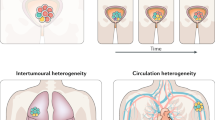
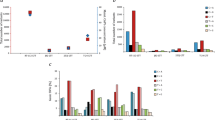
Abstract
Chemotherapy-resistant urothelial carcinoma has no uniformly curative therapy. Understanding how selective pressure from chemotherapy directs the evolution of urothelial carcinoma and shapes its clonal architecture is a central biological question with clinical implications. To address this question, we performed whole-exome sequencing and clonality analysis of 72 urothelial carcinoma samples, including 16 matched sets of primary and advanced tumors prospectively collected before and after chemotherapy. Our analysis provided several insights: (i) chemotherapy-treated urothelial carcinoma is characterized by intra-patient mutational heterogeneity, and the majority of mutations are not shared; (ii) both branching evolution and metastatic spread are very early events in the natural history of urothelial carcinoma; (iii) chemotherapy-treated urothelial carcinoma is enriched with clonal mutations involving L1 cell adhesion molecule (L1CAM) and integrin signaling pathways; and (iv) APOBEC-induced mutagenesis is clonally enriched in chemotherapy-treated urothelial carcinoma and continues to shape the evolution of urothelial carcinoma throughout its lifetime.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Howlader, N. et al. SEER Cancer Statistics Review, 1975–2010 (National Cancer Institute, Bethesda, Maryland, USA, 2013).
von der Maase, H. et al. Long-term survival results of a randomized trial comparing gemcitabine plus cisplatin, with methotrexate, vinblastine, doxorubicin, plus cisplatin in patients with bladder cancer. J. Clin. Oncol. 23, 4602–4608 (2005).
Clark, P.E. et al. Bladder cancer. J. Natl. Compr. Canc. Netw. 11, 446–475 (2013).
Sternberg, C.N. et al. Randomized phase III trial of high-dose-intensity methotrexate, vinblastine, doxorubicin, and cisplatin (MVAC) chemotherapy and recombinant human granulocyte colony-stimulating factor versus classic MVAC in advanced urothelial tract tumors: European Organization for Research and Treatment of Cancer Protocol no. 30924. J. Clin. Oncol. 19, 2638–2646 (2001).
Sternberg, C.N. et al. Seven year update of an EORTC phase III trial of high-dose intensity M-VAC chemotherapy and G-CSF versus classic M-VAC in advanced urothelial tract tumours. Eur. J. Cancer 42, 50–54 (2006).
Bellmunt, J. et al. Phase III trial of vinflunine plus best supportive care compared with best supportive care alone after a platinum-containing regimen in patients with advanced transitional cell carcinoma of the urothelial tract. J. Clin. Oncol. 27, 4454–4461 (2009).
Bellmunt, J. & Petrylak, D.P. New therapeutic challenges in advanced bladder cancer. Semin. Oncol. 39, 598–607 (2012).
Cancer Genome Atlas Research Network. Comprehensive molecular characterization of urothelial bladder carcinoma. Nature 507, 315–322 (2014).
Prandi, D. et al. Unraveling the clonal hierarchy of somatic genomic aberrations. Genome Biol. 15, 439 (2014).
Nixon, K.C. The parsimony ratchet, a new method for rapid parsimony analysis. Cladistics 15, 407–414 (1999).
Zheng, Y. et al. Identification of Pold2 as a novel interaction partner of protein inhibitor of activated STAT2. Int. J. Mol. Med. 30, 884–888 (2012).
Lee, Y.S., Gregory, M.T. & Yang, W. Human Pol ζ purified with accessory subunits is active in translesion DNA synthesis and complements Pol η in cisplatin bypass. Proc. Natl. Acad. Sci. USA 111, 2954–2959 (2014).
van Boxtel, R. et al. FOXP1 acts through a negative feedback loop to suppress FOXO-induced apoptosis. Cell Death Differ. 20, 1219–1229 (2013).
Choi, E.J. et al. FOXP1 functions as an oncogene in promoting cancer stem cell-like characteristics in ovarian cancer cells. Oncotarget 7, 3506–3519 (2016).
Greco, C. et al. E-cadherin/p120-catenin and tetraspanin Co-029 cooperate for cell motility control in human colon carcinoma. Cancer Res. 70, 7674–7683 (2010).
Wei, L., Li, Y. & Suo, Z. TSPAN8 promotes gastric cancer growth and metastasis via ERK MAPK pathway. Int. J. Clin. Exp. Med. 8, 8599–8607 (2015).
