Abstract
Most disease-associated genetic variants are noncoding, making it challenging to design experiments to understand their functional consequences1,2. Identification of expression quantitative trait loci (eQTLs) has been a powerful approach to infer the downstream effects of disease-associated variants, but most of these variants remain unexplained3,4. The analysis of DNA methylation, a key component of the epigenome5,6, offers highly complementary data on the regulatory potential of genomic regions7,8. Here we show that disease-associated variants have widespread effects on DNA methylation in trans that likely reflect differential occupancy of trans binding sites by cis-regulated transcription factors. Using multiple omics data sets from 3,841 Dutch individuals, we identified 1,907 established trait-associated SNPs that affect the methylation levels of 10,141 different CpG sites in trans (false discovery rate (FDR) < 0.05). These included SNPs that affect both the expression of a nearby transcription factor (such as NFKB1, CTCF and NKX2-3) and methylation of its respective binding site across the genome. Trans methylation QTLs effectively expose the downstream effects of disease-associated variants.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Manolio, T.A. Genomewide association studies and assessment of the risk of disease. N. Engl. J. Med. 363, 166–176 (2010).
Visscher, P.M., Brown, M.A., McCarthy, M.I. & Yang, J. Five years of GWAS discovery. Am. J. Hum. Genet. 90, 7–24 (2012).
Westra, H.-J. et al. Systematic identification of trans eQTLs as putative drivers of known disease associations. Nat. Genet. 45, 1238–1243 (2013).
Wright, F.A. et al. Heritability and genomics of gene expression in peripheral blood. Nat. Genet. 46, 430–437 (2014).
Bernstein, B.E., Meissner, A. & Lander, E.S. The mammalian epigenome. Cell 128, 669–681 (2007).
Mill, J. & Heijmans, B.T. From promises to practical strategies in epigenetic epidemiology. Nat. Rev. Genet. 14, 585–594 (2013).
Gutierrez-Arcelus, M. et al. Passive and active DNA methylation and the interplay with genetic variation in gene regulation. eLife 2, e00523 (2013).
Tsankov, A.M. et al. Transcription factor binding dynamics during human ES cell differentiation. Nature 518, 344–349 (2015).
Tigchelaar, E.F. et al. Cohort profile: LifeLines DEEP, a prospective, general population cohort study in the northern Netherlands: study design and baseline characteristics. BMJ Open 5, e006772 (2015).
van Greevenbroek, M.M.J. et al. The cross-sectional association between insulin resistance and circulating complement C3 is partly explained by plasma alanine aminotransferase, independent of central obesity and general inflammation (the CODAM study). Eur. J. Clin. Invest. 41, 372–379 (2011).
Schoenmaker, M. et al. Evidence of genetic enrichment for exceptional survival using a family approach: the Leiden Longevity Study. Eur. J. Hum. Genet. 14, 79–84 (2006).
Willemsen, G. et al. The Adult Netherlands Twin Register: twenty-five years of survey and biological data collection. Twin Res. Hum. Genet. 16, 271–281 (2013).
Hofman, A. et al. The Rotterdam Study: 2014 objectives and design update. Eur. J. Epidemiol. 28, 889–926 (2013).
Hu, S. et al. DNA methylation presents distinct binding sites for human transcription factors. eLife 2, e00726 (2013).
Yao, C. et al. Integromic analysis of genetic variation and gene expression identifies networks for cardiovascular disease phenotypes. Circulation 131, 536–549 (2015).
Huan, T. et al. A meta-analysis of gene expression signatures of blood pressure and hypertension. PLoS Genet. 11, e1005035 (2015).
Lemire, M. et al. Long-range epigenetic regulation is conferred by genetic variation located at thousands of independent loci. Nat. Commun. 6, 6326 (2015).
Orrù, V. et al. Genetic variants regulating immune cell levels in health and disease. Cell 155, 242–256 (2013).
Roederer, M. et al. The genetic architecture of the human immune system: a bioresource for autoimmunity and disease pathogenesis. Cell 161, 387–403 (2015).
