary meristem formation. Genes Dev. 17, 1175–1187 (2003).
Zhao, Y. et al. A role for flavin monooxygenase–like enzymes in auxin biosynthesis. Science 291, 306–309 (2001).
van der Graaff, E., Laux, T. & Rensing, S.A. The WUS homeobox-containing (WOX) protein family. Genome Biol. 10, 248 (2009).
Lin, T. et al. Genomic analyses provide insights into the history of tomato breeding. Nat. Genet. 46, 1220–1226 (2014).
Wolf, J.B.W. & Ellegren, H. Making sense of genomic islands of differentiation in light of speciation. Nat. Rev. Genet. 18, 87–100 (2017).
Schomburg, F.M., Patton, D.A., Meinke, D.W. & Amasino, R.M. FPA, a gene involved in floral induction in Arabidopsis, encodes a protein containing RNA-recognition motifs. Plant Cell 13, 1427–1436 (2001).
Köszegi, D. et al. Members of the RKD transcription factor family induce an egg cell-like gene expression program. Plant J. 67, 280–291 (2011).
Pagnussat, G.C. et al. Genetic and molecular identification of genes required for female gametophyte development and function in Arabidopsis. Development 132, 603–614 (2005).
Dreni, L. & Zhang, D. Flower development: the evolutionary history and functions of the AGL6 subfamily MADS-box genes. J. Exp. Bot. 67, 1625–1638 (2016).
Gifford, M.L., Dean, S. & Ingram, G.C. The Arabidopsis ACR4 gene plays a role in cell layer organisation during ovule integument and sepal margin development. Development 130, 4249–4258 (2003).
Miller, A.J. & Gross, B.L. From forest to field: perennial fruit crop domestication. Am. J. Bot. 98, 1389–1414 (2011).
Jeong, S., Palmer, T.M. & Lukowitz, W. The RWP-RK factor GROUNDED promotes embryonic polarity by facilitating YODA MAP kinase signaling. Curr. Biol. 21, 1268–1276 (2011).
Waki, T., Hiki, T., Watanabe, R., Hashimoto, T. & Nakajima, K. The Arabidopsis RWP-RK protein RKD4 triggers gene expression and pattern formation in early embryogenesis. Curr. Biol. 21, 1277–1281 (2011).
Schmidt, A. et al. Apomictic and sexual germline development differ with respect to cell cycle, transcriptional, hormonal and epigenetic regulation. PLoS Genet. 10, e1004476 (2014).
Pasternak, T.P. et al. The role of auxin, pH, and stress in the activation of embryogenic cell division in leaf protoplast-derived cells of alfalfa. Plant Physiol. 129, 1807–1819 (2002).
Okada, T. et al. Enlarging cells initiating apomixis in Hieracium praealtum transition to an embryo sac program prior to entering mitosis. Plant Physiol. 163, 216–231 (2013).
Shi, M., Guo, X., Chen, Y., Zhou, L. & Zhang, D. Isolation and characterization of 19 polymorphic microsatellite loci for Atalantia buxifolia (Rutaceae), a traditional medicinal plant. Conserv. Genet. Resour. 6, 857–859 (2014).
Chin, C.S. et al. Nonhybrid, finished microbial genome assemblies from long-read SMRT sequencing data. Nat. Methods 10, 563–569 (2013).
Kelley, D.R., Schatz, M.C. & Salzberg, S.L. Quake: quality-aware detection and correction of sequencing errors. Genome Biol. 11, R116 (2010).
Boetzer, M., Henkel, C.V., Jansen, H.J., Butler, D. & Pirovano, W. Scaffolding pre-assembled contigs using SSPACE. Bioinformatics 27, 578–579 (2011).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
Kurtz, S. et al. Versatile and open software for comparing large genomes. Genome Biol. 5, R12 (2004).
Du, Z., Zhou, X., Ling, Y., Zhang, Z. & Su, Z. agriGO: a GO analysis toolkit for the agricultural community. Nucleic Acids Res. 38, W64–W70 (2010).
Altschul, S.F., Gish, W., Miller, W., Myers, E.W. & Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Li, L., Stoeckert, C.J. Jr. & Roos, D.S. OrthoMCL: identification of ortholog groups for eukaryotic genomes. Genome Res. 13, 2178–2189 (2003).
Simillion, C., Janssens, K., Sterck, L. & Van de Peer, Y. i-ADHoRe 2.0: an improved tool to detect degenerated genomic homology using genomic profiles. Bioinformatics 24, 127–128 (2008).
Finn, R.D., Clements, J. & Eddy, S.R. HMMER web server: interactive sequence similarity searching. Nucleic Acids Res. 39, W29–W37 (2011).
