Abstract
Landraces (traditional varieties) of domesticated species preserve useful genetic variation, yet they remain untapped due to the genetic linkage between the few useful alleles and hundreds of undesirable alleles1. We integrated two approaches to characterize the diversity of 4,471 maize landraces. First, we mapped genomic regions controlling latitudinal and altitudinal adaptation and identified 1,498 genes. Second, we used F-one association mapping (FOAM) to map the genes that control flowering time, across 22 environments, and identified 1,005 genes. In total, we found that 61.4% of the single-nucleotide polymorphisms (SNPs) associated with altitude were also associated with flowering time. More than half of the SNPs associated with altitude were within large structural variants (inversions, centromeres and pericentromeric regions). The combined mapping results indicate that although floral regulatory network genes contribute substantially to field variation, over 90% of the contributing genes probably have indirect effects. Our dual strategy can be used to harness the landrace diversity of plants and animals.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Change history
20 February 2017
In the version of this article initially published online, the name of author Martha Willcox was misspelled as Martha Wilcox. The error has been corrected in the print, PDF and HTML versions of this article.
References
Warburton, M.L. et al. Genetic diversity in CIMMYT nontemperate maize germplasm: landraces, open pollinated varieties and inbred lines. Crop Sci. 48, 617–624 (2008).
Wallace, J.G., Larsson, S.J. & Buckler, E.S. Entering the second century of maize quantitative genetics. Heredity 112, 30–38 (2014).
Remington, D.L. et al. Structure of linkage disequilibrium and phenotypic associations in the maize genome. Proc. Natl. Acad. Sci. USA 98, 11479–11484 (2001).
Romay, M.C. et al. Comprehensive genotyping of the USA national maize inbred seed bank. Genome Biol. 14, R55 (2013).
Hufford, M.B. et al. Comparative population genomics of maize domestication and improvement. Nat. Genet. 44, 808–811 (2012).
Mir, C. et al. Out of America: tracing the genetic footprints of the global diffusion of maize. Theor. Appl. Genet. 126, 2671–2682 (2013).
van Heerwaarden, J. et al. Genetic signals of origin, spread and introgression in a large sample of maize landraces. Proc. Natl. Acad. Sci. USA 108, 1088–1092 (2011).
Hufford, M.B. et al. The genomic signature of crop—wild introgression in maize. PLoS Genet. 9, e1003477 (2013).
Warburton, M.L. et al. Gene flow among different teosinte taxa and into the domesticated maize gene pool. Genet. Resour. Crop Evol. 58, 1243–1261 (2011).
McMullen, M.D. et al. Genetic properties of the maize nested-association-mapping population. Science 325, 737–740 (2009).
Li, C. et al. Quantitative trait loci mapping for yield components and kernel-related traits in multiple connected RIL populations in maize. Euphytica 193, 303–316 (2013).
Flint-Garcia, S.A. et al. Maize association population: a high-resolution platform for quantitative trait locus dissection. Plant J. 44, 1054–1064 (2005).
Peiffer, J.A. et al. The genetic architecture of maize height. Genetics 196, 1337–1356 (2014).
Buckler, E.S. et al. The genetic architecture of maize flowering time. Science 325, 714–718 (2009).
Harjes, C.E. et al. Natural genetic variation in lycopene ɛ-cyclase tapped for maize biofortification. Science 319, 330–333 (2008).
Tian, F. et al. Genome-wide association study of leaf architecture in the maize nested-association-mapping population. Nat. Genet. 43, 159–162 (2011).
Arteaga, M.C. et al. Genomic variation in recently collected maize landraces from Mexico. Genom. Data 7, 38–45 (2015).
Strigens, A., Schipprack, W., Reif, J.C. & Melchinger, A.E. Unlocking the genetic diversity of maize landraces with doubled haploids opens new avenues for breeding. PLoS One 8, e57234 (2013).
Salhuana, W., Jones, Q. & Sevilla, R. The Latin American Maize Project: model for rescue and use of irreplaceable germplasm. Diversity (Basel) 7, 40–42 (1991).
Pollak, L.M. The history and success of the public–private project on germplasm enhancement of maize (GEM). Adv. Agron. 78, 45–87 (2003).
