Abstract
Crop domestication provided the calories that fueled the rise of civilization1,2,3. For many crop species, domestication was accompanied by the evolution of weedy crop relatives, which aggressively outcompete crops and reduce harvests4,5,6. Understanding the genetic mechanisms that underlie the evolution of weedy crop relatives is critical for agricultural weed management and food security. Here we use whole-genome sequences to examine the origin and adaptation of the two major strains of weedy rice found in the United States. We find that de-domestication from cultivated ancestors has had a major role in their evolution, with relatively few genetic changes required for the emergence of weediness traits. Weed strains likely evolved both early and late in the history of rice cultivation and represent an under-recognized component of the domestication process. Genomic regions identified here that show evidence of selection can be considered candidates for future genetic and functional analyses for rice improvement.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Doebley, J.F., Gaut, B.S. & Smith, B.D. The molecular genetics of crop domestication. Cell 127, 1309–1321 (2006).
Ross-Ibarra, J., Morrell, P.L. & Gaut, B.S. Plant domestication, a unique opportunity to identify the genetic basis of adaptation. Proc. Natl. Acad. Sci. USA 104, 8641–8648 (2007).
Larson, G. et al. Current perspectives and the future of domestication studies. Proc. Natl. Acad. Sci. USA 111, 6139–6146 (2014).
Vigueira, C., Olsen, K. & Caicedo, A. The red queen in the corn: agricultural weeds as models of rapid adaptive evolution. Heredity 110, 303–311 (2013).
De Wet, J.M. & Harlan, J.R. Weeds and domesticates: evolution in the man-made habitat. Econ. Bot. 29, 99–108 (1975).
Ellstrand, N.C. et al. Crops gone wild: evolution of weeds and invasives from domesticated ancestors. Evol. Appl. 3, 494–504 (2010).
Oerke, E.C. Crop losses to pests. J. Agric. Sci. 144, 31–43 (2006).
Estorninos, L.E. Jr., Gealy, D.R., Gbur, E.E., Talbert, R.E. & McClelland, M.R. Rice and red rice interference. II. Rice response to population densities of three red rice (Oryza sativa) ecotypes. Weed Sci. 53, 683–689 (2005).
Gealy, D.R. & Yan, W. Weed suppression potential of 'Rondo' and other indica rice germplasm lines. Weed Technol. 26, 517–524 (2012).
Chauhan, B.S. Strategies to manage weedy rice in Asia. Crop Prot. 48, 51–56 (2013).
Hill, J., Smith, R.J. & Bayer, D. Rice weed control: current technology and emerging issues in temperate rice. Anim. Prod. Sci. 34, 1021–1029 (1994).
Ziska, L.H. et al. Chapter three—weedy (red) rice: an emerging constraint to global rice production. Adv. Agron. 129, 181–228 (2015).
Basu, C., Halfhill, M.D., Mueller, T.C. & Stewart, C.N. Weed genomics: new tools to understand weed biology. Trends Plant Sci. 9, 391–398 (2004).
Burgos, N.R., Norman, R.J., Gealy, D.R. & Black, H. Competitive N uptake between rice and weedy rice. Field Crops Res. 99, 96–105 (2006).
Gealy, D.R. in Crop Ferality and Volunteerism (ed. Gressel, J.) 323–354 (CRRC Press, 2005).
Shivrain, V.K. et al. Gene flow between Clearfield™ rice and red rice. Crop Prot. 26, 349–356 (2007).
Shivrain, V.K., Burgos, N.R., Gealy, D.R., Moldenhauer, K.A. & Baquireza, C.J. Maximum outcrossing rate and genetic compatibility between red rice (Oryza sativa) biotypes and Clearfield™ rice. Weed Sci. 56, 807–813 (2008).
Burgos, N.R. et al. The impact of herbicide-resistant rice technology on phenotypic diversity and population structure of United States weedy rice. Plant Physiol. 166, 1208–1220 (2014).
Lu, B.R., Yang, X. & Ellstrand, N.C. Fitness correlates of crop transgene flow into weedy populations: a case study of weedy rice in China and other examples. Evol. Appl. 9, 857–870 (2016).
Merotto, A. et al. Evolutionary and social consequences of introgression of nontransgenic herbicide resistance from rice to weedy rice in Brazil. Evol. Appl. 9, 837–846 (2016).
