Abstract
The extent of heterogeneity among driver gene mutations present in naturally occurring metastases—that is, treatment-naive metastatic disease—is largely unknown. To address this issue, we carried out 60× whole-genome sequencing of 26 metastases from four patients with pancreatic cancer. We found that identical mutations in known driver genes were present in every metastatic lesion for each patient studied. Passenger gene mutations, which do not have known or predicted functional consequences, accounted for all intratumoral heterogeneity. Even with respect to these passenger mutations, our analysis suggests that the genetic similarity among the founding cells of metastases was higher than that expected for any two cells randomly taken from a normal tissue. The uniformity of known driver gene mutations among metastases in the same patient has critical and encouraging implications for the success of future targeted therapies in advanced-stage disease.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Greaves, M. & Maley, C.C. Clonal evolution in cancer. Nature 481, 306–313 (2012).
Alizadeh, A.A. et al. Toward understanding and exploiting tumor heterogeneity. Nat. Med. 21, 846–853 (2015).
Yates, L.R. et al. Subclonal diversification of primary breast cancer revealed by multiregion sequencing. Nat. Med. 21, 751–759 (2015).
Gerlinger, M. et al. Genomic architecture and evolution of clear cell renal cell carcinomas defined by multiregion sequencing. Nat. Genet. 46, 225–233 (2014).
de Bruin, E.C. et al. Spatial and temporal diversity in genomic instability processes defines lung cancer evolution. Science 346, 251–256 (2014).
Zhang, J. et al. Intratumor heterogeneity in localized lung adenocarcinomas delineated by multiregion sequencing. Science 346, 256–259 (2014).
Gundem, G. et al. The evolutionary history of lethal metastatic prostate cancer. Nature 520, 353–357 (2015).
Hong, M.K.H. et al. Tracking the origins and drivers of subclonal metastatic expansion in prostate cancer. Nat. Commun. 6, 6605 (2015).
Kumar, A. et al. Substantial interindividual and limited intraindividual genomic diversity among tumors from men with metastatic prostate cancer. Nat. Med. 22, 369–378 (2016).
Jones, S. et al. Comparative lesion sequencing provides insights into tumor evolution. Proc. Natl. Acad. Sci. USA 105, 4283–4288 (2008).
Yachida, S. et al. Distant metastasis occurs late during the genetic evolution of pancreatic cancer. Nature 467, 1114–1117 (2010).
Campbell, P.J. et al. The patterns and dynamics of genomic instability in metastatic pancreatic cancer. Nature 467, 1109–1113 (2010).
Sanborn, J.Z. et al. Phylogenetic analyses of melanoma reveal complex patterns of metastatic dissemination. Proc. Natl. Acad. Sci. USA 112, 10995–11000 (2015).
Navin, N. et al. Tumour evolution inferred by single-cell sequencing. Nature 472, 90–94 (2011).
Tomasetti, C., Vogelstein, B. & Parmigiani, G. Half or more of the somatic mutations in cancers of self-renewing tissues originate prior to tumor initiation. Proc. Natl. Acad. Sci. USA 110, 1999–2004 (2013).
Vogelstein, B. et al. Cancer genome landscapes. Science 339, 1546–1558 (2013).
Makohon-Moore, A. & Iacobuzio-Donahue, C.A. Pancreatic cancer biology and genetics from an evolutionary perspective. Nat. Rev. Cancer 16, 553–565 (2016).
Barber, L.J. et al. Secondary mutations in BRCA2 associated with clinical resistance to a PARP inhibitor. J. Pathol. 229, 422–429 (2013).
Diaz, L.A. Jr. et al. The molecular evolution of acquired resistance to targeted EGFR blockade in colorectal cancers. Nature 486, 537–540 (2012).
Misale, S. et al. Emergence of KRAS mutations and acquired resistance to anti-EGFR therapy in colorectal cancer. Nature 486, 532–536 (2012).
Waddell, N. et al. Whole genomes redefine the mutational landscape of pancreatic cancer. Nature 518, 495–501 (2015).
