Abstract
It has been proposed that the CLOCK–ARNTL (BMAL1) complex drives circadian transcription of thousands of genes, including Per and Cry family genes that encode suppressors of CLOCK–ARNTL-dependent transcription1,2,3. However, recent studies demonstrated that 70–80% of circadian-oscillating mRNAs have no obvious rhythms in their de novo transcription4,5, indicating the potential importance of post-transcriptional regulation. Our CLOCK-ChIP-seq analysis identified rhythmic expression of adenosine deaminase, RNA-specific, B1 (Adarb1, also known as Adar2), an adenosine-to-inosine (A-to-I) RNA-editing enzyme. RNA-seq showed circadian rhythms of ADARB1-mediated A-to-I editing in a variety of transcripts. In Adarb1-knockout mice, rhythms of large populations of mRNA were attenuated, indicating a profound impact of ADARB1-mediated A-to-I editing on RNA rhythms. Furthermore, Adarb1-knockout mice exhibited short-period rhythms in locomotor activity and gene expression. These phenotypes were associated with abnormal accumulation of CRY2. The present study identifies A-to-I RNA editing as a key mechanism of post-transcriptional regulation in the circadian clockwork.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Dunlap, J.C. Molecular bases for circadian clocks. Cell 96, 271–290 (1999).
Hastings, M.H., Reddy, A.B. & Maywood, E.S. A clockwork web: circadian timing in brain and periphery, in health and disease. Nat. Rev. Neurosci. 4, 649–661 (2003).
Hughes, M.E. et al. Harmonics of circadian gene transcription in mammals. PLoS Genet. 5, e1000442 (2009).
Koike, N. et al. Transcriptional architecture and chromatin landscape of the core circadian clock in mammals. Science 338, 349–354 (2012).
Menet, J.S., Rodriguez, J., Abruzzi, K.C. & Rosbash, M. Nascent-Seq reveals novel features of mouse circadian transcriptional regulation. eLife 1, e00011 (2012).
Yoshitane, H. et al. CLOCK-controlled polyphonic regulation of circadian rhythms through canonical and noncanonical E-boxes. Mol. Cell. Biol. 34, 1776–1787 (2014).
Vollmers, C. et al. Circadian oscillations of protein-coding and regulatory RNAs in a highly dynamic mammalian liver epigenome. Cell Metab. 16, 833–845 (2012).
Hogg, M., Paro, S., Keegan, L.P. & O'Connell, M.A. RNA editing by mammalian ADARs. Adv. Genet. 73, 87–120 (2011).
Nishikura, K. Functions and regulation of RNA editing by ADAR deaminases. Annu. Rev. Biochem. 79, 321–349 (2010).
Shimba, S. et al. Deficient of a clock gene, brain and muscle Arnt-like protein-1 (BMAL1), induces dyslipidemia and ectopic fat formation. PLoS One 6, e25231 (2011).
Barbon, A., Vallini, I., La Via, L., Marchina, E. & Barlati, S. Glutamate receptor RNA editing: a molecular analysis of GluR2, GluR5 and GluR6 in human brain tissues and in NT2 cells following in vitro neural differentiation. Brain Res. Mol. Brain Res. 117, 168–178 (2003).
Huang, H. et al. RNA editing of the IQ domain in Ca(v)1.3 channels modulates their Ca2+-dependent inactivation. Neuron 73, 304–316 (2012).
Rueter, S.M., Dawson, T.R. & Emeson, R.B. Regulation of alternative splicing by RNA editing. Nature 399, 75–80 (1999).
Maas, S., Patt, S., Schrey, M. & Rich, A. Underediting of glutamate receptor GluR-B mRNA in malignant gliomas. Proc. Natl. Acad. Sci. USA 98, 14687–14692 (2001).
Higuchi, M. et al. Point mutation in an AMPA receptor gene rescues lethality in mice deficient in the RNA-editing enzyme ADAR2. Nature 406, 78–81 (2000).
Kiran, A.M., O'Mahony, J.J., Sanjeev, K. & Baranov, P.V. Darned in 2013: inclusion of model organisms and linking with Wikipedia. Nucleic Acids Res. 41, D258–D261 (2013).
Ramaswami, G. & Li, J.B. RADAR: a rigorously annotated database of A-to-I RNA editing. Nucleic Acids Res. 42, D109–D113 (2014).