Elshire, R.J. et al. A robust, simple genotyping-by-sequencing (GBS) approach for high-diversity species. PLoS One 6, e19379 (2011).
Browning, B.L. & Browning, S.R. Improving the accuracy and efficiency of identity-by-descent detection in population data. Genetics 194, 459–471 (2013).
Mantel, N. The detection of disease clustering and a generalized regression approach. Cancer Res. 27, 209–220 (1967).
Rodgers-Melnick, E. et al. Recombination in diverse maize is stable, predictable and associated with genetic load. Proc. Natl. Acad. Sci. USA 112, 3823–3828 (2015).
Chia, J.-M. et al. Maize HapMap2 identifies extant variation from a genome in flux. Nat. Genet. 44, 803–807 (2012).
Bertin, P., Madur, D., Combes, V., Dumas, F. & Brunel, D. Adaptation of maize to temperate climates: mid-density genome-wide association genetics and diversity patterns reveal key genomic regions, with a major contribution of the Vgt2 (ZCN8) locus. PLoS One 8, e71377 (2013).
Ducrocq, S. et al. Key impact of Vgt1 on flowering-time adaptation in maize: evidence from association-mapping and ecogeographical information. Genetics 178, 2433–2437 (2008).
Hirsch, C.N. et al. Insights into the maize pan-genome and pan-transcriptome. Plant Cell 26, 121–135 (2014).
Chardon, F. et al. Genetic architecture of flowering time in maize as inferred from quantitative trait loci meta-analysis and synteny conservation with the rice genome. Genetics 168, 2169–2185 (2004).
Hung, H.-Y. et al. ZmCCT and the genetic basis of day-length adaptation underlying the postdomestication spread of maize. Proc. Natl. Acad. Sci. USA 109, E1913–E1921 (2012).
Salvi, S. et al. Conserved noncoding genomic sequences associated with a flowering-time quantitative trait locus in maize. Proc. Natl. Acad. Sci. USA 104, 11376–11381 (2007).
Ziska, L.H., Teramura, A.H. & Sullivan, J.H. Physiological sensitivity of plants along an elevational gradient to UV-B radiation. Am. J. Bot. 79, 863–871 (1992).
Crimmins, T.M., Crimmins, M.A. & David Bertelsen, C. Complex responses to climate drivers in onset of spring flowering across a semi-arid elevation gradient. J. Ecol. 98, 1042–1051 (2010).
Ziello, C., Estrella, N., Kostova, M., Koch, E. & Menzel, A. Influence of altitude on phenology of selected plant species in the Alpine region (1971–2000). Clim. Res. 39, 227–234 (2009).
Pyhäjärvi, T., Hufford, M.B., Mezmouk, S. & Ross-Ibarra, J. Complex patterns of local adaptation in teosinte. Genome Biol. Evol. 5, 1594–1609 (2013).
Dobzhansky, T. & Sturtevant, A.H. Inversions in the chromosomes of Drosophila pseudoobscura. Genetics 23, 28–64 (1938).
Rieseberg, L.H., Whitton, J. & Gardner, K. Hybrid zones and the genetic architecture of a barrier to gene flow between two sunflower species. Genetics 152, 713–727 (1999).
Lowry, D.B. & Willis, J.H. A widespread chromosomal inversion polymorphism contributes to a major life-history transition, local adaptation and reproductive isolation. PLoS Biol. 8, e1000500 (2010).
Stuber, C.W., Lincoln, S.E., Wolff, D.W., Helentjaris, T. & Lander, E.S. Identification of genetic factors contributing to heterosis in a hybrid from two elite maize inbred lines using molecular markers. Genetics 132, 823–839 (1992).
Castelletti, S., Tuberosa, R., Pindo, M. & Salvi, S. A MITE transposon insertion is associated with differential methylation at the maize flowering time QTL Vgt1. G3 (Bethesda) 4, 805–812 (2014).
Meng, X., Muszynski, M.G. & Danilevskaya, O.N. The FT-like ZCN8 gene functions as a floral activator and is involved in photoperiod sensitivity in maize. Plant Cell 23, 942–960 (2011).
