Abstract
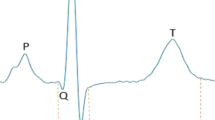
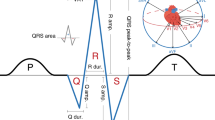
QT interval duration, reflecting myocardial repolarization on the electrocardiogram, is a heritable risk factor for sudden cardiac death and drug-induced arrhythmias. We conducted a meta-analysis of three genome-wide association studies in 13,685 individuals of European ancestry from the Framingham Heart Study, the Rotterdam Study and the Cardiovascular Health Study, as part of the QTGEN consortium. We observed associations at P < 5 × 10−8 with variants in NOS1AP, KCNQ1, KCNE1, KCNH2 and SCN5A, known to be involved in myocardial repolarization and mendelian long-QT syndromes. Associations were found at five newly identified loci, including 16q21 near NDRG4 and GINS3, 6q22 near PLN, 1p36 near RNF207, 16p13 near LITAF and 17q12 near LIG3 and RFFL. Collectively, the 14 independent variants at these 10 loci explain 5.4–6.5% of the variation in QT interval. These results, together with an accompanying paper, offer insights into myocardial repolarization and suggest candidate genes that could predispose to sudden cardiac death and drug-induced arrhythmias.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Straus, S.M. et al. Prolonged QTc interval and risk of sudden cardiac death in a population of older adults. J. Am. Coll. Cardiol. 47, 362–367 (2006).
Newton-Cheh, C. et al. QT interval is a heritable quantitative trait with evidence of linkage to chromosome 3 in a genome-wide linkage analysis: The Framingham Heart Study. Heart Rhythm 2, 277–284 (2005).
Newton-Cheh, C. & Shah, R. Genetic determinants of QT interval variation and sudden cardiac death. Curr. Opin. Genet. Dev. 17, 213–221 (2007).
Splawski, I. et al. Spectrum of mutations in long-QT syndrome genes. KVLQT1, HERG, SCN5A, KCNE1, and KCNE2. Circulation 102, 1178–1185 (2000).
Arking, D.E. et al. A common genetic variant in the NOS1 regulator NOS1AP modulates cardiac repolarization. Nat. Genet. 38, 644–651 (2006).
Bezzina, C.R. et al. A common polymorphism in KCNH2 (HERG) hastens cardiac repolarization. Cardiovasc. Res. 59, 27–36 (2003).
Pfeufer, A. et al. Common variants in myocardial ion channel genes modify the QT interval in the general population: results from the KORA study. Circ. Res. 96, 693–701 (2005).
Newton-Cheh, C. et al. Common genetic variation in KCNH2 is associated with QT interval duration: The Framingham Heart Study. Circulation 116, 1128–1136 (2007).
Gouas, L. et al. Association of KCNQ1, KCNE1, KCNH2 and SCN5A polymorphisms with QTc interval length in a healthy population. Eur. J. Hum. Genet. 13, 1213–1222 (2005).
Psaty, B.M. et al. Cohorts for Heart and Aging Research in Genomic Epidemiology (CHARGE) Consortium: design of prospective meta-analyses of genome-wide association studies from five cohorts. Circulation: Cardiovascular Genetics 2, 73–80 (2009).
Servin, B. & Stephens, M. Imputation-based analysis of association studies: candidate regions and quantitative traits. PloS Genet. 3, e114 (2007).
Devlin, B. & Roeder, K. Genomic control for association studies. Biometrics 55, 997–1004 (1999).
Pfeufer, A. et al. Common variants in ten loci modulate QT interval duration in the QTSCD study. Nat. Genet. advance online publication, doi:10.1038/ng.362 (22 March 2008).
Aarnoudse, A.J. et al. Common NOS1AP variants are associated with a prolonged QTc interval in the Rotterdam Study. Circulation 116, 10–16 (2007).
Lehtinen, A.B. et al. Association of NOS1AP genetic variants with QT interval duration in families from the Diabetes Heart Study. Diabetes 57, 1108–1114 (2008).