Song, B.K., Chuah, T.S., Tam, S.M. & Olsen, K.M. Malaysian weedy rice shows its true stripes: wild Oryza and elite rice cultivars shape agricultural weed evolution in Southeast Asia. Mol. Ecol. 23, 5003–5017 (2014).
Pusadee, T., Schaal, B.A., Rerkasem, B. & Jamjod, S. Population structure of the primary gene pool of Oryza sativa in Thailand. Genet. Resour. Crop Evol. 60, 335–353 (2013).
Londo, J. & Schaal, B. Origins and population genetics of weedy red rice in the USA. Mol. Ecol. 16, 4523–4535 (2007).
Reagon, M. et al. Genomic patterns of nucleotide diversity in divergent populations of US weedy rice. BMC Evol. Biol. 10, 180 (2010).
Grimm, A., Fogliatto, S., Nick, P., Ferrero, A. & Vidotto, F. Microsatellite markers reveal multiple origins for Italian weedy rice. Ecol. Evol. 3, 4786–4798 (2013).
Thurber, C.S., Jia, M.H., Jia, Y. & Caicedo, A.L. Similar traits, different genes? Examining convergent evolution in related weedy rice populations. Mol. Ecol. 22, 685–698 (2013).
Vigueira, C., Li, W. & Olsen, K. The role of Bh4 in parallel evolution of hull colour in domesticated and weedy rice. J. Evol. Biol. 26, 1738–1749 (2013).
Qi, X. et al. More than one way to evolve a weed: parallel evolution of US weedy rice through independent genetic mechanisms. Mol. Ecol. 24, 3329–3344 (2015).
Qiu, J. et al. Genome re-sequencing suggested a weedy rice origin from domesticated indica–japonica hybridization: a case study from southern China. Planta 240, 1353–1363 (2014).
Kumar, S., Stecher, G. & Tamura, K. MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 33, 1870–1874 (2016).
Alexander, D.H., Novembre, J. & Lange, K. Fast model-based estimation of ancestry in unrelated individuals. Genome Res. 19, 1655–1664 (2009).
Drummond, A.J., Suchard, M.A., Xie, D. & Rambaut, A. Bayesian phylogenetics with BEAUti and the BEAST 1.7. Mol. Biol. Evol. 29, 1969–1973 (2012).
Meyer, R.S. & Purugganan, M.D. Evolution of crop species: genetics of domestication and diversification. Nat. Rev. Genet. 14, 840–852 (2013).
Olsen, K.M. & Wendel, J.F. A bountiful harvest: genomic insights into crop domestication phenotypes. Annu. Rev. Plant Biol. 64, 47–70 (2013).
Olsen, K.M. & Wendel, J.F. Crop plants as models for understanding plant adaptation and diversification. Front. Plant Sci. 4, 1–16 (2013).
Cui, Y. et al. Little white lies: pericarp color provides insights into the origins and evolution of southeast Asian weedy rice. G3 6, 4105–4114 (2016).
Gu, X.Y. et al. Association between seed dormancy and pericarp color is controlled by a pleiotropic gene that regulates abscisic acid and flavonoid synthesis in weedy red rice. Genetics 189, 1515–1524 (2011).
Reagon, M., Thurber, C.S., Olsen, K.M., Jia, Y. & Caicedo, A.L. The long and the short of it: SD1 polymorphism and the evolution of growth trait divergence in US weedy rice. Mol. Ecol. 20, 3743–3756 (2011).
Pavlidis, P., Živković, D., Stamatakis, A. & Alachiotis, N. SweeD: likelihood-based detection of selective sweeps in thousands of genomes. Mol. Biol. Evol. 30, 2224–2234 (2013).
Huang, X. et al. A map of rice genome variation reveals the origin of cultivated rice. Nature 490, 497–501 (2012).
The 3,000 Rice Genomes Project Group. The 3,000 Rice Genomes Project. Gigascience 3, 7 (2014).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
McKenna, A. et al. The genome analysis toolkit: a mapreduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Xu, X. et al. Resequencing 50 accessions of cultivated and wild rice yields markers for identifying agronomically important genes. Nat. Biotechnol. 30, 105–111 (2012).
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2158 (2011).
Watterson, G.A. On the number of segregating sites in genetical models without recombination. Theor. Popul. Biol. 7, 256–276 (1975).
Larkin, M.A. et al. Clustal W and Clustal X version 2.0. Bioinformatics 23, 2947–2948 (2007).