Danilevskaya, O.N., Meng, X., Hou, Z., Ananiev, E.V. & Simmons, C.R. A genomic and expression compendium of the expanded PEBP gene family from maize. Plant Physiol. 146, 250–264 (2008).
Camus-Kulandaivelu, L. et al. Patterns of molecular evolution associated with two selective sweeps in the Tb1–Dwarf8 region in maize. Genetics 180, 1107–1121 (2008).
Dong, Z. et al. A gene regulatory network model for floral transition of the shoot apex in maize and its dynamic modeling. PLoS One 7, e43450 (2012).
Federer, W.T. & Crossa, J.I. 4 screening experimental designs for quantitative trait loci, association mapping, genotype-by-environment interaction and other investigations. Front. Physiol. 3, 156 (2012).
Glaubitz, J.C. et al. TASSEL–GBS: a high-capacity genotyping-by-sequencing analysis pipeline. PLoS One 9, e90346 (2014).
Swarts, K., Li, H., Romero Navarro, J.A. & An, D. Novel methods to optimize genotypic imputation for low-coverage, next-generation sequence data in crop plants. Plant Genome 7, doi:10.3835/plantgenome2014.05.0023 (2014).
Dray, S. & Dufour, A.B. The ade4 package: implementing the duality diagram for ecologists. J. Stat. Softw. 22, 1–20 (2007).
Aulchenko, Y.S., de Koning, D.-J. & Haley, C. Genome-wide rapid association using mixed model and regression: a fast and simple method for genome-wide pedigree-based quantitative-trait-loci association analysis. Genetics 177, 577–585 (2007).
Zhang, Z. et al. Mixed linear model approach adapted for genome-wide association studies. Nat. Genet. 42, 355–360 (2010).
Lawrence, M. et al. Software for computing and annotating genomic ranges. PLOS Comput. Biol. 9, e1003118 (2013).
Lipka, A.E. et al. GAPIT: genome association and prediction integrated tool. Bioinformatics 28, 2397–2399 (2012).
Speed, D. & Balding, D.J. MultiBLUP: improved SNP-based prediction for complex traits. Genome Res. 24, 1550–1557 (2014).
Yang, J., Lee, S.H., Goddard, M.E. & Visscher, P.M. GCTA: a tool for genome-wide complex-trait analysis. Am. J. Hum. Genet. 88, 76–82 (2011).
Sekhon, R.S. et al. Genome-wide atlas of transcription during maize development. Plant J. 66, 553–563 (2011).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis (Springer-Verlag, 2009).
Acknowledgements
J.A.R.N., M.W., J.B., S.T., E.P., A.T., H.V.D., V.V., A.O., A.E.B., N.O.G.M., I.O.-M., F.S.V., A.G.E., G.A., P.W. and S.H. were supported by La Secretaría de Agricultura, Ganadería, Desarrollo Rural, Pesca y Alimentación (SAGARPA), Mexico under the MasAgro (Sustainable Modernization of Traditional Agriculture) initiative. J.A.R.N., C.R., K.S. and E.S.B. were supported by the US National Science Foundation (grant no. 1238014 and 0922493), and the USDA–ARS. We would like to thank ICAMEX and DuPont Pioneer Mexico for assistance in establishing the phenotypic trials.
Author information
Authors and Affiliations
Contributions
J.A.R.N. conducted the GWAS analyses; M.W. coordinated the execution of the phenotypic trials, and the collection and curation of the phenotypic data; J.B. developed the phenotypic experimental designs, formulated models and determined landrace parent–environment estimates; C.R. assisted with the GWAS analysis and data interpretation; K.S. performed genotype imputation; S.T., E.P., A.T., H.V.D.,V.V., A.O., A.E.B., N.O.G.M., I.O.-M. and A.G.E. conducted the phenotypic trials; F.S.V. and A.G.E. developed the test-cross germplasm; G.A., P.W. and E.S.B. developed the project concept; S.H. coordinated the genotypic data collection, meta-data creation and passport data curation; and J.A.R.N., S.H. and E.S.B. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
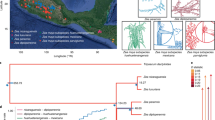
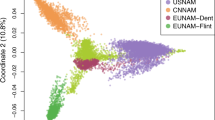
Supplementary Figure 2 First two principal coordinates from multidimensional scaling of the genetic distance among accessions.