Hoogstraat, M. et al. Genomic and transcriptomic plasticity in treatment-naive ovarian cancer. Genome Res. 24, 200–211 (2014).
Krasnitz, A., Sun, G., Andrews, P. & Wigler, M. Target inference from collections of genomic intervals. Proc. Natl. Acad. Sci. USA 110, E2271–E2278 (2013).
Davoli, T. et al. Cumulative haploinsufficiency and triplosensitivity drive aneuploidy patterns and shape the cancer genome. Cell 155, 948–962 (2013).
Siegel, R.L., Miller, K.D. & Jemal, A. Cancer statistics, 2016. CA Cancer J. Clin. 66, 7–30 (2016).
Embuscado, E.E. et al. Immortalizing the complexity of cancer metastasis: genetic features of lethal metastatic pancreatic cancer obtained from rapid autopsy. Cancer Biol. Ther. 4, 548–554 (2005).
Bailey, P. et al. Genomic analyses identify molecular subtypes of pancreatic cancer. Nature 531, 47–52 (2016).
Chang, M.T. et al. Identifying recurrent mutations in cancer reveals widespread lineage diversity and mutational specificity. Nat. Biotechnol. 34, 155–163 (2016).
Douville, C. et al. CRAVAT: cancer-related analysis of variants toolkit. Bioinformatics 29, 647–648 (2013).
Iacobuzio-Donahue, C.A., Velculescu, V.E., Wolfgang, C.L. & Hruban, R.H. Genetic basis of pancreas cancer development and progression: insights from whole-exome and whole-genome sequencing. Clin. Cancer Res. 18, 4257–4265 (2012).
Borazanci, E. et al. Adenosquamous carcinoma of the pancreas: molecular characterization of 23 patients along with a literature review. World J. Gastrointest. Oncol. 7, 132–140 (2015).
Kinde, I., Wu, J., Papadopoulos, N., Kinzler, K.W. & Vogelstein, B. Detection and quantification of rare mutations with massively parallel sequencing. Proc. Natl. Acad. Sci. USA 108, 9530–9535 (2011).
Yachida, S. & Iacobuzio-Donahue, C.A. The pathology and genetics of metastatic pancreatic cancer. Arch. Pathol. Lab. Med. 133, 413–422 (2009).
Olive, K.P. et al. Inhibition of Hedgehog signaling enhances delivery of chemotherapy in a mouse model of pancreatic cancer. Science 324, 1457–1461 (2009).
Campbell, P.J. et al. Subclonal phylogenetic structures in cancer revealed by ultra-deep sequencing. Proc. Natl. Acad. Sci. USA 105, 13081–13086 (2008).
Jones, S. et al. Core signaling pathways in human pancreatic cancers revealed by global genomic analyses. Science 321, 1801–1806 (2008).
Biankin, A.V. et al. Pancreatic cancer genomes reveal aberrations in axon guidance pathway genes. Nature 491, 399–405 (2012).
Schutte, M. et al. Abrogation of the Rb/p16 tumor-suppressive pathway in virtually all pancreatic carcinomas. Cancer Res. 57, 3126–3130 (1997).
Santarius, T., Shipley, J., Brewer, D., Stratton, M.R. & Cooper, C.S. A census of amplified and overexpressed human cancer genes. Nat. Rev. Cancer 10, 59–64 (2010).
Maley, C.C. et al. Genetic clonal diversity predicts progression to esophageal adenocarcinoma. Nat. Genet. 38, 468–473 (2006).
Fernández, L.C., Torres, M. & Real, F.X. Somatic mosaicism: on the road to cancer. Nat. Rev. Cancer 16, 43–55 (2016).
Blokzijl, F. et al. Tissue-specific mutation accumulation in human adult stem cells during life. Nature 538, 260–264 (2016).
Reiter, J.G. et al. Reconstructing metastatic seeding patterns of human cancers. Nat. Commun. http://dx.doi.org/10.1038/ncomms14114 (2017).
Sottoriva, A. et al. A Big Bang model of human colorectal tumor growth. Nat. Genet. 47, 209–216 (2015).