Nielsen, R. et al. Darwinian and demographic forces affecting human protein coding genes. Genome Res. 19, 838–849 (2009).
Acknowledgements
We thank M. Dyer and staff of the University greenhouse for assistance in growing plants and L. Small for technical laboratory assistance. We are grateful to members of the Olsen laboratory for helpful comments on this manuscript. This work is supported by the National Science Foundation Plant Genome Research Program (IOS-1032023). The USDA is an equal opportunity provider and employer.
Author information
Authors and Affiliations
Contributions
K.M.O., Y.J. and A.L.C. designed the experiments. L.-F.L. and Y.-L.L. analyzed the data. L.-F.L., K.M.O. and A.L.C. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
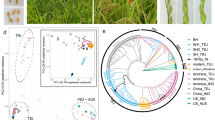
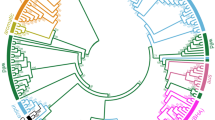
Supplementary Figure 1 Neighbor-joining tree of the 183 wild, cultivated, and weedy rice accessions.
Relationships of cultivated and wild rice correspond to previously observed relationships40. Wild rice accessions (dark green) are divided into different groups. The japonica (orange) and aromatic (light green) rice varieties form a clade. The BHA (red), SH (purple), and Chinese (black) weedy rice strains cluster with indica (light blue) and aus (pink), respectively. Bootstrap values (>50) are shown on each branch.
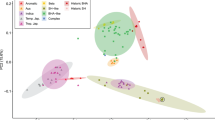
Supplementary Figure 2 Ancestral population structures of 183 rice accessions.
(a,b) Analyses of the full data set (a) and no-missing data set (b) were performed separately. Each vertical bar represents one accession, and different colors indicate distinct ancestry states. Cross-validation error was estimated for diverse K values from two to five. K = 3 minimizes the cross-validation error.
Supplementary Figure 3 Distributions of wild- and crop-specific private SNPs in SH (orange), BHA (red), and Chinese (green) weedy rice at the whole-genome level.
The box corresponds to the 95% confidence interval, and the black bar within each box is the mean value. The blue horizontal dashed line represents the equal proportion of wild- and crop-specific private SNPs. Positive and negative values represent crop-like and wild-like, respectively.
Supplementary Figure 4 Distribution patterns of wild- and crop-specific private SNPs within each chromosome of SH (orange), BHA (red), and Chinese (green) weedy rice.
The box corresponds to the 95% confidence interval, and the black bar within each box is the mean value for each chromosome. The blue horizontal dotted line represents zero. Positive and negative values represent the presence of crop-like and wild-like SNPs, respectively.
Supplementary Figure 5 Decrease in nucleotide diversity (θ) in SH and BHA weedy rice as compared to their crop ancestors.
The y axis represents the ratio of nucleotide diversity (θ) between weedy rice and their crop ancestors. Each bar is a 100-kb window, and the red solid line corresponds to the threshold for the top 5%. The numbers on the red line are the exact thresholds for each comparison.
Supplementary Figure 8 Ratio of nucleotide diversity (π) between indica and SH weedy rice across the genome.
(a) The criteria MQ ≥ 10 and DP ≥ 1 were used to filter the reported variants. (b) The criteria MQ ≥ 20 and DP ≥ 3 were used to filter the reported variants. (c) The criteria MQ ≥ 10 and DP ≥ 1 were used, excluding low-frequency (<2%) heterozygosity. Each bar represents a 100-kb window.
Supplementary Figure 9 Genome-wide genetic differentiation (FST) between indica and SH weedy rice across the genome.
(a) The criteria MQ ≥ 10 and DP ≥ 1 were used to filter the reported variants. (b) The criteria MQ ≥ 20 and DP ≥ 3 were used to filter the reported variants. (c) The criteria MQ ≥ 10 and DP ≥ 1 were used, excluding low-frequency (<2%) heterozygosity. Each bar represents a 100-kb window.
Supplementary Figure 10 Ratio of nucleotide diversity (π) between BHA and aus across the genome.
(a) The criteria MQ ≥ 10 and DP ≥ 1 were used to filter the reported variants. (b) The criteria MQ ≥ 20 and DP ≥ 3 were used to filter the reported variants. (c) The criteria MQ ≥ 10 and DP ≥ 1 were used, excluding low-frequency (<2%) heterozygosity. Each bar represents a 100-kb window.