Colors correspond to country of origin according to passport information of the accession.
Supplementary Figure 3 Landrace adaptation classes in MDS space.
LT corresponds to lowland tropical, ST to subtropical and HL to highland. Highland landraces span most of the first principal coordinate, displaying incomplete differentiation from middle- and low-elevation populations. Those landraces appear admixed with subtropical and low-elevation landraces and are almost entirely exclusive to Mexican material. In contrast, subtropical and tropical landraces are present admixed across both axes.
Supplementary Figure 4 Genome-wide view of the LD empirical threshold.
Red shaded areas represent the centromeres. In general, for all chromosomes, centromeres and the pericentromeric regions display higher LD than the rest of the genome. Two additional regions, shaded in gray, are the inversions on chromosomes 3 and 4. The dashed horizontal line represents the empirical LD threshold used to define the set of high-LD regions.
Supplementary Figure 5 Overlap rate with flowering time.
Overlap rate is estimated for the top associating SNPs of altitude and latitude at various P-value thresholds.
Supplementary Figure 6 MDS of the centromere of chromosome 5.
MDS for FOAM landrace accessions (blue) and the NAM founders (red). Most NAM founders cluster together, sharing one allele, with the other alleles corresponding to Il14H (top right), P39 (bottom) and CML333 (middle).
Supplementary Figure 10 GBS distribution of missing data.
Proportion missing before imputation by site and by individual for the GBS genotypes on the FOAM landrace parents.
Supplementary Figure 12 Median LD across windows for FOAM landrace parents.
The LD threshold was chosen based on change in slope and corresponds to the red line.
Supplementary Figure 14 QQ plot for the altitude GWAS across models.
Models correspond to the generalized linear model (GLM), a model with a population structure covariate in the from of 10 PCs (GLM + Q), a mixed model with a relatedness matrix (MLM + K) and a mixed model with relatedness and population structure (MLM Q + K).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–14 (PDF 1557 kb)
Supplementary Table 1
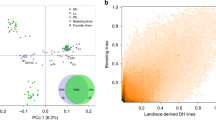
Phenotypic evaluation information. Lists years of evaluation, trial name, location, adaptation class, number of accessions evaluated, row length in meters, number of plants per row, and broad sense heritability estimates from variance partitioning model (XLSX 10 kb)
Supplementary Table 2
Top genes associated with female flowering (XLSX 61 kb)
Supplementary Table 3
Top genes associated with male flowering (XLSX 61 kb)
Supplementary Table 4
Gene Ontology enrichment for flowering time candidate genes (XLSX 12 kb)
Supplementary Table 5
Gene Ontology enrichment for genes associated with male and female flowering time in FOAM landrace panel (XLSX 10 kb)
Supplementary Table 6
Top SNPs associated with altitude (XLSX 117 kb)
Supplementary Table 7
Top SNPs associated with latitude (XLSX 248 kb)
Supplementary Table 8
CIMMYT inbred lines homozygous for the chromosome 4 inversion (XLSX 8 kb)
Rights and permissions
About this article
Cite this article
Romero Navarro, J., Willcox, M., Burgueño, J. et al. A study of allelic diversity underlying flowering-time adaptation in maize landraces. Nat Genet 49, 476–480 (2017). https://doi.org/10.1038/ng.3784
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3784
This article is cited by
-
Cytoplasmic genome contributions to domestication and improvement of modern maize
BMC Biology (2024)
-
Are cereal grasses a single genetic system?
Nature Plants (2024)
-
Plant pangenomes for crop improvement, biodiversity and evolution
Nature Reviews Genetics (2024)
-
Multi-locus genome-wide association studies reveal the dynamic genetic architecture of flowering time in chrysanthemum
Plant Cell Reports (2024)
-
Heritable microbiome variation is correlated with source environment in locally adapted maize varieties
Nature Plants (2024)