Wulff, B.-E., Sakurai, M. & Nishikura, K. Elucidating the inosinome: global approaches to adenosine-to-inosine RNA editing. Nat. Rev. Genet. 12, 81–85 (2011).
Eggington, J.M., Greene, T. & Bass, B.L. Predicting sites of ADAR editing in double-stranded RNA. Nat. Commun. 2, 319 (2011).
Riedmann, E.M., Schopoff, S., Hartner, J.C. & Jantsch, M.F. Specificity of ADAR-mediated RNA editing in newly identified targets. RNA 14, 1110–1118 (2008).
Liscovitch, N., Bazak, L., Levanon, E.Y. & Chechik, G. Positive correlation between ADAR expression and its targets suggests a complex regulation mediated by RNA editing in the human brain. RNA Biol. 11, 1447–1456 (2014).
Kanehisa, M., Goto, S., Sato, Y., Furumichi, M. & Tanabe, M. KEGG for integration and interpretation of large-scale molecular data sets. Nucleic Acids Res. 40, D109–D114 (2012).
Jepson, J.E.C. et al. Engineered alterations in RNA editing modulate complex behavior in Drosophila: regulatory diversity of adenosine deaminase acting on RNA (ADAR) targets. J. Biol. Chem. 286, 8325–8337 (2011).
Du, N.-H., Arpat, A.B., De Matos, M. & Gatfield, D. MicroRNAs shape circadian hepatic gene expression on a transcriptome-wide scale. eLife 3, e02510 (2014).
Garcia, D.M. et al. Weak seed-pairing stability and high target-site abundance decrease the proficiency of lsy-6 and other microRNAs. Nat. Struct. Mol. Biol. 18, 1139–1146 (2011).
Betel, D., Wilson, M., Gabow, A., Marks, D.S. & Sander, C. The microRNA.org resource: targets and expression. Nucleic Acids Res. 36, D149–D153 (2008).
Kawahara, Y. et al. Frequency and fate of microRNA editing in human brain. Nucleic Acids Res. 36, 5270–5280 (2008).
Vesely, C., Tauber, S., Sedlazeck, F.J., von Haeseler, A. & Jantsch, M.F. Adenosine deaminases that act on RNA induce reproducible changes in abundance and sequence of embryonic miRNAs. Genome Res. 22, 1468–1476 (2012).
Kawahara, Y., Zinshteyn, B., Chendrimada, T.P., Shiekhattar, R. & Nishikura, K. RNA editing of the microRNA-151 precursor blocks cleavage by the Dicer-TRBP complex. EMBO Rep. 8, 763–769 (2007).
Kojima, S. & Green, C.B. Circadian genomics reveal a role for post-transcriptional regulation in mammals. Biochemistry 54, 124–133 (2015).
Kojima, S., Shingle, D.L. & Green, C.B. Post-transcriptional control of circadian rhythms. J. Cell Sci. 124, 311–320 (2011).
Lim, C. & Allada, R. Emerging roles for post-transcriptional regulation in circadian clocks. Nat. Neurosci. 16, 1544–1550 (2013).
Hughes, M.E., Grant, G.R., Paquin, C., Qian, J. & Nitabach, M.N. Deep sequencing the circadian and diurnal transcriptome of Drosophila brain. Genome Res. 22, 1266–1281 (2012).
Wang, Q. et al. Altered G protein-coupling functions of RNA editing isoform and splicing variant serotonin2C receptors. J. Neurochem. 74, 1290–1300 (2000).
Yoo, S.-H. et al. PERIOD2:LUCIFERASE real-time reporting of circadian dynamics reveals persistent circadian oscillations in mouse peripheral tissues. Proc. Natl. Acad. Sci. USA 101, 5339–5346 (2004).
Kon, N. et al. Activation of TGF-β/activin signalling resets the circadian clock through rapid induction of Dec1 transcripts. Nat. Cell Biol. 10, 1463–1469 (2008).
Yoshitane, H. et al. Roles of CLOCK phosphorylation in suppression of E-box-dependent transcription. Mol. Cell. Biol. 29, 3675–3686 (2009).
Sasaki, M., Yoshitane, H., Du, N.-H., Okano, T. & Fukada, Y. Preferential inhibition of BMAL2-CLOCK activity by PER2 reemphasizes its negative role and a positive role of BMAL2 in the circadian transcription. J. Biol. Chem. 284, 25149–25159 (2009).
