Abstract
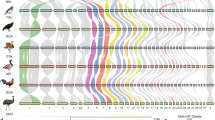
After birds diverged from mammals, different ancestral autosomes evolved into sex chromosomes in each lineage. In birds, females are ZW and males are ZZ, but in mammals females are XX and males are XY. We sequenced the chicken W chromosome, compared its gene content with our reconstruction of the ancestral autosomes, and followed the evolutionary trajectory of ancestral W-linked genes across birds. Avian W chromosomes evolved in parallel with mammalian Y chromosomes, preserving ancestral genes through selection to maintain the dosage of broadly expressed regulators of key cellular processes. We propose that, like the human Y chromosome, the chicken W chromosome is essential for embryonic viability of the heterogametic sex. Unlike other sequenced sex chromosomes, the chicken W chromosome did not acquire and amplify genes specifically expressed in reproductive tissues. We speculate that the pressures that drive the acquisition of reproduction-related genes on sex chromosomes may be specific to the male germ line.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Nanda, I. et al. 300 million years of conserved synteny between chicken Z and human chromosome 9. Nat. Genet. 21, 258–259 (1999).
Ross, M.T. et al. The DNA sequence of the human X chromosome. Nature 434, 325–337 (2005).
Fridolfsson, A.K. et al. Evolution of the avian sex chromosomes from an ancestral pair of autosomes. Proc. Natl. Acad. Sci. USA 95, 8147–8152 (1998).
Bellott, D.W. et al. Convergent evolution of chicken Z and human X chromosomes by expansion and gene acquisition. Nature 466, 612–616 (2010).
Handley, L.J., Ceplitis, H. & Ellegren, H. Evolutionary strata on the chicken Z chromosome: implications for sex chromosome evolution. Genetics 167, 367–376 (2004).
Lahn, B.T. & Page, D.C. Four evolutionary strata on the human X chromosome. Science 286, 964–967 (1999).
Zhou, Q. et al. Complex evolutionary trajectories of sex chromosomes across bird taxa. Science 346, 1246338 (2014).
Mueller, J.L. et al. Independent specialization of the human and mouse X chromosomes for the male germ line. Nat. Genet. 45, 1083–1087 (2013).
Charlesworth, B. & Charlesworth, D. The degeneration of Y chromosomes. Phil. Trans. R. Soc. Lond. B 355, 1563–1572 (2000).
Bellott, D.W. et al. Mammalian Y chromosomes retain widely expressed dosage-sensitive regulators. Nature 508, 494–499 (2014).
Soh, Y.Q. et al. Sequencing the mouse Y chromosome reveals convergent gene acquisition and amplification on both sex chromosomes. Cell 159, 800–813 (2014).
Hughes, J.F. et al. Strict evolutionary conservation followed rapid gene loss on human and rhesus Y chromosomes. Nature 483, 82–86 (2012).
Hughes, J.F. et al. Chimpanzee and human Y chromosomes are remarkably divergent in structure and gene content. Nature 463, 536–539 (2010).
Li, G. et al. Comparative analysis of mammalian Y chromosomes illuminates ancestral structure and lineage-specific evolution. Genome Res. 23, 1486–1495 (2013).
Skaletsky, H. et al. The male-specific region of the human Y chromosome is a mosaic of discrete sequence classes. Nature 423, 825–837 (2003).
Moghadam, H.K., Pointer, M.A., Wright, A.E., Berlin, S. & Mank, J.E. W chromosome expression responds to female-specific selection. Proc. Natl. Acad. Sci. USA 109, 8207–8211 (2012).
Mank, J.E., Hosken, D.J. & Wedell, N. Conflict on the sex chromosomes: cause, effect, and complexity. Cold Spring Harb. Perspect. Biol. 6, a017715 (2014).