Waclaw, B. et al. A spatial model predicts that dispersal and cell turnover limit intratumour heterogeneity. Nature 525, 261–264 (2015).
Maddipati, R. & Stanger, B.Z. Pancreatic cancer metastases harbor evidence of polyclonality. Cancer Discov. 5, 1086–1097 (2015).
Gibson, W.J. et al. The genomic landscape and evolution of endometrial carcinoma progression and abdominopelvic metastasis. Nat. Genet. 48, 848–855 (2016).
Gerlinger, M. et al. Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. N. Engl. J. Med. 366, 883–892 (2012).
Jones, P.A. & Baylin, S.B. The epigenomics of cancer. Cell 128, 683–692 (2007).
Williams, M.J., Werner, B., Barnes, C.P., Graham, T.A. & Sottoriva, A. Identification of neutral tumor evolution across cancer types. Nat. Genet. 48, 238–244 (2016).
Bozic, I., Gerold, J.M. & Nowak, M.A. Quantifying clonal and subclonal passenger mutations in cancer evolution. PLoS Comput. Biol. 12, e1004731 (2016).
Witkiewicz, A.K. et al. Whole-exome sequencing of pancreatic cancer defines genetic diversity and therapeutic targets. Nat. Commun. 6, 6744 (2015).
Jiao, Y. et al. Exome sequencing identifies frequent inactivating mutations in BAP1, ARID1A and PBRM1 in intrahepatic cholangiocarcinomas. Nat. Genet. 45, 1470–1473 (2013).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J.R. Stat. Soc. 57, 289–300 (1995).
Salari, R. et al. Inference of tumor phylogenies with improved somatic mutation discovery. J. Comput. Biol. 20, 933–944 (2013).
El-Kebir, M., Oesper, L., Acheson-Field, H. & Raphael, B.J. Reconstruction of clonal trees and tumor composition from multi-sample sequencing data. Bioinformatics 31, i62–i70 (2015).
Popic, V. et al. Fast and scalable inference of multi-sample cancer lineages. Genome Biol. 16, 91 (2015).
Boeva, V. et al. Control-FREEC: a tool for assessing copy number and allelic content using next-generation sequencing data. Bioinformatics 28, 423–425 (2012).
Krzywinski, M. et al. Circos: an information aesthetic for comparative genomics. Genome Res. 19, 1639–1645 (2009).
Lappalainen, I. et al. The European Genome-phenome Archive of human data consented for biomedical research. Nat. Genet. 47, 692–695 (2015).
Acknowledgements
We thank the Memorial Sloan Kettering Cancer Center Molecular Cytology core facility for immunohistochemistry staining. This work was supported by Office of Naval Research grant N00014-16-1-2914, the Bill and Melinda Gates Foundation (OPP1148627), and a gift from B. Wu and E. Larson (M.A.N.), National Institutes of Health grants CA179991 (C.A.I.-D. and I.B.), F31 CA180682 (A.P.M.-M.), CA43460 (B.V.), and P50 CA62924, the Monastra Foundation, the Virginia and D.K. Ludwig Fund for Cancer Research, the Lustgarten Foundation for Pancreatic Cancer Research, the Sol Goldman Center for Pancreatic Cancer Research, the Sol Goldman Sequencing Center, ERC Start grant 279307: Graph Games (J.G.R., D.K., and C.K.), Austrian Science Fund (FWF) grant P23499-N23 (J.G.R., D.K., and C.K.), and FWF NFN grant S11407-N23 RiSE/SHiNE (J.G.R., D.K., and C.K.).
Author information
Authors and Affiliations
Contributions
C.I.D. and A.M.M. performed the autopsies. C.I.D., A.P.M.-M., R.H.H., L.D.W., B.V., K.W.K., N.P., M.Z., F.W., and Y.J. designed experiments. A.M.M., J.R., I.B., F.W., J.H., and M.A. performed biostatistical analyses. A.M.M., M.Z., B.J., and Z.A.K. performed the experiments. J.G.R., I.B., J.H., D.K., and K.C. performed computational analysis. J.R., I.B., B.A., and M.A.N. performed modeling. All authors interpreted the data. C.A.I.-D., A.M.M., and B.V. wrote the manuscript, J.R., I.B., and M.A.N. provided input to the manuscript, and all authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Hierarchical clustering for four cases.