Rao, S.S.P. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Alipanahi, B., Delong, A., Weirauch, M.T. & Frey, B.J. Predicting the sequence specificities of DNA- and RNA-binding proteins by deep learning. Nat. Biotechnol. 33, 831–838 (2015).
Zuin, J. et al. Cohesin and CTCF differentially affect chromatin architecture and gene expression in human cells. Proc. Natl. Acad. Sci. USA 111, 996–1001 (2014).
Splinter, E. et al. CTCF mediates long-range chromatin looping and local histone modification in the β-globin locus. Genes Dev. 20, 2349–2354 (2006).
Jostins, L. et al. Host–microbe interactions have shaped the genetic architecture of inflammatory bowel disease. Nature 491, 119–124 (2012).
Pers, T.H. et al. Biological interpretation of genome-wide association studies using predicted gene functions. Nat. Commun. 6, 5890 (2015).
Kristiansson, K. et al. Genome-wide screen for metabolic syndrome susceptibility loci reveals strong lipid gene contribution but no evidence for common genetic basis for clustering of metabolic syndrome traits. Circ Cardiovasc Genet 5, 242–249 (2012).
Lettre, G. et al. Genome-wide association study of coronary heart disease and its risk factors in 8,090 African Americans: the NHLBI CARe Project. PLoS Genet. 7, e1001300 (2011).
Soranzo, N. et al. Meta-analysis of genome-wide scans for human adult stature identifies novel loci and associations with measures of skeletal frame size. PLoS Genet. 5, e1000445 (2009).
Filion, G.J.P. et al. A family of human zinc finger proteins that bind methylated DNA and repress transcription. Mol. Cell. Biol. 26, 169–181 (2006).
Sasai, N. & Defossez, P.A. Many paths to one goal? The proteins that recognize methylated DNA in eukaryotes. Int. J. Dev. Biol. 53, 323–334 (2009).
Shiraishi, K. et al. A genome-wide association study identifies two new susceptibility loci for lung adenocarcinoma in the Japanese population. Nat. Genet. 44, 900–903 (2012).
Qiu, Z. et al. Functional interactions between NURF and Ctcf regulate gene expression. Mol. Cell. Biol. 35, 224–237 (2015).
van Dam, R.M., Boer, J.M.A., Feskens, E.J.M. & Seidell, J.C. Parental history of diabetes modifies the association between abdominal adiposity and hyperglycemia. Diabetes Care 24, 1454–1459 (2001).
Scholtens, S. et al. Cohort profile: LifeLines, a three-generation cohort study and biobank. Int. J. Epidemiol. 44, 1172–1180 (2015).
Boomsma, D.I. et al. Netherlands Twin Register: a focus on longitudinal research. Twin Res. 5, 401–406 (2002).
Boomsma, D.I. et al. Genome-wide association of major depression: description of samples for the GAIN Major Depressive Disorder Study: NTR and NESDA biobank projects. Eur. J. Hum. Genet. 16, 335–342 (2008).
Deelen, J. et al. Genome-wide association meta-analysis of human longevity identifies a novel locus conferring survival beyond 90 years of age. Hum. Mol. Genet. 23, 4420–4432 (2014).
Anonymous. Credit for code. Nat. Genet. 46, 1 (2014).
Deelen, P. et al. Genotype harmonizer: automatic strand alignment and format conversion for genotype data integration. BMC Res. Notes 7, 901 (2014).
Howie, B.N., Donnelly, P. & Marchini, J. A flexible and accurate genotype imputation method for the next generation of genome-wide association studies. PLoS Genet. 5, e1000529 (2009).
Deelen, P. et al. Improved imputation quality of low-frequency and rare variants in European samples using the 'Genome of The Netherlands'. Eur. J. Hum. Genet. 22, 1321–1326 (2014).
Bonder, M.J. et al. Genetic and epigenetic regulation of gene expression in fetal and adult human livers. BMC Genomics 15, 860 (2014).
