Abstract
Meiotic recombination involves a combination of gene conversion and crossover events that, along with mutations, produce germline genetic diversity. Here we report the discovery of 3,176 SNP and 61 indel gene conversions. Our estimate of the non-crossover (NCO) gene conversion rate (G) is 7.0 for SNPs and 5.8 for indels per megabase per generation, and the GC bias is 67.6%. For indels, we demonstrate a 65.6% preference for the shorter allele. NCO gene conversions from mothers are longer than those from fathers, and G is 2.17 times greater in mothers. Notably, G increases with the age of mothers, but not the age of fathers. A disproportionate number of NCO gene conversions in older mothers occur outside double-strand break (DSB) regions and in regions with relatively low GC content. This points to age-related changes in the mechanisms of meiotic gene conversion in oocytes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
20 September 2018
In the version of this article published, statements about the impact of insertions and deletions on gene conversions were incorrect. We reported a bias toward deletions, whereas in fact the bias was toward insertions. We are deeply indebted to Laurent Duret and Brice Letcher for noticing this mistake in our manuscript. Full details can be found in the correction notice.
References
Sun, H., Treco, D., Schultes, N.P. & Szostak, J.W. Double-strand breaks at an initiation site for meiotic gene conversion. Nature 338, 87–90 (1989).
Lam, I. & Keeney, S. Mechanism and regulation of meiotic recombination initiation. Cold Spring Harb. Perspect. Biol. 7, a016634 (2014).
Baudat, F. et al. PRDM9 is a major determinant of meiotic recombination hotspots in humans and mice. Science 327, 836–840 (2010).
Haber, J. Genome Stability (Garland Science, 2013).
McMahill, M.S., Sham, C.W. & Bishop, D.K. Synthesis-dependent strand annealing in meiosis. PLoS Biol. 5, e299 (2007).
Jeffreys, A.J. & May, C.A. Intense and highly localized gene conversion activity in human meiotic crossover hot spots. Nat. Genet. 36, 151–156 (2004).
Odenthal-Hesse, L., Berg, I.L., Veselis, A., Jeffreys, A.J. & May, C.A. Transmission distortion affecting human noncrossover but not crossover recombination: a hidden source of meiotic drive. PLoS Genet. 10, e1004106 (2014).
Cole, F. et al. Mouse tetrad analysis provides insights into recombination mechanisms and hotspot evolutionary dynamics. Nat. Genet. 46, 1072–1080 (2014).
Allers, T. & Lichten, M. Differential timing and control of noncrossover and crossover recombination during meiosis. Cell 106, 47–57 (2001).
Szostak, J.W., Orr-Weaver, T.L., Rothstein, R.J. & Stahl, F.W. The double-strand-break repair model for recombination. Cell 33, 25–35 (1983).
Galtier, N., Piganeau, G., Mouchiroud, D. & Duret, L. GC-content evolution in mammalian genomes: the biased gene conversion hypothesis. Genetics 159, 907–911 (2001).
Glémin, S. et al. Quantification of GC-biased gene conversion in the human genome. Genome Res. 25, 1215–1228 (2015).
Duret, L. & Galtier, N. Biased gene conversion and the evolution of mammalian genomic landscapes. Annu. Rev. Genomics Hum. Genet. 10, 285–311 (2009).
Narasimhan, V.M. et al. A direct multi-generational estimate of the human mutation rate from autozygous segments seen in thousands of parentally related individuals. Preprint at bioRxiv http://dx.doi.org/10.1101/059436 (2016).
Palamara, P.F. et al. Leveraging distant relatedness to quantify human mutation and gene-conversion rates. Am. J. Hum. Genet. 97, 775–789 (2015).
Lachance, J. & Tishkoff, S.A. Biased gene conversion skews allele frequencies in human populations, increasing the disease burden of recessive alleles. Am. J. Hum. Genet. 95, 408–420 (2014).
Kong, A. et al. Fine-scale recombination rate differences between sexes, populations and individuals. Nature 467, 1099–1103 (2010).
Kong, A. et al. A high-resolution recombination map of the human genome. Nat. Genet. 31, 241–247 (2002).
Kong, A. et al. Recombination rate and reproductive success in humans. Nat. Genet. 36, 1203–1206 (2004).
Williams, A.L. et al. Non-crossover gene conversions show strong GC bias and unexpected clustering in humans. eLife 4, e04637 (2015).
Guillon, H., Baudat, F., Grey, C., Liskay, R.M. & de Massy, B. Crossover and noncrossover pathways in mouse meiosis. Mol. Cell 20, 563–573 (2005).