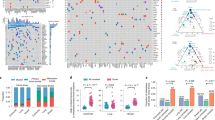
Samples are indicated along the y axis, and variants are listed along the top of the diagram. Colors correspond to discrete tumor samples and follow the rainbow spectrum from the Treeomics phylogenies, scaling from ancestral to descendant, as indicated by the evolutionary relationships. Samples that are uncolored represent those analyzed by hierarchical clustering only. The variant status in each sample is shown in blue (for present), dark red (for absent), or light red (for unknown due to low coverage). Hierarchical clustering using UPGMA (unweighted pair group method with arithmetic mean) is indicated on the right y axes. Primary tumors are labeled as “PT” followed by a number, lymph node metastases are labeled as “NoM” followed by a number, liver metastases are labeled as “LiM” followed by a number, lung metastases are labeled as “LuM” followed by a number, and peritoneal metastases are labeled as “PeM” followed by a number. (a) Pam01. (b) Pam02. (c) Pam03. (d) Pam04.
Supplementary Figure 2 Immunohistochemical analysis of proteins encoded by four major driver genes.
Protein expression was evaluated in tumor tissues. Driver genes evaluated by immunohistochemistry are listed in columns, and tumor tissues from each case are presented in rows. In all images, red arrows point to positively staining non-neoplastic cells (internal controls) and black arrows point to positively staining nuclei in neoplastic cells. The patterns observed for the individual metastases sequenced in each patient were reproducibly observed in the primary carcinoma and other metastases of the same patient, as we have previously demonstrated59. Scale bar, 10 μm.
Supplementary Figure 3 Distributions of copy number variations in Pam02.
Circos plots showing statistically significant CNVs in Pam02 whole-genome samples. For each sample ring, the y axis spans –2 to 2, with 0 representing a normal diploid copy number in unaffected regions, deletions represented as –1 or –2, and amplifications represented as 1 or 2. CNVs of >2 were scored as 2. All values were log2 transformed for visualization. The outermost ring shows the chromosomes in clockwise order. Deletions are shown in blue, while amplifications are shown in red. Gene names are those described in Supplementary Table 7. From innermost to outermost, the samples are PT18, PT4, PT9, LiM6, LiM5, LiM2, LiM8, LiM3, LiM7, and LiM1.
Supplementary Figure 4 Distributions of copy number variations in Pam03.
Circos plots showing statistically significant CNVs in whole-genome samples. For each sample ring, the y axis goes from –2 to 2, with a central black line representing a normal diploid copy number in unaffected regions, deletions represented as –1 or –2, and amplifications represented as 1 or 2. CNVs of >2 were scored as 2. The values are log2 transformed for visualization. The outermost ring shows the chromosomes in clockwise order. Deletions are shown in blue, while amplifications are shown in red. Gene names are those described in Supplementary Table 7. The innermost ring is PT12, followed by PT10, PT11, LuM3, LiM2, LiM4, LiM5, LiM3, LiM1, LuM1, and LuM2.
Supplementary Figure 5 Distributions of copy number variations in Pam04.
Circos plots showing statistically significant CNVs in whole-genome samples. For each sample ring, the y axis goes from –2 to 2, with a central black line representing a normal diploid copy number in unaffected regions, deletions represented as –1 or –2, and amplifications represented as 1 or 2. CNVs of >2 were scored as 2. The values are log2 transformed for visualization. The outermost ring shows the chromosomes in clockwise order. Deletions are shown in blue, while amplifications are shown in red. Gene names are those described in Supplementary Table 7. The innermost ring is PT27, followed by PT2, PT26, PeM3, PeM2, PeM1, PeM6, PeM5, and PeM4.
Supplementary Figure 6 B-allele frequencies for four Pam01 metastases.