Touleimat, N. & Tost, J. Complete pipeline for Infinium(®) Human Methylation 450K BeadChip data processing using subset quantile normalization for accurate DNA methylation estimation. Epigenomics 4, 325–341 (2012).
Pidsley, R. et al. A data-driven approach to preprocessing Illumina 450K methylation array data. BMC Genomics 14, 293 (2013).
Westra, H.J. et al. MixupMapper: correcting sample mix-ups in genome-wide datasets increases power to detect small genetic effects. Bioinformatics 27, 2104–2111 (2011).
Fehrmann, R.S.N. et al. Trans-eQTLs reveal that independent genetic variants associated with a complex phenotype converge on intermediate genes, with a major role for the HLA. PLoS Genet. 7, e1002197 (2011).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet.journal 17, 10–12 (2011).
Dobin, A. et al. STAR: ultrafast universal RNA–seq aligner. Bioinformatics 29, 15–21 (2013).
Robinson, M.D. & Oshlack, A. A scaling normalization method for differential expression analysis of RNA–seq data. Genome Biol. 11, R25 (2010).
Zhernakova, D.V. et al. Identification of context-dependent expression quantitative trait loci in whole blood. Nat. Genet. http://dx.doi.org/10.1038/ng.3737 (2016).
Flicek, P. et al. Ensembl 2013. Nucleic Acids Res. 41, D48–D55 (2013).
Kent, W.J., Sugnet, C.W., Furey, T.S. & Roskin, K.M. The Human Genome Browser at UCSC. Genome Res. 12, 996–1006 (2002).
Kundaje, A. et al. Integrative analysis of 111 reference human epigenomes. Nature 518, 317–330 (2015).
Heinz, S. et al. Effect of natural genetic variation on enhancer selection and function. Nature 503, 487–492 (2013).
Hofmann, M. & Klinkenberg, R. Rapid Miner Data Mining Use Cases and Business Analytics Applications (Chapman & Hall/CRC, 2013).
de Ronde, J.J., Bonder, M.J., Lips, E.H., Rodenhuis, S. & Wessels, L.F.A. Breast cancer subtype specific classifiers of response to neoadjuvant chemotherapy do not outperform classifiers trained on all subtypes. PLoS One 9, e88551 (2014).
Acknowledgements
This work was performed within the framework of the Biobank-Based Integrative Omics Studies (BIOS) consortium funded by BBMRI-NL, a research infrastructure financed by the Dutch government (NWO 184.021.007). Samples were contributed by LifeLines, the Leiden Longevity Study, the Netherlands Twin Registry (NTR), the Rotterdam Study, the Genetic Research in Isolated Populations program, the CODAM study and the PAN study. We thank the participants of all aforementioned biobanks and acknowledge the contributions of the investigators to this study (Supplementary Note). This work was carried out on the Dutch national e-infrastructure with the support of SURF Cooperative. L.F. is supported by a grant from the Dutch Research Council (ZonMW-VIDI 917.14.374) and is supported by FP7/2007–2013, grant agreement 259867 and by an ERC Starting Grant, grant agreement 637640 (ImmRisk).
Author information
Authors and Affiliations
Consortia
Contributions
B.T.H., P.A.C.'t.H., J.B.J.v.M., A.I., R.J. and L.F. formed the management team of the BIOS consortium. D.I.B., R.P., J.v.D., J.J.H., M.M.J.v.G., C.D.A.S., C.J.H.v.d.K., C.G.S., C.W., L.F., A.Z., E.F.T., P.E.S., M.B., J.D., D.v.H., J.H.V., L.H.v.d.B., C.M.v.D., A.H., A.I. and A.G.U. managed and organized the biobanks. J.B.J.v.M., P.M.J., M. Verbiest, H.E.D.S., M. Verkerk, R.v.d.B., J.v.R. and N.L. generated RNA–seq and Illumina 450K data. H.M., M.v.I., M.v.G., J.B., D.V.Z., R.J., P.v.'t.H., P.D., I.N., P.A.C.'t.H., B.T.H. and M.M. were responsible for data management and the computational infrastructure. M.J.B., R.L., M. Vermaat, D.V.Z., R.C.S., I.J., M.v.I., P.D., F.v.D., M.v.G., W.A., S.M.K., M.A.S., E.W.v.Z., Y.L., M.L., T.J.H., R.J., P.A.C.'t.H., L.F. and B.T.H. performed the data analysis. M.J.B., R.L., L.F. and B.T.H. drafted the manuscript. D.V.Z., M.M., P.D. and M. Vermaat contributed equally. A.I., R.J. and J.B.J.v.M. contributed equally.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
A full list of members and affiliations appears in the Supplementary Note.