Wright, A.E., Harrison, P.W., Montgomery, S.H., Pointer, M.A. & Mank, J.E. Independent stratum formation on the avian sex chromosomes reveals inter-chromosomal gene conversion and predominance of purifying selection on the W chromosome. Evolution 68, 3281–3295 (2014).
Ayers, K.L. et al. RNA sequencing reveals sexually dimorphic gene expression before gonadal differentiation in chicken and allows comprehensive annotation of the W-chromosome. Genome Biol. 14, R26 (2013).
Smeds, L. et al. Evolutionary analysis of the female-specific avian W chromosome. Nat. Commun. 6, 7330 (2015).
Yazdi, H.P. & Ellegren, H. Old but not (so) degenerated—slow evolution of largely homomorphic sex chromosomes in ratites. Mol. Biol. Evol. 31, 1444–1453 (2014).
Hori, T., Asakawa, S., Itoh, Y., Shimizu, N. & Mizuno, S. Wpkci, encoding an altered form of PKCI, is conserved widely on the avian W chromosome and expressed in early female embryos: implication of its role in female sex determination. Mol. Biol. Cell 11, 3645–3660 (2000).
Alföldi, J. et al. The genome of the green anole lizard and a comparative analysis with birds and mammals. Nature 477, 587–591 (2011).
St John, J.A. et al. Sequencing three crocodilian genomes to illuminate the evolution of archosaurs and amniotes. Genome Biol. 13, 415 (2012).
Green, R.E. et al. Three crocodilian genomes reveal ancestral patterns of evolution among archosaurs. Science 346, 1254449 (2014).
Zhang, J., Li, C., Zhou, Q. & Zhang, G. Improving the ostrich genome assembly using optical mapping data. Gigascience 4, 24 (2015).
Vicoso, B., Kaiser, V.B. & Bachtrog, D. Sex-biased gene expression at homomorphic sex chromosomes in emus and its implication for sex chromosome evolution. Proc. Natl. Acad. Sci. USA 110, 6453–6458 (2013).
Li, S. et al. Genomic signatures of near-extinction and rebirth of the crested ibis and other endangered bird species. Genome Biol. 15, 557 (2014).
Lahn, B.T. & Page, D.C. Functional coherence of the human Y chromosome. Science 278, 675–680 (1997).
Kaiser, V.B., Zhou, Q. & Bachtrog, D. Nonrandom gene loss from the Drosophila miranda neo-Y chromosome. Genome Biol. Evol. 3, 1329–1337 (2011).
White, M.A., Kitano, J. & Peichel, C.L. Purifying selection maintains dosage-sensitive genes during degeneration of the threespine stickleback Y chromosome. Mol. Biol. Evol. 32, 1981–1995 (2015).
Huang, N., Lee, I., Marcotte, E.M. & Hurles, M.E. Characterising and predicting haploinsufficiency in the human genome. PLoS Genet. 6, e1001154 (2010).
Merkin, J., Russell, C., Chen, P. & Burge, C.B. Evolutionary dynamics of gene and isoform regulation in mammalian tissues. Science 338, 1593–1599 (2012).
Yan, L. et al. Single-cell RNA–Seq profiling of human preimplantation embryos and embryonic stem cells. Nat. Struct. Mol. Biol. 20, 1131–1139 (2013).
Cunningham, F. et al. Ensembl 2015. Nucleic Acids Res. 43, D662–D669 (2015).
Zweier, C. et al. Haploinsufficiency of TCF4 causes syndromal mental retardation with intermittent hyperventilation (Pitt–Hopkins syndrome). Am. J. Hum. Genet. 80, 994–1001 (2007).
Tonkin, E.T., Wang, T.J., Lisgo, S., Bamshad, M.J. & Strachan, T. NIPBL, encoding a homolog of fungal Scc2-type sister chromatid cohesion proteins and fly Nipped-B, is mutated in Cornelia de Lange syndrome. Nat. Genet. 36, 636–641 (2004).