Zöller, M. Tetraspanins: push and pull in suppressing and promoting metastasis. Nat. Rev. Cancer 9, 40–55 (2009).
Kinsella, R.J. et al. Ensembl BioMarts: a hub for data retrieval across taxonomic space. Database (Oxford) 2011, bar030 (2011).
Plimack, E.R. et al. Defects in DNA repair genes predict response to neoadjuvant cisplatin-based chemotherapy in muscle-invasive bladder cancer. Eur. Urol. 68, 959–967 (2015).
Eke, I. & Cordes, N. Focal adhesion signaling and therapy resistance in cancer. Semin. Cancer Biol. 31, 65–75 (2015).
Maiuthed, A. & Chanvorachote, P. Cisplatin at sub-toxic levels mediates integrin switch in lung cancer cells. Anticancer Res. 34, 7111–7117 (2014).
Seguin, L., Desgrosellier, J.S., Weis, S.M. & Cheresh, D.A. Integrins and cancer: regulators of cancer stemness, metastasis, and drug resistance. Trends Cell Biol. 25, 234–240 (2015).
Tsutsumi, S. et al. L1 cell adhesion molecule (L1CAM) expression at the cancer invasive front is a novel prognostic marker of pancreatic ductal adenocarcinoma. J. Surg. Oncol. 103, 669–673 (2011).
Weidle, U.H., Eggle, D. & Klostermann, S. L1-CAM as a target for treatment of cancer with monoclonal antibodies. Anticancer Res. 29, 4919–4931 (2009).
Meier, B. et al. C. elegans whole-genome sequencing reveals mutational signatures related to carcinogens and DNA repair deficiency. Genome Res. 24, 1624–1636 (2014).
Szikriszt, B. et al. A comprehensive survey of the mutagenic impact of common cancer cytotoxics. Genome Biol. 17, 99 (2016).
Alexandrov, L.B. et al. Signatures of mutational processes in human cancer. Nature 500, 415–421 (2013).
Nik-Zainal, S. et al. Mutational processes molding the genomes of 21 breast cancers. Cell 149, 979–993 (2012).
Kim, J. et al. Somatic ERCC2 mutations are associated with a distinct genomic signature in urothelial tumors. Nat. Genet. 48, 600–606 (2016).
Roberts, S.A. et al. An APOBEC cytidine deaminase mutagenesis pattern is widespread in human cancers. Nat. Genet. 45, 970–976 (2013).
Liu, D. et al. Clinical validation of chemotherapy response biomarker ERCC2 in muscle-invasive urothelial bladder carcinoma. JAMA Oncol. 2, 1094–1096 (2016).
Van Allen, E.M. et al. Somatic ERCC2 mutations correlate with cisplatin sensitivity in muscle-invasive urothelial carcinoma. Cancer Discov. 4, 1140–1153 (2014).
Hoopes, J.I. et al. APOBEC3A and APOBEC3B preferentially deaminate the lagging strand template during DNA replication. Cell Rep. 14, 1273–1282 (2016).
Lackey, L. et al. APOBEC3B and AID have similar nuclear import mechanisms. J. Mol. Biol. 419, 301–314 (2012).
Chelico, L., Pham, P. & Goodman, M.F. Stochastic properties of processive cytidine DNA deaminases AID and APOBEC3G. Phil. Trans. R. Soc. Lond. B 364, 583–593 (2009).
Harris, R.S. & Liddament, M.T. Retroviral restriction by APOBEC proteins. Nat. Rev. Immunol. 4, 868–877 (2004).
Faltas, B.M., Karir, B.S., Tagawa, S.T. & Rosenberg, J.E. Novel molecular targets for urothelial carcinoma. Expert Opin. Ther. Targets 19, 515–525 (2015).
Knowles, M.A. & Hurst, C.D. Molecular biology of bladder cancer: new insights into pathogenesis and clinical diversity. Nat. Rev. Cancer 15, 25–41 (2015).
Yoon, H. et al. L1 cell adhesion molecule and epidermal growth factor receptor activation confer cisplatin resistance in intrahepatic cholangiocarcinoma cells. Cancer Lett. 316, 70–76 (2012).
Stoeck, A. et al. L1-CAM in a membrane-bound or soluble form augments protection from apoptosis in ovarian carcinoma cells. Gynecol. Oncol. 104, 461–469 (2007).
Sebens Müerköster, S. et al. Drug-induced expression of the cellular adhesion molecule L1CAM confers anti-apoptotic protection and chemoresistance in pancreatic ductal adenocarcinoma cells. Oncogene 26, 2759–2768 (2007).