Integrated supplementary information
Supplementary Figure 1 Density of distances between CpG sites and the most strongly associated meQTL SNP.
Density plot of the distances between the 139,566 CpGs harboring a cis-meQTL and the most strongly associated SNP. Most SNP–CpG pairs are in close proximity (median distance = 10 kb), as indicated by the narrow peak around zero.
Supplementary Figure 2 Relationship between methylation variation and meQTL-associated CpGs.
The proportion of CpGs harboring an identified trans-meQTL increases with increasing variability in DNA methylation. The proportion of CpGs with evidence of a trans-meQTL is calculated per decile of variability in methylation (x axis).
Supplementary Figure 3 Characterization of cis-meQTLs.
(a) The number of cis-meQTLs found is strongly dependent on the variability in DNA methylation at a CpG site. Variances for 405,709 CpGs interrogated in the analyses were calculated using the 3,841 samples for which 450K data were available. Next, the CpGs were divided into deciles and the number of effects was counted for each decile. The different stacked colors correspond to primary, secondary, etc., effects. (b) The proportion of variance explained remains limited, even for highly variable CpG sites. The x axis shows the variances calculated for the 405,709 CpGs interrogated. The y axis shows the proportion of that variance explained by our identified cis-meQTLs. The limited proportion of variance explained, even for highly variable probes, suggests that increased statistical power contributes to but does not fully explain the increased number of cis-meQTLs identified. (c) DNA methylation variability differs across genomic contexts. Each line represents the proportion of the 405,709 CpGs used present in each genomic region. This clearly shows that some CpGs on the array are over-represented in certain genomic contexts. For example, weakly variable CpGs (0–10%) are over-represented in CpG islands. This may confound any enrichment analyses if variability in DNA methylation is influencing the likelihood of a given CpG harboring a meQTL. (d) DNA methylation variability seems to be the driving factor for identifying cis-meQTLs, even within genomic contexts. Each line again represents a distinct genomic context. (e) Reported enrichments of cis-meQTL effects for certain genomic contexts are strongly attenuated after accounting for the differential variability in DNA methylation between those genomic regions. Gray bars show uncorrected odds ratios. Blue bars show odds ratios corrected for methylation variability and the distance to the nearest SNP.
Supplementary Figure 4 Characterization of cis-eQTMs in relation to the direction of the eQTM effect.
Over-representation of positive (blue bars) and negative (red bars) e-CpGs in CpG islands and predicted chromatin states. The x axis shows this over-representation in terms of odds ratios and error bars (95% confidence intervals). e-CpGs with negative associations are over-represented in active regions (for example, active TSSs and enhancers), whereas e-CpGs with positive associations are often found in repressed regions (for example, quiescent regions). CGI, CpG island; TssA, active TSS; TssAFlnk, flanking active TSS; TxFlnk, transcribed at gene 5′ or 3′ end; Tx, strong transcription; TxWk, weak transcription; EnhG, genic enhancer; Enh, enhancer; ZNF/Rpts, ZNF genes and repeats; Het, heterochromatin; TssBiv, bivalent/poised TSS; BivFlnk, flanking bivalent TSS/enhancer; EnhBiv: bivalent enhancer.