Supplementary Figure 11 Genome-wide genetic differentiation (FST) between BHA and aus across the genome.
(a) The criteria MQ ≥ 10 and DP ≥ 1 were used to filter the reported variants. (b) The criteria MQ ≥ 20 and DP ≥ 3 were used to filter the reported variants. (c) The criteria MQ ≥ 10 and DP ≥ 1 were used, excluding low-frequency (<2%) heterozygosity. Each bar represents a 100-kb window.
Supplementary Figure 12 Ratio of nucleotide diversity (π) between wild and cultivated rice across the genome.
(a) The criteria MQ ≥ 10 and DP ≥ 1 were used to filter the reported variants. (b) The criteria MQ ≥ 20 and DP ≥ 3 were used to filter the reported variants. (c) The criteria MQ ≥ 10 and DP ≥ 1 were used, excluding low-frequency (<2%) heterozygosity. Each bar represents a 100-kb window.
Supplementary Figure 13 Genome-wide genetic differentiation (FST) between wild and cultivated rice across the genome.
(a) The criteria MQ ≥ 10 and DP ≥ 1 were used to filter the reported variants. (b) The criteria MQ ≥ 20 and DP ≥ 3 were used to filter the reported variants. (c) The criteria MQ ≥ 10 and DP ≥ 1 were used, excluding low-frequency (<2%) heterozygosity. Each bar represents a 100-kb window.
Supplementary Figure 14 Ratio of nucleotide diversity (π) between wild and cultivated rice.
The identified domestication genes are shown for each chromosome.
Supplementary Figure 15 Ratio of nucleotide diversities (θ and π) between wild, cultivated, and weedy rice.
The y and x axes are the ratio of θ and π, respectively.
Supplementary Figure 16 Neighbor-joining tree based on the 54 selected rice accessions.
Relationships of cultivated and wild rice correspond to previously observed relationships40. Wild rice accessions (dark green) are divided into different groups. The japonica (orange) and aromatic (light green) rice varieties form a clade. The BHA (red), SH (purple), and Chinese (black) weedy rice strains cluster with indica (light blue) and aus (pink), respectively. Bootstrap values (>50) are shown on each branch.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–16 and Supplementary Tables 2, 4–6 and 14 (PDF 2846 kb)
Supplementary Table 1
The list of 183 accessions of wild, cultivated, and weedy rice species sampled in the collection. (XLSX 59 kb)
Supplementary Table 3
Genetic assignments at wild, cultivated, and weedy rice accessions from two to five. (XLSX 59 kb)
Supplementary Table 7
Causative mutations of domestication genes in the three types of weedy rice. (XLSX 37 kb)
Supplementary Table 8
Genome-wide detection of regions of low nucleotide diversity in SH compared to indica. (XLSX 52 kb)
Supplementary Table 9
Genome-wide detection of regions of low nucleotide diversity in BHA compared to aus. (XLSX 53 kb)
Supplementary Table 10
Functionally characterized genes showing low nucleotide diversity in SH and high genetic differentiation between SH and indica. (XLSX 57 kb)
Supplementary Table 11
Functionally characterized genes showing low nucleotide diversity in BHA and high genetic differentiation between BHA and aus. (XLSX 70 kb)
Supplementary Table 12
Number of raw variants in wild, cultivated, and weedy rice with different filter criteria. (XLSX 42 kb)
Supplementary Table 13
Genome-wide detection and functional annotation of selective sweep regions in the full crop populations. (XLSX 54 kb)
Supplementary Data
Sequence alignment of Rc genes in wild, cultivated, and weedy rice. (TXT 2000 kb)
Rights and permissions
About this article
Cite this article
Li, LF., Li, YL., Jia, Y. et al. Signatures of adaptation in the weedy rice genome. Nat Genet 49, 811–814 (2017). https://doi.org/10.1038/ng.3825
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3825
This article is cited by
-
Porous borders at the wild-crop interface promote weed adaptation in Southeast Asia
Nature Communications (2024)
-
Transcriptomic analysis provides insight into the genetic regulation of shade avoidance in Aegilops tauschii
BMC Plant Biology (2023)
-
A syntelog-based pan-genome provides insights into rice domestication and de-domestication
Genome Biology (2023)
-
Common evolutionary trajectory of short life-cycle in Brassicaceae ruderal weeds
Nature Communications (2023)
-
Weed genomics: yielding insights into the genetics of weedy traits for crop improvement
aBIOTECH (2023)