Sebens Müerköster, S. et al. alpha5-integrin is crucial for L1CAM-mediated chemoresistance in pancreatic adenocarcinoma. Int. J. Oncol. 34, 243–253 (2009).
Kiefel, H. et al. L1CAM: a major driver for tumor cell invasion and motility. Cell Adh. Migr. 6, 374–384 (2012).
Gast, D. et al. The RGD integrin binding site in human L1-CAM is important for nuclear signaling. Exp. Cell Res. 314, 2411–2418 (2008).
Voura, E.B., Ramjeesingh, R.A., Montgomery, A.M. & Siu, C.H. Involvement of integrin alpha(v)beta(3) and cell adhesion molecule L1 in transendothelial migration of melanoma cells. Mol. Biol. Cell 12, 2699–2710 (2001).
Burgett, M.E. et al. Direct contact with perivascular tumor cells enhances integrin αvβ3 signaling and migration of endothelial cells. Oncotarget http://dx.doi.org/10.18632/oncotarget.9700 (2016).
Hodkinson, P.S. et al. ECM overrides DNA damage-induced cell cycle arrest and apoptosis in small-cell lung cancer cells through beta1 integrin-dependent activation of PI3-kinase. Cell Death Differ. 13, 1776–1788 (2006).
Sethi, T. et al. Extracellular matrix proteins protect small cell lung cancer cells against apoptosis: a mechanism for small cell lung cancer growth and drug resistance in vivo. Nat. Med. 5, 662–668 (1999).
Zhang, H. et al. Beta 1-integrin protects hepatoma cells from chemotherapy induced apoptosis via a mitogen-activated protein kinase dependent pathway. Cancer 95, 896–906 (2002).
Argyris, E.G. et al. The interferon-induced expression of APOBEC3G in human blood-brain barrier exerts a potent intrinsic immunity to block HIV-1 entry to central nervous system. Virology 367, 440–451 (2007).
Shain, K.H. & Dalton, W.S. Cell adhesion is a key determinant in de novo multidrug resistance (MDR): new targets for the prevention of acquired MDR. Mol. Cancer Ther. 1, 69–78 (2001).
Damiano, J.S. Integrins as novel drug targets for overcoming innate drug resistance. Curr. Cancer Drug Targets 2, 37–43 (2002).
Hazlehurst, L.A. & Dalton, W.S. Mechanisms associated with cell adhesion mediated drug resistance (CAM-DR) in hematopoietic malignancies. Cancer Metastasis Rev. 20, 43–50 (2001).
Hehlgans, S., Haase, M. & Cordes, N. Signalling via integrins: implications for cell survival and anticancer strategies. Biochim. Biophys. Acta 1775, 163–180 (2007).
Cho, S. et al. Generation, characterization and preclinical studies of a human anti-L1CAM monoclonal antibody that cross-reacts with rodent L1CAM. MAbs 8, 414–425 (2016).
Min, J.K. et al. L1 cell adhesion molecule is a novel therapeutic target in intrahepatic cholangiocarcinoma. Clin. Cancer Res. 16, 3571–3580 (2010).
Schäfer, H. et al. Combined treatment of L1CAM antibodies and cytostatic drugs improve the therapeutic response of pancreatic and ovarian carcinoma. Cancer Lett. 319, 66–82 (2012).
Golubovskaya, V.M. Targeting FAK in human cancer: from finding to first clinical trials. Front. Biosci. (Landmark Ed.) 19, 687–706 (2014).
Schultze, A. & Fiedler, W. Therapeutic potential and limitations of new FAK inhibitors in the treatment of cancer. Expert Opin. Investig. Drugs 19, 777–788 (2010).
Rezaee, M., Sanche, L. & Hunting, D.J. Cisplatin enhances the formation of DNA single- and double-strand breaks by hydrated electrons and hydroxyl radicals. Radiat. Res. 179, 323–331 (2013).
Zellweger, R. et al. Rad51-mediated replication fork reversal is a global response to genotoxic treatments in human cells. J. Cell Biol. 208, 563–579 (2015).
Roberts, S.A. & Gordenin, D.A. Hypermutation in human cancer genomes: footprints and mechanisms. Nat. Rev. Cancer 14, 786–800 (2014).
Chan, K. et al. An APOBEC3A hypermutation signature is distinguishable from the signature of background mutagenesis by APOBEC3B in human cancers. Nat. Genet. 47, 1067–1072 (2015).