Amberger, J.S., Bocchini, C.A., Schiettecatte, F., Scott, A.F. & Hamosh, A. OMIM.org: Online Mendelian Inheritance in Man (OMIM®), an online catalog of human genes and genetic disorders. Nucleic Acids Res. 43, D789–D798 (2015).
UniProt Consortium. UniProt: a hub for protein information. Nucleic Acids Res. 43, D204–D212 (2015).
Skromne, I. & Stern, C.D. Interactions between Wnt and Vg1 signalling pathways initiate primitive streak formation in the chick embryo. Development 128, 2915–2927 (2001).
Papp, B., Pál, C. & Hurst, L.D. Dosage sensitivity and the evolution of gene families in yeast. Nature 424, 194–197 (2003).
Aitken, R.J. & Marshall Graves, J.A. The future of sex. Nature 415, 963 (2002).
Cocquet, J. et al. A genetic basis for a postmeiotic X versus Y chromosome intragenomic conflict in the mouse. PLoS Genet. 8, e1002900 (2012).
Cocquet, J. et al. The multicopy gene Sly represses the sex chromosomes in the male mouse germline after meiosis. PLoS Biol. 7, e1000244 (2009).
Conway, S.J. et al. Y353/B: a candidate multiple-copy spermiogenesis gene on the mouse Y chromosome. Mamm. Genome 5, 203–210 (1994).
Mueller, J.L. et al. The mouse X chromosome is enriched for multicopy testis genes showing postmeiotic expression. Nat. Genet. 40, 794–799 (2008).
Cockwell, A., MacKenzie, M., Youings, S. & Jacobs, P. A cytogenetic and molecular study of a series of 45,X fetuses and their parents. J. Med. Genet. 28, 151–155 (1991).
Hassold, T., Benham, F. & Leppert, M. Cytogenetic and molecular analysis of sex-chromosome monosomy. Am. J. Hum. Genet. 42, 534–541 (1988).
Hook, E.B. & Warburton, D. The distribution of chromosomal genotypes associated with Turner's syndrome: livebirth prevalence rates and evidence for diminished fetal mortality and severity in genotypes associated with structural X abnormalities or mosaicism. Hum. Genet. 64, 24–27 (1983).
Fechheimer, N.S. Origins of heteroploidy in chicken embryos. Poult. Sci. 60, 1365–1371 (1981).
Bloom, S.E. Chromosome abnormalities in chicken (Gallus domesticus) embryos: types, frequencies and phenotypic effects. Chromosoma 37, 309–326 (1972).
Zhao, D. et al. Somatic sex identity is cell autonomous in the chicken. Nature 464, 237–242 (2010).
Dunford, A. et al. Tumor-suppressor genes that escape from X-inactivation contribute to cancer sex bias. Nat. Genet. 49, 10–16 (2017).
Lee, M.K. et al. Construction and characterization of three BAC libraries for analysis of the chicken genome. Anim. Genet. 34, 151–152 (2003).
Wallis, J.W. et al. A physical map of the chicken genome. Nature 432, 761–764 (2004).
International Chicken Genome Sequencing Consortium. Sequence and comparative analysis of the chicken genome provide unique perspectives on vertebrate evolution. Nature 432, 695–716 (2004).
Zody, M.C. Investigation of Mechanics of Mutation and Selection by Comparative Sequencing. PhD thesis, Uppsala Univ. (2009).
Chen, N., Bellott, D.W., Page, D.C. & Clark, A.G. Identification of avian W-linked contigs by short-read sequencing. BMC Genomics 13, 183 (2012).
Itoh, Y. & Mizuno, S. Molecular and cytological characterization of SspI-family repetitive sequence on the chicken W chromosome. Chromosome Res. 10, 499–511 (2002).
Solovei, I., Ogawa, A., Naito, M., Mizuno, S. & Macgregor, H. Specific chromomeres on the chicken W lampbrush chromosome contain specific repetitive DNA sequence families. Chromosome Res. 6, 323–327 (1998).