Yang, Z. PAML 4: phylogenetic analysis by maximum likelihood. Mol. Biol. Evol. 24, 1586–1591 (2007).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Tamura, K. et al. MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 28, 2731–2739 (2011).
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2158 (2011).
Otto, T.D., Sanders, M., Berriman, M. & Newbold, C. Iterative Correction of Reference Nucleotides (iCORN) using second generation sequencing technology. Bioinformatics 26, 1704–1707 (2010).
Larkin, M.A. et al. Clustal W and Clustal X version 2.0. Bioinformatics 23, 2947–2948 (2007).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Trapnell, C. et al. Transcript assembly and quantification by RNA-seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 28, 511–515 (2010).
Livak, K.J. & Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25, 402–408 (2001).
Acknowledgements
We are grateful to the staff from the Citrus Experiment Stations of the MATS program of the Ministry of Agriculture of China (CARS-27) for providing the citrus samples, D.-X. Zhang and M.-M. Shi (South China Botanical Garden of Chinese Academy of Sciences) for the Atalantia samples, R. Krueger (USDA-ARS National Clonal Germplasm Repository for Citrus and Dates) and T. Siebert (University of California, Riverside) for citrus samples. We thank H. Kuang, W. Xie, Y. Ouyang and L. Chai for helpful discussions, and S.-N. Hu, R.-Y. Jia and the sequencing platform from Beijing Institute of Genomics of Chinese Academy of Sciences (CAS) for the PacBio sequencing. This project was supported by the National Natural Science Foundation of China (31630065, 31572105, 31330066, 31272148 and 31521092 to X.D. and Q.X.), the fundamental research funds for the central universities (2662015PY109 to Q.X.) and the National Basic Research Program of China (973 project 2013CB127103 to B.-G.M.).
Author information
Authors and Affiliations
Contributions
Q.X. conceived and designed the project and the strategy. X.W. performed comparative and population genomics and genetic mapping analyses and annotated three draft genomes. Y.X. performed transcriptome and genetic analyses. S.Z analyzed candidate genes and molecular markers in the populations. L.C. provided the haploid pummelo samples. Y.H. assembled the haploid genome and managed the database. J.C. and S.T. assembled the draft genomes. G.W. performed the genome annotations and anchor. Y.L. annotated one draft genome. C.C. and H.L. performed cytological analyses. H.Y., L.W., X.J., X.Y. and J.X. collected and evaluated the samples for genome and transcriptome analyses. Q.X. coordinated the project with help from X.D., B.-G.M., L.-L.C. and R.M.L. Z.X. and M.T. generated and maintained the segregating population for this study. X.W., Y.X. and Q.X. wrote the manuscript with contributions from R.M.L., S.Z., Y.H., J.C., G.W., S.T., C.C., Y.L., N.W., L.-L.C., B.-G.M., Y.R. and X.D.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–34, Supplementary Tables 1–7, 9–12, 14, 15, 24, 28, 30, 39, 41 and Supplementary Notes 1–11 (PDF 20959 kb)
Supplementary Data Set 1
The concatenated sequences containing 103 single copy genes for phylogenetic analysis. (TXT 1238 kb)
Supplementary Data Set 2
The protein sequences of the MADS-box transcription factor of six citrus species and Arabidopsis. (TXT 145 kb)
Supplementary Data Set 3
The SNP data set used for tree construction for 108 citrus accessions. (TXT 95119 kb)
Supplementary Data Set 4
The SNP data set used for tree construction for atalantia, papeda and pummelo populations. (TXT 38884 kb)
Supplementary Tables
Supplementary Tables 8, 13, 16–23, 25–27, 29, 31–38, 40, 42 and 43 (XLSX 16788 kb)
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International (CC BY 4.0) License. The images or other third party material in this article is included in the article’s Creative Commons license, unless otherwise indicated in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Wang, X., Xu, Y., Zhang, S. et al. Genomic analyses of primitive, wild and cultivated citrus provide insights into asexual reproduction. Nat Genet 49, 765–772 (2017). https://doi.org/10.1038/ng.3839
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3839
This article is cited by
-
QTL analysis on a lemon population provides novel insights on the genetic regulation of the tolerance to the two-spotted spider mite attack
BMC Plant Biology (2024)
-
An integrated multi-omics approach reveals polymethoxylated flavonoid biosynthesis in Citrus reticulata cv. Chachiensis
Nature Communications (2024)
-
Haplotype-resolved chromosome-level genome assembly of Huyou (Citrus changshanensis)
Scientific Data (2024)
-
A tetraploid-dominated cytochimera developed from a natural bud mutant of the nonapomictic mandarin variety ‘Orah’
Molecular Breeding (2024)
-
Workflow for development of CAPS markers with one type of restriction enzyme to identify citrus cultivars
Tree Genetics & Genomes (2024)