For tumors that were whole-genome sequenced, B-allele frequencies are plotted for >3,000 SNPs per chromosome. Each chromosome is aligned sequentially and colored according to the color spectrum. The y axis represents frequency; the normal range is represented by the middle blue bar. Major loss-of-heterozygosity events (black arrowheads) are observable in all metastases. The differences in the patterns of changes in B-allele frequency are likely caused by the varying neoplastic cell content in the different samples as well as other artifacts.
Supplementary Figure 7 B-allele frequencies for Pam02 primary tumor sections and metastases.
For tumors that were whole-genome sequenced, B-allele frequencies are plotted for >3,000 SNPs per chromosome. Each chromosome is aligned sequentially and colored according to the color spectrum. The y axis represents frequency; the normal range is represented by the middle blue bar. Major loss-of-heterozygosity events (black arrowheads) are observable in all primary tumor sections and metastases. The differences in the patterns of change in B-allele frequency are likely caused by the varying neoplastic cell content in the different samples as well as other artifacts.
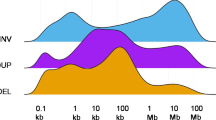
Supplementary Figure 8 Structural variants identified in Pam01–Pam04 samples.
Each type of structural variant is assigned a distinct color. Samples are labeled along the x axis, while numbers of structural variants are shown along the y axis.
Supplementary Figure 9 Distributions of metastatic disease in the Pam13 and Pam16 patients with cancer.
Anatomical locations of the primary carcinomas and discrete metastases used for whole-exome sequencing.
Supplementary Figure 10 Inferred phylogeny of primary tumor sections and metastases for patient Pam03.
(a) Time is represented on the left axis, and divergence is represented on the x axis. Colors correspond to discrete tumor samples and follow the rainbow spectrum, scaling from ancestral to descendant, as indicated by the evolutionary relationships. The ATM mutation was originally inferred to be wild type in primary tumor section PT1; however, targeted sequencing had insufficient coverage in PT1 and hence Treeomics misplaced the mutation (shown correctly here). (b) See Supplementary Table 3 for sample identity. Primary tumors are labeled at “PT” followed by a number, and the remaining samples are metastases labeled by organ. Hypothetical subclones are indicated by “SC” followed by the subclone number. The numbers of acquired mutations are in blue with a plus sign. Percentages denote bootstrapping values. Phylogeny was inferred by Treeomics43. (c) The dimensions of the original primary tumor in centimeters. (d) Primary tumor slices are numbered according to the original sectioning and plane order. See Supplementary Table 3 for sample identity. Metastases are labeled by organ followed by a metastasis number.
Supplementary Figure 11 Inferred phylogeny of primary tumor sections and metastases for patient Pam04.
(a) Time is represented on the left axis, and divergence is represented on the x axis. Colors correspond to discrete tumor samples and follow the rainbow spectrum, scaling from ancestral to descendant, as indicated by the evolutionary relationships. The KRAS mutation was originally inferred to be wild type in primary tumor section PT1; however, targeted sequencing had insufficient coverage in PT1 and hence Treeomics misplaced the mutation (shown correctly here). (b) See Supplementary Table 3 for sample identity. Primary tumors are labeled at “PT” followed by a number, and the remaining samples are metastases labeled by organ. Hypothetical subclones are indicated by “SC” followed by the subclone number. The numbers of acquired mutations are in blue with a plus sign. Percentages denote bootstrapping values. Phylogeny was inferred by Treeomics43. (c) The dimensions of the original primary tumor in centimeters. (d) Primary tumor slices are numbered according to the original sectioning and plane order. See Supplementary Table 3 for sample identity. Metastases are labeled by organ followed by a metastasis number.
Supplementary Figure 12 Inferred phylogeny for Pam01.
Time is represented on the left axis, and divergence is represented on the x axis. See Supplementary Table 3 for sample identity. Colors correspond to discrete tumor samples and follow the rainbow spectrum, scaling from ancestral to descendant, as indicated by the evolutionary relationships. Hypothetical subclones are indicated by “SC” followed by the subclone number. The numbers of acquired mutations are in blue with a plus sign. Percentages denote bootstrapping values. Phylogeny was inferred by Treeomics43.