Hirano, A. et al. FBXL21 regulates oscillation of the circadian clock through ubiquitination and stabilization of cryptochromes. Cell 152, 1106–1118 (2013).
Yang, J.H., Sklar, P., Axel, R. & Maniatis, T. Editing of glutamate receptor subunit B pre-mRNA in vitro by site-specific deamination of adenosine. Nature 374, 77–81 (1995).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet.journal 17, 10–12 (2011).
Wang, K. et al. MapSplice: accurate mapping of RNA-seq reads for splice junction discovery. Nucleic Acids Res. 38, e178 (2010).
Barnett, D.W., Garrison, E.K., Quinlan, A.R., Strömberg, M.P. & Marth, G.T. BamTools: a C++ API and toolkit for analyzing and managing BAM files. Bioinformatics 27, 1691–1692 (2011).
Oster, H., Damerow, S., Hut, R.A. & Eichele, G. Transcriptional profiling in the adrenal gland reveals circadian regulation of hormone biosynthesis genes and nucleosome assembly genes. J. Biol. Rhythms 21, 350–361 (2006).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Trapnell, C. et al. Differential analysis of gene regulation at transcript resolution with RNA-seq. Nat. Biotechnol. 31, 46–53 (2013).
Yang, R. & Su, Z. Analyzing circadian expression data by harmonic regression based on autoregressive spectral estimation. Bioinformatics 26, i168–i174 (2010).
Hughes, M.E., Hogenesch, J.B. & Kornacker, K. JTK_CYCLE: an efficient nonparametric algorithm for detecting rhythmic components in genome-scale data sets. J. Biol. Rhythms 25, 372–380 (2010).
Vacic, V., Iakoucheva, L.M. & Radivojac, P. Two Sample Logo: a graphical representation of the differences between two sets of sequence alignments. Bioinformatics 22, 1536–1537 (2006).
Jiang, M., Anderson, J., Gillespie, J. & Mayne, M. uShuffle: a useful tool for shuffling biological sequences while preserving the k-let counts. BMC Bioinformatics 9, 192 (2008).
Kent, W.J. et al. The human genome browser at UCSC. Genome Res. 12, 996–1006 (2002).
Quinlan, A.R. & Hall, I.M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Chan, P.P. & Lowe, T.M. GtRNAdb: a database of transfer RNA genes detected in genomic sequence. Nucleic Acids Res. 37, D93–D97 (2009).
Acknowledgements
We thank K. Yugi, Y. Sugiura and T. Kokaji for their help with metabolic analysis, and we thank T. Nakagawa for his help with RNA-seq data analysis. We also thank J. Nakano for critical comments on the manuscript. Computations were partially performed on the NIG supercomputer at ROIS National Institute of Genetics. Adarb1−/−Gria2R/R mice were obtained from MMRRC 8U42OD010924-13. This work was partially supported by Grants-in-Aid for Scientific Research and Innovative Areas Genome Science from MEXT of Japan (to H.Y., W.I. and Y.F.), by the Japan Prize foundation (to H.Y.) and by Uchang Cho Institute of Science (to H.Y.). H.T. is supported by a JSPS research fellowship for young scientists.
Author information
Authors and Affiliations
Contributions
H.T. performed almost all the experiments. H.T., H.O. and Y.S. conducted the transcriptome experiments and performed bioinformatic analysis. S.S. generated Arntl-knockout mice. S.K. supervised the KEGG pathway analysis and the metabolic experiments. H.Y., W.I. and Y.F. supervised the project. H.T., H.Y., H.O. and Y.F. co-wrote the manuscript. All authors discussed the results and contributed to the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Circadian profiles of Adar b1 transcript and ADARB1 protein in mouse tissues.