Lipinski, K.A. et al. Cancer evolution and the limits of predictability in precision cancer medicine. Trends Cancer 2, 49–63 (2016).
Beltran, H. et al. Whole-exome sequencing of metastatic cancer and iomarkers of treatment response. JAMA Oncol. 1, 466–474 (2015).
Demichelis, F. et al. SNP panel identification assay (SPIA): a genetic-based assay for the identification of cell lines. Nucleic Acids Res. 36, 2446–2456 (2008).
Beltran, H. et al. Divergent clonal evolution of castration-resistant neuroendocrine prostate cancer. Nat. Med. 22, 298–305 (2016).
Futreal, P.A. et al. A census of human cancer genes. Nat. Rev. Cancer 4, 177–183 (2004).
Rubio-Perez, C. et al. In silico prescription of anticancer drugs to cohorts of 28 tumor types reveals targeting opportunities. Cancer Cell 27, 382–396 (2015).
Cibulskis, K. et al. Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nat. Biotechnol. 31, 213–219 (2013).
Jiang, Y., Soong, T.D., Wang, L., Melnick, A.M. & Elemento, O. Genome-wide detection of genes targeted by non-Ig somatic hypermutation in lymphoma. PLoS One 7, e40332 (2012).
Romanel, A., Lago, S., Prandi, D., Sboner, A. & Demichelis, F. ASEQ: fast allele-specific studies from next-generation sequencing data. BMC Med. Genomics 8, 9 (2015).
Ramos, A.H. et al. Oncotator: cancer variant annotation tool. Hum. Mutat. 36, E2423–E2429 (2015).
Eilbeck, K. et al. The sequence ontology: a tool for the unification of genome annotations. Genome Biol. 6, R44 (2005).
Acknowledgements
We would like to thank our patients and their families for participation in this study. We would like to thank B. Sleckman for constructive review of the manuscript. We would also like to acknowledge D. Scherr and C. Barbieri for contributing samples, our research and clinical pathology fellows J. Fontugne, M. Kossai, C. Pauli, K. Hennrick and K. Park for their assistance during rapid autopsies, and S.S. Chae and D. Wilkes for technical assistance and constructive comments. We would also like to thank T.Y. MacDonald, J. Padilla and T. Fedrizzi for technical assistance. Work was partially supported by the Translational Research Program at WCMC Pathology and Laboratory Medicine. This work was supported by a Conquer Cancer Foundation and the John and Elizabeth Leonard Family Foundation Young Investigator Award (B.M.F.), NIH/NCATS KL2TR000458 (B.M.F.), Early Detection Research Network US NCI 5U01 CA111275-09 (J.M.M., M.A.R. and F.D.), Damon Runyon Cancer Research Foundation Clinical Investigator Award CI-67-13 (H.B.), and H2020 European Research Council ERC-CoG 648670 (F.D.).
Author information
Authors and Affiliations
Contributions
Initiation and design of study: B.M.F., H.B., M.A.R. and F.D. Subject enrollment and sample and clinical data collection: B.M.F., H.B., D.M.N., S.T.T., A.M.M., C.S., J.M.M. and B.R. Statistical and bioinformatics analyses: D.P., O.E., A.S. and F.D. Supervision of research: B.M.F., M.A.R., J.R. and F.D. Writing of the first draft of the manuscript: B.M.F., D.P., H.B., M.A.R. and F.D. All authors contributed to the writing and editing of the revised manuscript and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
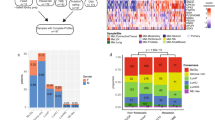
Supplementary Figure 1 Analysis of mutated genes as reported in TCGA bladder study.
Each row represents a gene harboring frequent SNVs (middle panel) or somatic copy number (bottom panel). Annotation rows include, from top to bottom, tumor ploidy, treatment information, biopsy site, patient’s gender, matched samples. Top bars report the otal number of non-silent SNVs in each sample. Bars on the left indicate the per-sample frequency of aberration in the cohort. NA, not available.
Supplementary Figure 2 SNVs burden comparison.
Per sample number of all SNVs (left boxplot) and only non-silent SNVs (right boxplot) in pre-chemotherapy and post-chemotherapy tumors. No statistical difference.
Supplementary Figure 3 Single nucleotide variants (SNV) validation by targeted sequencing (N250 panel).
Each dot represents a genomic position corresponding to an SNV sequenced with WES and with N250 approaches. Reported values are the ratios between the number of reads supporting the alternative base and the coverage. SNVs from the same patient are represented with the same color.