Supplementary Figure 13 Inferred phylogeny for Pam13.
Time is represented on the left axis, and divergence is represented on the x axis. See Supplementary Table 3 for sample identity. Colors correspond to discrete tumor samples and follow the rainbow spectrum, scaling from ancestral to descendant, as indicated by the evolutionary relationships. Hypothetical subclones are indicated by “SC” followed by the subclone number. The numbers of acquired mutations are in blue with a plus sign. Percentages denote bootstrapping values. Phylogeny was inferred by Treeomics43. The KRAS variant was visualized in every tumor sample during manual review.
Supplementary Figure 14 Inferred phylogeny for Pam16.
Time is represented on the left axis, and divergence is represented on the x axis. See Supplementary Table 3 for sample identity. Colors correspond to discrete tumor samples and follow the rainbow spectrum, scaling from ancestral to descendant, as indicated by the evolutionary relationships. Hypothetical subclones are indicated by “SC” followed by the subclone number. The numbers of acquired mutations are in blue with a plus sign. Percentages denote bootstrapping values. Phylogeny was inferred by Treeomics43.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–14, Supplementary Tables 1 and 2, and Supplementary Note (PDF 2788 kb)
Supplementary Table 3
Samples analyzed. (XLSX 24 kb)
Supplementary Table 4
Average coverage per base. (XLSX 45 kb)
Supplementary Table 5
Summary of somatic copy number alterations identified in whole-genome sequencing samples. (XLSX 41 kb)
Supplementary Table 6
Major driver gene mutations identified in each patient. (XLSX 49 kb)
Supplementary Table 7
SCNAs identified in known PDAC driver genes. (XLSX 177 kb)
Supplementary Table 8
Candidate structural variants identified in Pam01. (XLSX 298 kb)
Supplementary Table 9
Candidate structural variants identified in Pam02. (XLSX 943 kb)
Supplementary Table 10
Candidate structural variants identified in Pam03. (XLSX 1079 kb)
Supplementary Table 11
Candidate structural variants identified in Pam04. (XLSX 639 kb)
Supplementary Table 12
Variants validated by targeted sequencing. (XLSX 102 kb)
Supplementary Table 13
Jaccard similarity coefficients of metastases based on stringently filtered whole-genome sequencing and whole-exome sequencing. (XLSX 46 kb)
Supplementary Table 14
Similarity coefficients of normal organs from Blokzjil et al. (XLSX 9 kb)
Supplementary Table 15
Genetic distances among metastases based on targeted sequencing. (XLSX 44 kb)
Supplementary Table 16
Jaccard similarity coefficients of metastases based on targeted sequencing (founder mutations excluded). (XLSX 44 kb)
Supplementary Table 17
Genetic distances among metastases based on whole-genome sequencing. (XLSX 47 kb)
Supplementary Table 18
Variants identified by whole-exome sequencing in the validation set. (XLSX 87 kb)
Supplementary Table 19
Primers. (XLSX 41 kb)
Rights and permissions
About this article
Cite this article
Makohon-Moore, A., Zhang, M., Reiter, J. et al. Limited heterogeneity of known driver gene mutations among the metastases of individual patients with pancreatic cancer. Nat Genet 49, 358–366 (2017). https://doi.org/10.1038/ng.3764
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3764
This article is cited by
-
Optimization of a mouse model of pancreatic cancer to simulate the human phenotypes of metastasis and cachexia
BMC Cancer (2024)
-
Cancer, metastasis, and the epigenome
Molecular Cancer (2024)
-
HATCHet2: clone- and haplotype-specific copy number inference from bulk tumor sequencing data
Genome Biology (2024)
-
Transient loss of Polycomb components induces an epigenetic cancer fate
Nature (2024)
-
The glutathione S-transferase Gstt1 drives survival and dissemination in metastases
Nature Cell Biology (2024)