Webb, A.J., Berg, I.L. & Jeffreys, A. Sperm cross-over activity in regions of the human genome showing extreme breakdown of marker association. Proc. Natl. Acad. Sci. USA 105, 10471–10476 (2008).
Gudbjartsson, D.F. et al. Sequence variants from whole genome sequencing a large group of Icelanders. Sci. Data 2, 150011 (2015).
Pratto, F. et al. Recombination initiation maps of individual human genomes. Science 346, 1256442 (2014).
Kong, A. et al. Common and low-frequency variants associated with genome-wide recombination rate. Nat. Genet. 46, 11–16 (2014).
Padhukasahasram, B. & Rannala, B. Meiotic gene-conversion rate and tract length variation in the human genome. Eur. J. Hum. Genet. http://dx.doi.org/10.1038/ejhg.2013.30 (2013).
Barrett, J.C. & Cardon, L.R. Evaluating coverage of genome-wide association studies. Nat. Genet. 38, 659–662 (2006).
Pardo-Manuel de Villena, F. & Sapienza, C. Recombination is proportional to the number of chromosome arms in mammals. Mamm. Genome 12, 318–322 (2001).
Duret, L. & Arndt, P.F. The impact of recombination on nucleotide substitutions in the human genome. PLoS Genet. 4, e1000071 (2008).
Martin, H.C. et al. Multicohort analysis of the maternal age effect on recombination. Nat. Commun. 6, 7846 (2015).
Myers, S., Freeman, C., Auton, A., Donnelly, P. & McVean, G. A common sequence motif associated with recombination hot spots and genome instability in humans. Nat. Genet. 40, 1124–1129 (2008).
Frazer, K.A. et al. A second generation human haplotype map of over 3.1 million SNPs. Nature 449, 851–861 (2007).
Assis, R. & Kondrashov, A.S. A strong deletion bias in nonallelic gene conversion. PLoS Genet. 8, e1002508 (2012).
Leushkin, E.V. & Bazykin, G.A. Short indels are subject to insertion-biased gene conversion. Evolution 67, 2604–2613 (2013).
Gudbjartsson, D.F. et al. Large-scale whole-genome sequencing of the Icelandic population. Nat. Genet. 47, 435–444 (2015).
1000 Genomes Project Consortium. An integrated map of genetic variation from 1,092 human genomes. Nature 491, 56–65 (2012).
Handel, M.A. & Schimenti, J.C. Genetics of mammalian meiosis: regulation, dynamics and impact on fertility. Nat. Rev. Genet. 11, 124–136 (2010).
Subramanian, V.V. & Bickel, S.E. Aging predisposes oocytes to meiotic nondisjunction when the cohesin subunit SMC1 is reduced. PLoS Genet. 4, e1000263 (2008).
Weng, K.A., Jeffreys, C.A. & Bickel, S.E. Rejuvenation of meiotic cohesion in oocytes during prophase I is required for chiasma maintenance and accurate chromosome segregation. PLoS Genet. 10, e1004607 (2014).
Leland, S. et al. Heterozygosity for a Bub1 mutation causes female-specific germ cell aneuploidy in mice. Proc. Natl. Acad. Sci. USA 106, 12776–12781 (2009).
Hodges, C.A., Revenkova, E., Jessberger, R., Hassold, T.J. & Hunt, P.A. SMC1β-deficient female mice provide evidence that cohesins are a missing link in age-related nondisjunction. Nat. Genet. 37, 1351–1355 (2005).
Nagaoka, S.I., Hassold, T.J. & Hunt, P.A. Human aneuploidy: mechanisms and new insights into an age-old problem. Nat. Rev. Genet. 13, 493–504 (2012).
Campbell, C.L., Furlotte, N.A., Eriksson, N., Hinds, D. & Auton, A. Escape from crossover interference increases with maternal age. Nat. Commun. 6, 6260 (2015).
Martini, E. et al. Genome-wide analysis of heteroduplex DNA in mismatch repair–deficient yeast cells reveals novel properties of meiotic recombination pathways. PLoS Genet. 7, e1002305 (2011).
Tsaponina, O. & Haber, J.E. Frequent interchromosomal template switches during gene conversion in S. cerevisiae. Mol. Cell 55, 615–625 (2014).
de Boer, E., Jasin, M. & Keeney, S. Local and sex-specific biases in crossover vs. noncrossover outcomes at meiotic recombination hot spots in mice. Genes Dev. 29, 1721–1733 (2015).