(a) Overlap of rhythmic transcripts identified in the previous transcriptome analyses. The mouse liver samples were prepared under DD conditions in Yoshitane et al.6, Koike et al.4 and Vollmers et al.7 and under LD conditions in Menet et al.5. (b) Circadian expression profiles of two isoforms of Adar p150 and p110 in the mouse liver examined by qRT-PCR analysis. The signals were normalized to Rps29 (mean ± SEM; n = 3). (c) Temporal profiles of Arntl and Adarb1 mRNAs (determined by qRT-PCR) and the editing levels at the Flnb Q/R site (determined by direct sequencing analysis) in mouse liver through 3 days in DD. (d) Temporal expression profiles of Adarb1, Adarb1 long isoform, Adar p150, Adar p110, Adarb2, Dbp and Arntl in mouse tissues. The signals were normalized to Rps29 (mean ± SEM; n = 3). (e) Temporal profiles of the editing levels at the Q/R site of Flnb, the Y/C and I/M sites of Cacna1d and the A and B sites of Htr2c in direct sequencing analysis (mean ± SEM; n = 3). (f) Temporal profiles of ADARB1 protein levels (top panel, indicated by an arrowhead) in control and Adarb1-KO liver nuclear extracts. Non-specific bands were indicated by asterisks. TBP served as a loading control (bottom panel). (b, d, e) The temporal changes were analyzed by one-way ANOVA, * p < 0.05, ** p < 0.01, *** p < 0.001 and n.s. p ≧ 0.05.
Supplementary Figure 2 Circadian regulation of A-to-I RNA editing in mouse liver.
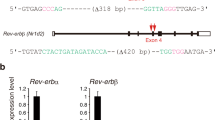
(a) Direct sequencing chromatograms from RT-PCR products of Cdk13 mRNA at various times of the day. (b) Circadian profiles of editing levels at the Cdk13 Q/R site in direct sequencing analysis (mean ± SEM; n = 3). (c) Regulation of alternative splicing of Adarb1 transcripts by Adarb1-mediated editing. The self-editing site is shown as a closed circle, and the indicated PCR primer set amplifies only the long isoform. (d) Circadian expression profiles of the long (top) and short isoforms (middle) of Adarb1 transcripts, determined by RT-PCR analyses. The relative band intensities [long / (long + short) (%)] were plotted (bottom). The gene specific primers used in this RT-PCR were described in Supplementary Table 6. (e) Direct sequencing chromatograms from RT-PCR products of Cog3 and Copa mRNAs at various times of the day. (f) Circadian profiles of editing levels [G / (A+G) (%)] at the indicated editing sites in RNA-Seq analysis (top and 3rd panels) and in direct sequencing analysis (2nd and bottom panels, mean ± SEM; n = 3). (g) Neighbor preferences of ADAR and ADARB1 represented by Two Sample Logo sequence motifs. Logo displays enriched bases above the upper line and depleted bases below the lower line on both sides of the central edited adenosine. (h) Trinucleotide preferences of ADAR and ADARB1. (b, f) The temporal changes were analyzed by one-way ANOVA, * p < 0.05, ** p < 0.01, *** p < 0.001, n.s. p ≧ 0.05 and N.D. not detected.
Supplementary Figure 3 Gene expression rhythms revealed by RNA-seq in Adarb1-knockout mice.
(a) Comparison between two biological replicates of transcript expression levels (FPKM) exhibited a high degree of reproducibility with an average R2 value of 0.98. (b) Overlap of rhythmic transcripts between control (black) and Adarb1-KO liver (yellow). A solid line circle indicates rhythmic transcripts with strong rhythmic expression, and a dotted circle indicates those with weak rhythmic expression. (c) Temporal expression profiles of Slcola4 and Aim1l in qRT-PCR analysis. The signals were normalized to Rps29 (mean ± SEM; n = 3). The temporal changes were analyzed by one-way ANOVA, * p < 0.05, ** p < 0.01 and n.s. p ≧ 0.05. (d) Temporal expression profiles of Cpeb2, Hmgxb4 and Gmeb1 in RNA-Seq analysis (Data are mean; n = 2). (e) Pie charts showing the enrichment of transcripts that were defined as arrhythmic at nascent RNA levels in the previous study. (f, g) Circadian expression profiles of typical clock genes in RNA-Seq analysis (f) and in qRT-PCR analysis (g) of control and Adarb1-KO mouse liver. In panel g, the signals were normalized to Rps29 (mean ± SEM; n = 3). (h) A heat map of rhythmically expressed genes both in control and Adarb1-KO mice (n = 659). Gene expression levels of two biological replicates across time points were shown in each lane corresponding to one gene, ordered by the phases. High expression levels were displayed in yellow and low in blue. (i) A histogram of acrophase (circadian peak phase) of the 659 rhythmic genes in both genotypes.