Post, W. et al. Associations between genetic variants in the NOS1AP (CAPON) gene and cardiac repolarization in the old order Amish. Hum. Hered. 64, 214–219 (2007).
Tobin, M.D. et al. Gender and effects of a common genetic variant in the NOS1 regulator NOS1AP on cardiac repolarization in 3761 individuals from two independent populations. Int. J. Epidemiol. 37, 1132–1141 (2008).
Eijgelsheim, M. et al. Identification of a common variant at the NOS1AP locus strongly associated to QT-interval duration. Hum. Mol. Genet. 18, 347–357 (2009).
Pietila, E. et al. Association between HERG K897T polymorphism and QT interval in middle-aged Finnish women. J. Am. Coll. Cardiol. 40, 511–514 (2002).
Sotoodehnia, N. et al. KCNE1 Gene D85N Variant, QT Interval, and Risk Of Mortality. Circulation 111, e231 (2005).
Marjamaa, A. et al. Common candidate gene variants are associated with QT interval duration in the general population. J. Intern. Med. advance online publication, doi:10.1111/j.1365-2796.2008.02026.x (25 October 2008).
Wei, J. et al. KCNE1 polymorphism confers risk of drug-induced long QT syndrome by altering kinetic properties of I-Ks potassium channels. Circulation 100, 495–495 (1999).
Paulussen, A.D. et al. Genetic variations of KCNQ1, KCNH2, SCN5A, KCNE1, and KCNE2 in drug-induced long QT syndrome patients. J. Mol. Med. 82, 182–188 (2004).
Salisbury, B.A. et al. The single nucleotide polymorphism D85N-KCNE1 is associated with both congenital and drug-induced long QT. Heart Rhythm 3(Suppl. 1), S98 (2006).
Splawski, I. et al. Variant of SCN5A sodium channel implicated in risk of cardiac arrhythmia. Science 297, 1333–1336 (2002).
Burke, A. et al. Role of SCN5A Y1102 polymorphism in sudden cardiac death in blacks. Circulation 112, 798–802 (2005).
Amsterdam, A. et al. Identification of 315 genes essential for early zebrafish development. Proc. Natl. Acad. Sci. USA 101, 12792–12797 (2004).
Qu, X. et al. Ndrg4 is required for normal myocyte proliferation during early cardiac development in zebrafish. Dev. Biol. 317, 486–496 (2008).
Milan, D.J. et al. Drugs that induce repolarization abnormalities cause bradycardia in zebrafish. Circulation 107, 1355–1358 (2003).
Luo, W. et al. Targeted ablation of the phospholamban gene is associated with markedly enhanced myocardial contractility and loss of beta-agonist stimulation. Circ. Res. 75, 401–409 (1994).
Schmitt, J.P. et al. Dilated cardiomyopathy and heart failure caused by a mutation in phospholamban. Science 299, 1410–1413 (2003).
Haghighi, K. et al. A mutation in the human phospholamban gene, deleting arginine 14, results in lethal, hereditary cardiomyopathy. Proc. Natl. Acad. Sci. USA 103, 1388–1393 (2006).
Splawski, I. et al. Severe arrhythmia disorder caused by cardiac L-type calcium channel mutations. Proc. Natl. Acad. Sci. USA 102, 8089–8096 (2005).
Street, V.A. et al. Mutation of a putative protein degradation gene LITAF/SIMPLE in Charcot-Marie-Tooth disease 1C. Neurology 60, 22–26 (2003).
Dawber, T., Kannel, W.B. & Lyell, L.P. An approach to longitudinal studies in a community: the Framingham Heart Study. Ann. NY Acad. Sci. 107, 539–556 (1963).
Kannel, W.B., Feinleib, M. & McNamara, P.M. An investigation of coronary heart disease in families: the Framingham Offspring Study. Am. J. Epidemiol. 110, 281–290 (1979).
Splansky, G.L. et al. The third generation cohort of the National Heart, Lung, and Blood Institute's Framingham Heart Study: design, recruitment, and initial examination. Am. J. Epidemiol. 165, 1328–1335 (2007).