Saitoh, Y. & Mizuno, S. Distribution of XhoI and EcoRI family repetitive DNA sequences into separate domains in the chicken W chromosome. Chromosoma 101, 474–477 (1992).
Morisson, M. et al. ChickRH6: a chicken whole-genome radiation hybrid panel. Genet. Sel. Evol. 34, 521–533 (2002).
Slonim, D., Kruglyak, L., Stein, L. & Lander, E. Building human genome maps with radiation hybrids. J. Comput. Biol. 4, 487–504 (1997).
Krasikova, A. et al. On the positions of centromeres in chicken lampbrush chromosomes. Chromosome Res. 14, 777–789 (2006).
Flicek, P., Keibler, E., Hu, P., Korf, I. & Brent, M.R. Leveraging the mouse genome for gene prediction in human: from whole-genome shotgun reads to a global synteny map. Genome Res. 13, 46–54 (2003).
Korf, I., Flicek, P., Duan, D. & Brent, M.R. Integrating genomic homology into gene structure prediction. Bioinformatics 17 (Suppl. 1), S140–S148 (2001).
Boardman, P.E. et al. A comprehensive collection of chicken cDNAs. Curr. Biol. 12, 1965–1969 (2002).
Hedges, S.B., Dudley, J. & Kumar, S. TimeTree: a public knowledge-base of divergence times among organisms. Bioinformatics 22, 2971–2972 (2006).
Bray, N.L., Pimentel, H., Melsted, P. & Pachter, L. Near-optimal probabilistic RNA–seq quantification. Nat. Biotechnol. 34, 525–527 (2016).
Robinson, M.D., McCarthy, D.J. & Smyth, G.K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Acknowledgements
We thank M. Delany (University of California, Davis) for UCD001 DNA, A. Vignal and M. Morrison (INRA Toulouse) for ChickRH6 radiation hybrid panel DNA, M. Lovett (Washington University, St. Louis) for chicken embryonic fibroblasts, C. Friedman and B. Trask for flow-sorted chicken W chromosomes, F. McCarthy for permission to use the Chickspress RNA–seq data set (PRJNA204941), and the “Chromas” Saint-Petersburg University Resource Center and L. Rapoport for technical assistance. This work was supported by the National Institutes of Health and the Howard Hughes Medical Institute. S.G. and E.G. were supported by the Russian Foundation of Basic Research (grant 16-04-01823a).
Author information
Authors and Affiliations
Contributions
D.W.B., H.S., W.C.W., A.G.C., E.G., R.K.W., and D.C.P. planned the project. D.W.B., H.S., T.-J.C., D.L., and N.C. developed female-specific sequence-tagged sites. D.W.B., H.S., T.-J.C., and L.B. performed clone mapping. D.W.B., T.-J.C., N.K., T.G., and C.K. performed clone sequencing. S.G. and T.P. performed FISH analyses. D.W.B. and T.-J.C. performed RH mapping. D.W.B. and H.S. performed sequence analyses. D.W.B. and D.C.P. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Phase-contrast and fluorescence images of lampbrush W-chromosome spreads
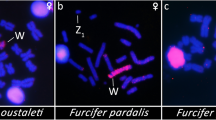
(a,f,k) Images of chicken Z–W lampbrush bivalents in phase contrast, oriented with the terminal giant lumpy (TGL) loops and the W chromosome at the top and the Z chromosome at the bottom. Scale bar, 10 μm. (b,g,l) FISH localization of BAC probes (red) on the same lampbrush chromosomes counterstained with DAPI (blue). (c–e,h–j,m–o) Magnification showing the DAPI-intense chromomeres of the W chromosome (c,h,m), the BAC hybridization signal (d,i,n), and the merged image (e,j,o). Chomomere 1 is furthest from the chiasma, and chromomere 7 is adjacent to the chiasma. CH261-75N4 localizes to chromomere 2 (b,e), CH261-107E4 localizes to chromomere 4 (g, j), and CH261-114G22 localizes to chromomere 7 (l,o).