Supplementary Figure 4 Short-period phenotype of ADARB1 deficiency.
(a) Differences in circadian peak phase (Δacrophase) of the 659 rhythmic genes between control and Adarb1-KO mouse liver (*** p < 0.001 by Paired t-test). The acrophase in control was set to 0. (b) In NIH3T3 cells transiently transfected with an shRNA expression vector (control sh, Adarb1_sh1, Adarb1_sh2 or Adarb1_sh3), Adarb1 mRNA levels were examined by qRT-PCR. The signals were normalized to Rps29 (mean ± SEM; n = 3; ** p < 0.01 by Student's t-test). (c) Unsynchronized NIH3T3 cells were transiently transfected with a Flag-ADARB1 expression vector in combination with an shRNA expression vector (control sh, Adarb1_sh1, Adarb1_sh2 or Adarb1_sh3). The cell lysates were subjected to immunoblotting. ACTB served as a loading control. (d, e) A representative set of bioluminescence rhythms of Fig.4c (d) and 4d (e) was shown. (f) Circadian profiles of CRY1 (arrowhead), PER2, CLOCK and ARNTL proteins in control and Adarb1-KO mice liver nuclear extracts (mean ± SEM; n = 3; * p < 0.05 by Student's t-test). Nonspecific bands are indicated by asterisks. (g) Dual luciferase reporter assay in unsynchronized NIH3T3 cells transiently transfected with a luciferase-Cry2 3’-UTR reporter (nt 1-2177) in combination with a let-7g expression vector (mean ± SEM; n = 3; * p < 0.05, ** p < 0.01 and n.s. p ≧ 0.05 by Student's t-test). The relative luciferase activities were normalized to the signals from the cells transiently transfected with an empty pcDNA3.1/V5-His plasmid, and the mean value of the empty luciferase reporter was set to 1. (h) Circadian periods of synchronized NIH3T3 cells transiently transfected with a Arntl-luciferase reporter in combination with control sh, Adarb1 shRNA and let-7g expression vectors (mean ± SEM; n = 6; * p < 0.05, *** p < 0.001 by Student's t-test). (i) A model for the ADARB1-mediated regulations of the rhythmical A-to-I RNA editing events and global circadian outputs. The circadian expression of Adarb1 by CLOCK-dependent transactivation produces the rhythmic profiles of ADARB1-dependent A-to-I RNA editing. ADARB1 plays an essential role for the robust circadian oscillation by suppressing CRY2 expression through the regulation of miRNA biogenesis. ADARB1-mediated post-transcriptional regulation contributed to circadian profiles of steady-state mRNA levels and global circadian outputs.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–4 and Supplementary Table 4 (PDF 2328 kb)
Supplementary Table 1
List of the rhythmic genes that are common in four published papers4–7, and their CLOCK ChIP-seq scores modified from our previous study6 (XLSX 14 kb)
Supplementary Table 2
Number of sequenced tags (XLSX 14 kb)
Supplementary Table 3
List of the rhythmic editing sites (XLSX 150 kb)
Supplementary Table 5
Gene expression profiles in RNA-seq of control and Adarb1-knockout mice (XLSX 13250 kb)
Supplementary Table 6
List of primer sequences (XLSX 13 kb)
Rights and permissions
About this article
Cite this article
Terajima, H., Yoshitane, H., Ozaki, H. et al. ADARB1 catalyzes circadian A-to-I editing and regulates RNA rhythm. Nat Genet 49, 146–151 (2017). https://doi.org/10.1038/ng.3731
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3731
This article is cited by
-
A-to-I RNA editing shows dramatic up-regulation in osteosarcoma and broadly regulates tumor-related genes by altering microRNA target regions
Journal of Applied Genetics (2023)
-
Rhythmic transcription of Bmal1 stabilizes the circadian timekeeping system in mammals
Nature Communications (2022)
-
Circadian Regulation of GluA2 mRNA Processing in the Rat Suprachiasmatic Nucleus and Other Brain Structures
Molecular Neurobiology (2021)
-
Membrane and synaptic defects leading to neurodegeneration in Adar mutant Drosophila are rescued by increased autophagy
BMC Biology (2020)
-
Genome-wide profiling of RNA editing sites in sheep
Journal of Animal Science and Biotechnology (2019)