Hofman, A. et al. The Rotterdam Study: objectives and design update. Eur. J. Epidemiol. 22, 819–829 (2007).
Hofman, A., Grobbee, D.E., de Jong, P.T. & van den Ouweland, F.A. Determinants of disease and disability in the elderly: the Rotterdam Elderly Study. Eur. J. Epidemiol. 7, 403–422 (1991).
Fried, L.P. et al. The Cardiovascular Health Study: design and rationale. Ann. Epidemiol. 1, 263–276 (1991).
van Bemmel, J.H., Kors, J.A. & van Herpen, G. Methodology of the modular ECG analysis system MEANS. Methods Inf. Med. 29, 346–353 (1990).
Rabbee, N. & Speed, T.P. A genotype calling algorithm for affymetrix SNP arrays. Bioinformatics 22, 7–12 (2006).
Li, Y. & Mach Abecasis, G.R. 1.0: Rapid haplotype reconstruction and missing genotype inference. Am. J. Hum. Genet. S79, 2290 (2006).
de Bakker, P.I. et al. Practical aspects of imputation-driven meta-analysis of genome-wide association studies. Hum. Mol. Genet. 17, R122–R128 (2008).
Pe'er, I., Yelensky, R., Altshuler, D. & Daly, M.J. Estimation of the multiple testing burden for genomewide association studies of nearly all common variants. Genet. Epidemiol. 32, 381–385 (2008).
Acknowledgements
The Framingham Heart Study work was conducted in part using data and resources from the Framingham Heart Study of the National Heart Lung and Blood Institute of the National Institutes of Health and Boston University School of Medicine (contract #N01-HC-25195) , its contract with Affymetrix for genotyping services (contract #N02-HL-6-4278), and the Doris Duke Charitable Foundation (C.N.-C.) and Burroughs Wellcome Fund (C.N.-C.). The Framingham analyses reflect intellectual input and resource development from the Framingham Heart Study investigators participating in the SNP Health Association Resource (SHARe) project. A portion of this research utilized the Linux Cluster for Genetic Analysis (LinGA-II) funded by the Robert Dawson Evans Endowment of the Department of Medicine at Boston University School of Medicine and Boston Medical Center. The measurement of ECG intervals in Framingham Heart Study generation 1 and 2 samples was done by eResearchTechnology and supported by an unrestricted grant from Pfizer. The measurement of ECG intervals in the Framingham Heart Study generation 3 sample was completed by A. Hirji and S. Kovvali using AMPS software provided through an unrestricted academic license by Analyzing Medical Parameters for Solutions (A.M.P.S., LLC). The Rotterdam Study is funded by Erasmus Medical Center and Erasmus University, Rotterdam, Netherlands Organization for the Health Research and Development (ZonMw), the Research Institute for Diseases in the Elderly (RIDE), the Ministry of Education, Culture and Science, the Ministry for Health, Welfare and Sports, the European Commission (DG XII), and the Municipality of Rotterdam. The generation and management of GWAS genotype data for the Rotterdam Study is supported by the Netherlands Organisation of Scientific Research NWO Investments (#175.010.2005.011, 911-03-012). This study is funded by the Research Institute for Diseases in the Elderly (014-93-015; RIDE2), and the Netherlands Genomics Initiative (NGI)/Netherlands Organisation for Scientific Research (NWO) project #050-060-810. The CHS research reported in this article was supported by contract numbers N01-HC-85079 through N01-HC-85086, N01-HC-35129, N01 HC-15103, N01 HC-55222, N01-HC-75150, N01-HC-45133, grant numbers U01 HL080295 and R01 HL087652 from the National Heart, Lung, and Blood Institute, with additional contribution from the National Institute of Neurological Disorders and Stroke. DNA handling and genotyping was supported in part by National Center for Research Resources grant M01RR00069 to the Cedars-Sinai General Clinical Research Center Genotyping core and National Institute