Supplementary Figure 4 Discordance in the SNVs between chemotherapy-naive and chemotherapy-treated UC tumors.
(a) Percentage of shared and unique SNVs in matched chemotherapy-naive and chemotherapy-treated UC tumors (same of Fig. 2a). (b) Detail of the specific amino acid change in matched chemotherapy-naive and chemotherapy-treated UC tumors. Each column represents a matched pair of pre- and post- chemotherapy tumors. Row reports the specific amino acid change. The figure highlights the wide divergence in the mutational landscape of pre- and post-chemotherapy samples.
Supplementary Figure 5 Phylogenetic trees of patients with two sequenced tumor samples.
Phylogenetic trees from 15 patients with two samples per patient.
Supplementary Figure 6 Hierarchical clusters of 131 UC tumor samples by copy-number alterations.
The plot extends Fig. 5a considering samples from both WCM and TCGA cohorts. Copy number gains are represented in red and copy number losses are represented in blue. Each column represents one tumor sample. Clinical annotations include treatment, biopsy site, cohort, and the presence of TP53 SNV.
Supplementary Figure 7 Hierarchical clusters of 86 UC tumor samples by copy-number alterations.
The plot shows allele specific copy number clusters as in Fig. 5a while considering samples from the TCGA cohort. Copy number gains are represented in red and copy number losses are represented in blue. Each column represents one tumor sample. Each row represents one of the 2503 genes used to compute TCGA clusters (Fig. S4.1 from Nature 507, 315–322, 2014)). Annotations include presence of TP53 SNV and TCGA clusters.
Supplementary Figure 8 Pairwise comparison of SNVs clonality.
Top right scatter plot reports the clonality of SNVs in matched pre-chemotherapy and post-chemotherapy samples. Bottom and left boxplots show clonality of SNVs that are private to pre-chemotherapy and post-chemotherapy samples, respectively.
Supplementary Figure 9 Comparison of FFPE and Fresh samples.
(left) Boxplot of the purity in FFPE and Fresh samples (p>0.05, Wilcoxon test). (right) Boxplot of the number of non-silent SNVs in FFPE and Fresh samples (p>0.05, Wilcoxon test).
Supplementary Figure 10 Landscape of somatic SNVs identified.
Landscape of somatic SNVs identified.
Supplementary Figure 11 Comparison of shared and local SNVs considering silent and non-silent SNVs.
Comparison of the percentage of shared and local SNVs in matched chemotherapy-naive and chemotherapy-treated UC tumors when considering only non-silent SNVs (top) or all SNVs (bottom).
Supplementary Figure 12 Early branching evolution in UC using silent and non-silent SNVs.
Phylogenetic trees (top), shared and private clonally-adjusted mutations (bottom) from 6 patients with three or more tumor samples per patient.
Supplementary Figure 13 Phylogenetic tree of case WCM117 using silent and non-silent SNVs.
Silent and non-silent SNVs across 12 tumor samples (top) collected at various time points and from various anatomical locations. Fractions of tumor cells harboring each mutation represented by shades of green. Reconstruction of evolutionary tree (bottom).
Supplementary Figure 14 Clonal enrichment of mutations in chemotherapy-treated UC considering silent and non-silent SNVs.
Clonal enrichment of mutations in chemotherapy-treated UC considering silent and non-silent SNVs.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–14 and Supplementary Note (PDF 3323 kb)
Supplementary Tables 1–11
Supplementary Tables 1–11 (XLSX 5139 kb)
Rights and permissions
About this article
Cite this article
Faltas, B., Prandi, D., Tagawa, S. et al. Clonal evolution of chemotherapy-resistant urothelial carcinoma. Nat Genet 48, 1490–1499 (2016). https://doi.org/10.1038/ng.3692
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3692
This article is cited by
-
Establishment and characterization of multiple patient-derived organoids from a case of advanced endometrial cancer
Human Cell (2024)
-
ARHGAP6 inhibits bladder cancer cell viability, migration, and invasion via β-catenin signaling and enhances mitomycin C sensitivity
Human Cell (2023)
-
The evolution of non-small cell lung cancer metastases in TRACERx
Nature (2023)
-
Stepwise molecular mechanisms responsible for chemoresistance in bladder cancer cells
Cell Death Discovery (2022)
-
Serial ctDNA analysis predicts clinical progression in patients with advanced urothelial carcinoma
British Journal of Cancer (2022)