Wong, W.S.W. et al. New observations on maternal age effect on germline de novo mutations. Nat. Commun. 7, 10486 (2016).
Goldmann, J.M. et al. Parent-of-origin-specific signatures of de novo mutations. Nat. Genet. 48, 935–939 (2016).
Kong, A. et al. Rate of de novo mutations and the importance of father's age to disease risk. Nature 488, 471–475 (2012).
Efron, B. & Tibshirani, R.J. An Introduction to the Bootstrap (CRC Press, 1994).
Ihaka, R. & Gentleman, R.R. A language for data analysis and graphics. J. Comput. Graph. Stat. 5, 299–314 (1996).
van Rossum, G. & Drake, F.L. PYTHON Reference Manual (Centrum voor Wiskunde en Informatica, 1995).
Acknowledgements
This work was supported in part by the NIH (NIDA) (R01-DA017932).
Author information
Authors and Affiliations
Contributions
B.V.H., D.F.G. and K.S. designed the experiments. B.V.H. wrote the first draft of the manuscript. B.V.H., M.T.H., B.K., U.S., P.S., A.H., A.K., D.F.G. and K.S. reviewed and contributed to subsequent drafts of the manuscript. B.V.H., M.T.H. and A.G. implemented the methodology. B.V.H., M.T.H. and B.K. prepared tables and figures. Aslaug Jonasdottir and Adalbjorg Jonasdottir performed the Sanger sequencing. U.T. oversaw the operations of the genotyping facility. B.V.H., M.T.H., F.Z., G.T., A.G. and G.M. processed the data. B.V.H. and M.T.H. analyzed the data. All authors contributed to the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
All authors are employees of deCODE Genetics/Amgen.
Integrated supplementary information
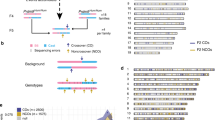
Supplementary Figure 1 Repair mechanisms of DNA double-strand breaks.
Small gray boxes show regions where heteroduplex DNA is formed. DSBs are resolved via either synthesis-dependent strand annealing (SDSA) or double Holliday junctions (dHjs). dHjs result in either dissolution or resolution; resolution results in either crossover (CO) or non-crossover (NCO).
Supplementary Figure 2 Odds ratio of an NCO gene conversion being in a male DSB region and a crossover recombination hotspot as a function of mother’s age in the chip data set.
Error bars, 95% confidence intervals. The dotted lines represent linear models.
Supplementary Figure 3 Crossover recombination rate of NCO gene conversions in the chip data set.
Error bars, 95% confidence intervals. The dotted line represents a linear model.
Supplementary Figure 4 MPPs by local GC content.
(a) Distribution of SNP MPPs by local GC content. (b) Probability that a SNP MPP is in a DSB region by local GC content.
Supplementary Figure 5 Number of verified gene-converted MPPs per event as a function of mother’s age in the chip data set.
Error bars, 95% confidence intervals. The dotted line represents a linear model.
Supplementary Figure 6 Detection of complex crossover gene conversions.
Two equally parsimonious locations exist to place a crossover recombination. The navy blue haplotype represents the background haplotype, and the cyan haplotype represents the other haplotype. The location of gene conversion depends on the assignment of the background haplotype.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7, Supplementary Tables 1–10 and Supplementary Note. (PDF 2505 kb)
Supplementary Data
Gene conversion MPPs. Data set of all gene conversion MPPs found and neighboring MPPs. (XLSX 1263 kb)
Rights and permissions
About this article
Cite this article
Halldorsson, B., Hardarson, M., Kehr, B. et al. The rate of meiotic gene conversion varies by sex and age. Nat Genet 48, 1377–1384 (2016). https://doi.org/10.1038/ng.3669
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3669
This article is cited by
-
Genetic architecture of individual meiotic crossover rate and distribution in Atlantic Salmon
Scientific Reports (2023)
-
Recombination rates in pigs differ between breeds, sexes and individuals, and are associated with the RNF212, SYCP2, PRDM7, MEI1 and MSH4 loci
Genetics Selection Evolution (2022)
-
Transmission ratio distortion of mutations in the master regulator of centriole biogenesis PLK4
Human Genetics (2022)
-
Comparative genome anatomy reveals evolutionary insights into a unique amphitriploid fish
Nature Ecology & Evolution (2022)
-
Analysis of the recombination landscape of hexaploid bread wheat reveals genes controlling recombination and gene conversion frequency
Genome Biology (2019)