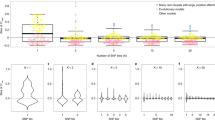
Supplementary Figure 5 Trans-meQTLs identified for a risk factor for inflammatory bowel disease, rs11190140, and the overlap with NKX2-3.
(a) Depiction of the NKX2-3 gene and rs11190140, associated with inflammatory bowel disease. The plot shows increased expression of NKX2-3 for the T risk allele. (b) In addition to influencing NKX2-3 expression, rs11190140 also influences DNA methylation at 228 CpGs in trans, decreasing methylation levels at 81.1% of the affected CpG sites (red). In addition, many of the CpG sites overlap with NKX2-1 and NKX2-5 motifs (there is no NKX2-3 motif or ChIP–seq data available). (c) Gene network of the genes associated with 15 of the 228 CpGs (6.6%) with a trans-meQTL: blue, cis-eQTL-affected gene; red, genes associated both in methylation and expression.
Supplementary Figure 6 Trans-meQTLs identified for a risk factor for height, rs6763931, and the overlap with ZBTB38.
(a) Depiction of the ZBTB38 gene and rs6763931, associated with height. The plot shows increased expression of ZBTB38 for the T risk allele. (b) In addition to influencing ZBTB38 expression, rs6763931 also influences DNA methylation at 267 CpGs in trans, decreasing methylation levels at 99.2% of the affected CpG sites (red). In addition, depletion of overlap with H3K27me3 is observed (7.4-fold depletion, P = 3.8 × 10–28), shown in the outer chart. (c) Gene network of the genes associated with 60 of the 779 CpGs (7.7%) with a trans-meQTL: blue, cis-eQTL effected gene; red, genes associated both in methylation and expression.
Supplementary Figure 7 Trans-meQTLs identified for a risk factor related to lung carcinoma, rs7216064, and overlap with BPTF.
(a) Depiction of the BPTF gene and rs7216064, associated with lung carcinoma. (b) rs7216064 influences DNA methylation at 64 CpGs in trans, decreasing methylation levels at 82.8% of the affected CpG sites (red). In addition, many of the CpG sites (81.3%) overlap with CTCF-binding sites (16.8-fold enrichment, P = 5.1 × 10–25), shown in the outer chart.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and Supplementary Note. (PDF 1911 kb)
Supplementary Table 1
Descriptions and number of samples per cohort. (XLSX 8 kb)
Supplementary Table 2
Number of independent cis-meQTLs per QTL mapping round. (XLSX 8 kb)
Supplementary Table 3
GWAS SNPs tested for trans-meQTLs. (XLSX 90 kb)
Supplementary Table 4
Replication of lymphocyte trans-meQTLs in blood and vice versa. (XLSX 1434 kb)
Supplementary Table 5
Results of trans-meQTLs in non-corrected data. (XLSX 651 kb)
Supplementary Table 6
Results of trans-meQTLs in data corrected for blood cell composition. (XLSX 722 kb)
Supplementary Table 7
Results of trans-meQTL mapping on SNPs related to blood cell composition. (XLSX 162 kb)
Supplementary Table 8
Trans-meQTL effects replicated in expression. (XLSX 350 kb)
Rights and permissions
About this article
Cite this article
Bonder, M., Luijk, R., Zhernakova, D. et al. Disease variants alter transcription factor levels and methylation of their binding sites. Nat Genet 49, 131–138 (2017). https://doi.org/10.1038/ng.3721
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3721
This article is cited by
-
DNA methylation may partly explain psychotropic drug-induced metabolic side effects: results from a prospective 1-month observational study
Clinical Epigenetics (2024)
-
Epigenome-wide association study of dietary fatty acid intake
Clinical Epigenetics (2024)
-
Kidney-specific methylation patterns correlate with kidney function and are lost upon kidney disease progression
Clinical Epigenetics (2024)
-
Epigenomic insights into common human disease pathology
Cellular and Molecular Life Sciences (2024)
-
Genetic control of DNA methylation is largely shared across European and East Asian populations
Nature Communications (2024)