Supplementary Figure 2 Dot plots of the pseudoautosomal region and the HINTW region.
Dot plots of nucleotide sequence identity in a window size of 50 bp and a step size of 1 bp. (a) Rectangular dot plot showing nucleotide identity between the most distal clones from the short arms of the W and Z chromosomes. The pseudoautosomal region of the Z and W chromosomes begins in telomeric repeats near the TCF4 gene. (b) Triangular dot plot showing nucleotide identity on the W chromosome in the 100 kb adjacent to the HINTW array. Two copies of HINTW are tandemly repeated outside the array.
Supplementary Figure 3 Ancestrally broad expression of Z–W pairs.
Violin plots marked with the median (black circle) and interquartile range (black bar) comparing the annotations of the human orthologs of ancestral Z–W gene pairs identified in chicken (dark pink); 4 species (chicken, collared flycatcher, crested ibis, and emu) (medium pink); and all 14 published female avian genomes (light pink) versus the human orthologs of the remainder of ancestral Z genes (light yellow). P values obtained using one-tailed Mann–Whitney U tests are reported with the associated test statistic, U. (a) The human orthologs of ancestral Z–W pairs are more broadly expressed in adult human tissues than other ancestral Z genes. Chicken Z–W pairs n = 26, other ancestral Z genes n = 516, P < 1.6 × 10–3, U = 9,012; 4 species Z–W pairs n = 70, other ancestral Z genes n = 472, P < 0.047, U = 18,563; 14 species Z–W pairs n = 133, other ancestral Z genes n = 409, P < 0.13, U = 28,960. (b) The human orthologs of ancestral Z–W pairs are more highly expressed in human blastocysts than other ancestral Z genes. Chicken Z–W pairs n = 26, other ancestral Z genes n = 495, P < 5.4 × 10–5, U = 9,333; 4 species Z–W pairs n = 68, other ancestral Z genes n = 453, P < 0.087, U = 18,156; 14 species Z–W pairs n = 129, other ancestral Z genes n = 392, P < 0.011, U = 28,720.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–3 (PDF 541 kb)
Supplementary Table 1
Tiling path accessions and coordinates. (XLSX 59 kb)
Supplementary Table 2
Ancestral Z genes and associated statistics. (XLSX 295 kb)
Supplementary Table 3
Ancestral ZW pairs from all 14 species. (XLSX 90 kb)
Supplementary Table 4
GO term enrichment. (XLSX 41 kb)
Supplementary Data 1
FASTA sequence of the chicken W chromosome assembly. (TXT 6913 kb)
Supplementary Data 2
FASTA sequence of transcripts of chicken Z–W gene pairs. (TXT 20 kb)
Supplementary Data 3
FASTA-formatted alignment of chicken Z–W gene pairs and human orthologs. (TXT 158 kb)
Supplementary Data 4
Radiation hybrid mapping data. (TXT 175 kb)
Rights and permissions
About this article
Cite this article
Bellott, D., Skaletsky, H., Cho, TJ. et al. Avian W and mammalian Y chromosomes convergently retained dosage-sensitive regulators. Nat Genet 49, 387–394 (2017). https://doi.org/10.1038/ng.3778
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3778
This article is cited by
-
Gene gain and loss from the Asian corn borer W chromosome
BMC Biology (2024)
-
Characterizing genetic variation on the Z chromosome in Schistosoma japonicum reveals host-parasite co-evolution
Parasites & Vectors (2024)
-
Evolution and expression patterns of the neo-sex chromosomes of the crested ibis
Nature Communications (2024)
-
Sex-limited chromosomes and non-reproductive traits
BMC Biology (2022)
-
Fragile, unfaithful and persistent Ys—on how meiosis can shape sex chromosome evolution
Heredity (2022)