of Diabetes and Digestive and Kidney Diseases grant DK063491 to the Southern California Diabetes Endocrinology Research Center. A full list of principal CHS investigators and institutions can be found at http://www.chs-nhlbi.org/pi.htm. The authors acknowledge the essential role of the CHARGE (Cohorts for Heart and Aging Research in Genome Epidemiology) Consortium in development and support of this manuscript. CHARGE members include the Netherland's Rotterdam Study, the NHLBI's Atherosclerosis Risk in Communities (ARIC) Study, Cardiovascular Health Study (CHS) and Framingham Heart Study (FHS), and the NIA's Iceland Age, Gene/Environment Susceptibility (AGES) Study. C.N.-C. is supported by US National Institutes of Health grant K23-HL-080025, a Doris Duke Charitable Foundation Clinical Scientist Development Award, and a Burroughs Wellcome Fund Career Award for Medical Scientists. M.E. is funded by the Netherlands Heart Foundation (#2007B221). J.I.R. is supported by the Cedars-Sinai Board of Governors' Chair in Medical Genetics. The authors wish to thank the following people: G. Crawford and C. Guiducci (Broad Institute of Harvard and Massachusetts Institute of Technology) who completed the Sequenom-based technical validation genotyping of the Framingham Heart Study samples; P. Arp and M. Jhamai (Erasmus Medical Center) for Illumina array genotyping and Taqman-based technical validation genotyping of the Rotterdam Study samples; and Dr. M. Moorhouse, M. Verkerk and S. Bervoets (Erasmus Medical Center) for database management in the Rotterdam Study; and the study participants, the staff from the Rotterdam Study and the participating general practitioners and pharmacists. The QTGEN consortium would like to thank the QTSCD consortium for the opportunity to exchange top results pre-publication.
Author information
Authors and Affiliations
Contributions
Framingham Heart Study: M.G.L., C.N.-C., P.A.N., C.J.O., X.Y.
Rotterdam Study: M.E., K.E., A.H., J.A.K., F.R., B.H.Ch.S., A.G.U., J.C.M.W.
Cardiovascular Health Study: J.C.B., S.R.H., T.L., K.M., C.N.-C., B.M.P., K.M.R., J.I.R., N.L.S., N.S.
Broad Institute of Harvard and Massachusetts Institute of Technology: P.I.W.dB., C.N.-C.
Design of QTGEN study: P.I.W.dB., M.E., M.G.L., T.L., C.N.-C., C.J.O., B.M.P., K.M.R., B.H.Ch.S. Genotyping: Affymetrix, C.N.-C., F.R., J.I.R., A.G.U. Statistical analysis and informatics: J.C.B., P.I.W.dB., M.E., K.E., T.L., K.M., C.N.-C., K.M.R., F.R., A.G.U., X.Y. Drafting of manuscript: C.N.-C. Critical revision of manuscript: J.C.B., P.I.W.dB., M.E., K.E., S.R.H., A.H., J.A.K., P.A.N., B.M.P., K.M.R., J.I.R., N.L.S., N.S., B.H.Ch.S., J.C.M.W.
Corresponding authors
Supplementary information
Supplementary Text and Figures
Supplementary Figure 1, Supplementary Tables 1–4 and Supplementary Methods (PDF 464 kb)
Rights and permissions
About this article
Cite this article
Newton-Cheh, C., Eijgelsheim, M., Rice, K. et al. Common variants at ten loci influence QT interval duration in the QTGEN Study. Nat Genet 41, 399–406 (2009). https://doi.org/10.1038/ng.364
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.364
This article is cited by
-
Dosage of the pseudoautosomal gene SLC25A6 is implicated in QTc interval duration
Scientific Reports (2023)
-
Association of T66A polymorphism in CASQ2 with PR interval in a Chinese population
Herz (2021)
-
Rat models of human diseases and related phenotypes: a systematic inventory of the causative genes
Journal of Biomedical Science (2020)
-
Serum neurofilament light levels in normal aging and their association with morphologic brain changes
Nature Communications (2020)
-
Exome sequencing identifies a novel nonsense mutation of Ring Finger Protein 207 in a Chinese family with Long QT syndrome and syncope
Journal of Human Genetics (2019)